4.4: Exercise 1 - Streak plates
- Page ID
- 17513
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Microbiologists like to begin their experiments with a single colony, because the cells in a colony are the progeny of a single cell. A concern in all genetic experiments is unknown mutations that arise spontaneously and may affect the phenotype being studied. Spontaneous mutations arise constantly in all cells, with a rate of approximately 10-8/base/ generation. For S. cerevisiae, with a genome of 12 Mbp, most cells will have accumulated at least one mutation by the time that they have undergone 9-10 divisions. A colony, which has hundreds of millions of cells, is therefore a population of genetically very similar, but not completely identical, organisms.
Researchers commonly use streak plates to isolate single colonies. A streak plate is actually a serial dilution of an existing culture on solid media. Researchers begin a streak by picking up a small sample of yeast or another microorganism with a sterile instrument, which could be a platinum loop, a toothpick or micropipette tip. They then spread the culture by making a series of zig-zag strokes across the surface of the plate. The number of cells on the loop or toothpick decreases as the streak progresses. Consequently, streaks appear thickest at their starting points, and the streak thickness decreases until it is possible to detect well-isolated single colonies near the end of the streak. Because it may be difficult to resolve colonies from a single streak, many labs use a series of streaks on the same plate to separate colonies. Each new streak
is done with a freshly sterilized loop that picks up cells by crossing over the tracks of the previous streak, before beginning a new series of zig-zags. In our experiments, we will use a multi-streak protocol, which allows us to culture multiple strains on a single plate of culture medium. (See the figure below.) The streak plates that you prepare will contain the stocks for future experiments. As you streak your strains, pay careful attention to detail to avoid cross-contamination or confusion about the identities of individual strains.
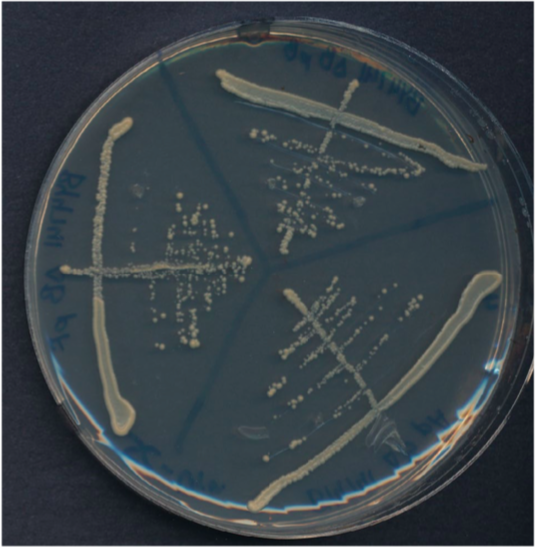
Streak plate with three sectors.
Plate has been divided into three clearly labeled sectors. Three streaks were used to spread the cells in each sector. The third streak in each sector contains well-separated colonies that can be used for genetics experiments.
Preparing a streak plate
- Your team will be assigned three different S. cerevisiae met strains to culture. The stock strains that you will use have been grown to a high density in liquid YPD medium. Gather the YPD cultures of the parent strains to be propagated, a platinum inoculation loop, and an agar plate with fresh YPD media. This plate will serve as your team’s stock plate for the semester.
- Divide your team’s stock plate into sectors by marking the bottom of the plate with a magic marker. CLEARLY label each sector with a code for the strain that will be streaked in it. As series of three streaks will be made within each sector, as described on the previous page. Keep the labels at the rim of the plate and use small letters. Note your initials and the date.

- Sterilize the inoculation loop by holding it in the flame of a Bunsen burner until it glows red. Cool the loop by briefly touching the surface of your agar plate before proceeding. (A hot loop will kill yeast cells that it contacts!)
- The cells in your stock cultures may have settled over time. Resuspend the cells using the vortex mixer. Working quickly, remove the cap the first culture tube. Dip the sterilized loop into the culture, remove the loop and place the cap back on the culture tube.
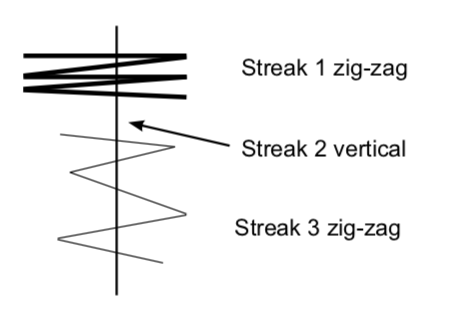
- Transfer cells to the appropriately labeled sector on your stock plate. First streak: make several zigzags across the outside edge of the sector with the loop. LIGHTLY touch the agar surface as you move the toothpick. Think of pushing a hockey puck across an ice rink, rather than digging a ditch. You do NOT want to put track marks in the agar! Replace the lid.
- Sterilize the loop in the flame and cool it on the agar surface before beginning the second streak. Make a second vertical streak from the rim of the plate toward the center, staying within the sector. The streak should cross the zigzags in the first streak. Close the lid and re-sterilize (and cool) the loop.
- Third streak: make a third series of zigzags that cross back and forth over the straight second streak, beginning at the outer edge of the plate and moving toward the center. Be careful to stay within the sector. Close the lid and sterilize the loop.
- Repeat steps 4-7 for each of your liquid cultures.
- Invert the plate and incubate it at 30 ̊C until individual colonies are visible, which is usually 24-48 hours.


