10.2: Making Metazoans
- Page ID
- 4860
As we move from quorum sensing between organisms of the same type, we come to what are known as biofilms. These are microbial communities, such as the plaque that forms on your teeth, and consist of a number of different types of often co-dependent organisms. While horizontal gene transfer may occur between these different organisms, they remain distinct and give rise to organisms genetically related to their parent(s).
The next obvious level of organization is what we will call a discrete colonial organism. In such organisms individual cells are attached to one another, generally through the extracellular materials they secrete. They gain advantages associated with larger size (for example, they may be able to swim faster or become too big to easily swallow) but these advantages are constrained by the fact that the individual cells retain their individuality.Moreover, in a pure colonial organism, each cell within the colony retains its ability to reproduce independently, either sexually or asexually. Previously we introduced the terms soma for the cells of the body that reproduce asexually and are responsible for the growth and repair of the organism, and the germ line,the cells that are responsible for producing the next generation of organisms. In a purely colonial organism, all cells are potential germ cells. There is no central system for coordinating behavior.
So we might ask, what is the next step in the evolution of a truly integrated multicellular organism, one with a soma and a germ line? It appears that there have been a number of paths to multicellularity: for example animals, plants, and fungi all appear to have independent strategies (lineages). We can use modern organisms a modern bestiary to illustrate these strategies310. This is not to claim that any represent real ancestors, all are modern organisms, well adapted to their current environment and the result of their own evolutionary history. Never the less they have dealt with various aspects of multicellular coordination and differentiation in interesting ways.
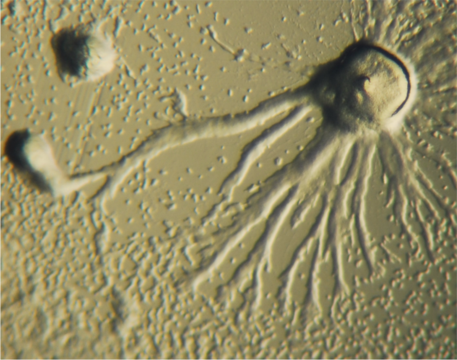
Figure \(\PageIndex{1}\): Dictyostelium showing aggregation behavior. Image from Wikimedia Commons (“Dictyostelium Aggregation” by Bruno in Columbus – Own work. Licensed under Public domain via Wikimedia Commons – http://commons.wikimedia.org/ wiki/File:Dictyostelium_Aggregation.JPG)
Consider the eukaryotic slime mold Dictyostelium discoideum. Cellular slime molds live in soil and eat bacteria - they are unicellular predators. Most of the time they are small, amoeba-like, haploid cells. Upon starvation they can undergo a dramatic aggregation process. Aggregation is triggered by the release from individual cells of pulses of cyclic adenosine monophosphate (cAMP); a process analogous to quorum sensing in bacteria (see above). The result is that individual cells begin to migrate up the cAMP concentration gradient, where they interact with and adhere to one another. Groups of cells produce more cAMP leading to the formation of cellular aggregates known as slugs that contain between 10,000 to 100,000 discrete cells. Slugs migrate in a coordinated manner. Eventually a slug will stop its migration and begin a process of known as differentiation, a process involves changes in gene expression and cellular behavior. Some of the cells of the slug differentiate to form stalk cells. The coordinated elongation of these stalk cells lifts the rest of the slug “body” into the air.The non-stalk cells differentiate to form spores, cells like the quiescent persisters we mentioned above. When released into the air, the spores are widely dispersed and, if they land in an appropriate environment, can go on to form single celled amoebae.
By now you may be able to generate a plausible scenario to explain exactly how the self-sacrificing behavior of stalk cells is possible. The answer lies in inclusive fitness. The purpose of the slug and stalk are to enable Dictyostelium cells to escape a hostile environment and colonize new, more hospitable environments. In fact, in a number of cases the spores carry with them bacteria that inoculate their new environments; these are bacteria that the amoeba can eat. The slime mold could be considered migrating bacterial farmers311. Since individual Dictyostelium amoeboid cells can not migrate far, most of the cells in any particular region, that is the cells that combine to form a slug, are likely to be closely related to one another - they are part of a clone. The sacrifice of the stalk cells is more than made up for by the increased chance that the spore cells will survive and produce lots of offspring. Of course there is a danger that some cells will diverge (through mutation) and cheat the system. That is, they will avoid becoming stalk cells. Such cheating has been observed in wild type Dictyostelium and cheating is a challenged faced by all multicellular systems. There are a number of strategies that are used to suppress cheaters, generally they are similar to those exploited in the context of quorum sensing312.
An organism that displays a distinct type of differentiation behavior is the bacterium Caulobacter crescentus. Under certain conditions it will produce a distinct type of cell, a stalk cell, which attaches to a surface. It canndivide to produce swimming cells that can migrate away and colonize new surfaces. The stalk cell can continue to produce swimming cells, and the swimming cells can settle down and transform into stalk cells. C. crecentus has established two different cell types designed to exploit two distinct environments.
References
310 The medieval bestiary: http://bestiary.ca
311 Small molecules mediate bacterial farming by social amoebae. http://www.ncbi.nlm.nih.gov/pubmed/23975931
312 Kin Recognition Protects Cooperators against Cheaters: http://www.ncbi.nlm.nih.gov/pubmed/23910661
Contributors and Attributions
Michael W. Klymkowsky (University of Colorado Boulder) and Melanie M. Cooper (Michigan State University) with significant contributions by Emina Begovic & some editorial assistance of Rebecca Klymkowsky.


