B9. Excitatory Neurotransmitters
- Page ID
- 5597
Glutamate is a major excitatory neurotransmitter in the brain. Four types of glutamate receptors are found in the central nervous system. They differ in the nature of ligands which bind to the receptor and which act as agonists.
Figure: Glutamate receptor ligands
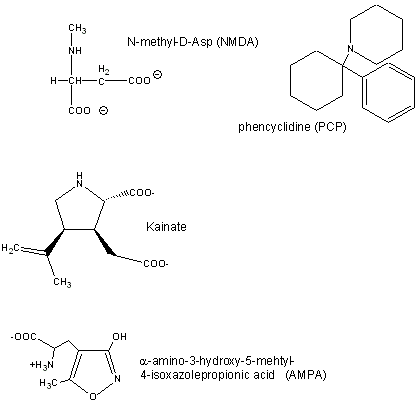
Excessive amounts of glutamate are neurotoxic. Three of the receptor types are briefly described below:
- AMPA receptor binds α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid and acts as a sodium and potassium ion channel.
- Kainate receptor, named after the molecule kainate which acts as an agonist, is a a sodium and potassium ion channel.
- NMDA (N-methyl-D-Asp) receptor allows flow of sodium, potassium and calcium and is regulated by glycine and Zn2+. The street drug PCP (phencyclidine) inhibits this channel.
Sobolevsky et al have recently analyzed the structure, symmetry and mechanism of the rat GluA2 receptor, a homotetramer, a bound competitive antagonist, ZK 200775. GluA2, an AMPA type receptor, has structural similarities to an NMDA type receptor. In order to determine the structure of the wild type GluA2 receptor, a model receptor polypeptide that was termed GluA2cryst (PDB# 3KG2) was crystallized. Isolated GluA2 subunits were used to piece together a detailed model of rat GluA2 as it would exist in its natural membrane bound form. The receptor has three main domains that are essential for its function. The first is a large extracellular amino-terminal domain (ATD). This domain serves as a guide for ligand binding in the next domain, the ligand-binding domain (LBD). The ligand binding domain participates in agonist/antagonist competitive binding that regulates the gating of the ion channel. The final domain is the transmembrane domain (TMD) that forms the ion channel for molecules to pass through the cell membrane.
The GluA2 receptor is shaped like a “Y” with the domains layered on top of each other. The TMDs form the transmembrane section and base on the outside of the membrane. The ATDs protrude outward like the “arms” of a Y and the LBD is on top of the TMD. This process gates the transmembrane channel as can be seen in Figure 1. Analysis of the crystallized protein uncovered a bound antagonistic ligand proving that the ligand binding occurs in a domain (LBD) rather than between domains, as was previously conceived.
The authors studied the gating mechanism between the LBDs and TMDs by analyzing the change in symmetry that occurs when the gate is opened and closed. When the gate is closed, the LBDs exhibit a two-fold symmetry and the TMDs four-fold symmetry. When the ligand binds to the LBDs an abrupt structural change occurs creating four fold symmetry in the LBDs. This transition was measured by calculating the r.m.s.d. of the alpha carbons between the subunits undergoing the structural change. The TMDs also underwent a conformational change when ligand binding occured and ions travelled down the channel. In order to accommodate the ion transfer, several subunits within the TMDs loosened their helix and provided an open pathway for the ion to travel in to the cytoplasm of the cell.


