A7. Biological Uses for Reactive Oxygen Species - ROS
- Page ID
- 5554
ROS can be used in the body as defense mechanisms. The can be generated by immune cells that phagocytize (engulf) bacteria and other foreign substances. After uptake, these immune cells undergo an oxidative burst, increasing the amount of dioxygen that they consume, leading to production of superoxide and hydrogen peroxide which can be used to kill the engulfed bacteria. Recent articles by Wentworth et al. go even farther. They demonstrated that antibody molecules (protein which bind to foreign molecules and target them for clearance or further immune response) can also generate ROS when they bind to their target. Such an outcome was totally unexpected, but not inconsistent with the observation that antibodies can have catalytic activity if made against transition state analogs of substrates. Over 100 different antibodies were found to generate hydrogen peroxide through the reaction of singlet oxygen (possibly generated by phagocytotic neutrophils during oxidative bursts) with water to form H2O2. (We have previously discussed the idea that singlet O2 can be generated from ground state O2 by excitation through UV light or through collisional activation with an excited state chromophore - a conjugated alkene or aromatic molecule). They postulate the following reaction to form hydrogen peroxide. The oxygen in the peroxide comes from water, as shown by labeling water with 18O:
\[\mathrm{2H_2O + 1O_2\: (singlet\: oxygen) \rightarrow H_2O_3 + H_2O \rightarrow H_2O_2 \:+\: ?}\]
Antibodies show this effect to a much greater extent that other proteins studied. Using UV irradiation as a source of energy, the rate of production of H2O2 was linear with time, but decreased from the initial rate as H2O2 concentrations increased. If it was removed with catalase, the initial rate returned to normal, suggesting that H2O2 was a product inhibitor of the reaction. The reactions also saturated with dioxygen as well. The maximal wavelength of UV irradiation to produce H2O2 was identical to the maximal wavelength of absorbance for tryptophan in the protein. The excited state tryptophan presumably deexcites by energy transfer to triplet dioxygen (3O2 ) in much the manner described earlier for photobleaching. A significant source of electrons needed to be found to account for the production of so much H2O2 which if it was made from singlet oxygen would require 2 electrons for each H2O2. These numbers of electrons could not come from photooxidation of aromatic amino acids in the protein. Hence they theorized that water would add as a nucleophile to the singlet oxygen, providing the source of electrons required to make H2O2. This reactions would go through the H2O3 intermediate.
In another recent study (Nov. 02), the same team discovered that antibodies can not only produce H2O2 but also ozone. O3 is used in killing pathogens but also might lead to inflammation. In their previous work described above, they generated singlet oxygen in a "non-physiological" way - through UV irradiation of the antibodies. They also didn't show that the ROS produced can kill bacteria without any other immune mediators. They used a new singlet oxygen generating system which led to insufficient peroxide formation to account for the extensive killing of E. Coli that they observed (95%). This led them to look for the generation of a more potent oxidizing agent - which they believe is consistent with the formation of ozone. They also showed that neutrophils were the source of the required singlet oxygen as described above.
In an additional study (Dec 02), the showed that antibodies in the presence of singlet oxygen and in the absence of immune cells or other immune proteins (like complement proteins) can "destroy antigenic target". They also confirmed that ozone is produced by antibodies during bacterial killing by neutrophils (which produce singlet oxygen) and in inflammatory states. Signaling through ozone might amplify other immune molecules that signal the activation (or hyperactivation) of the immune system.
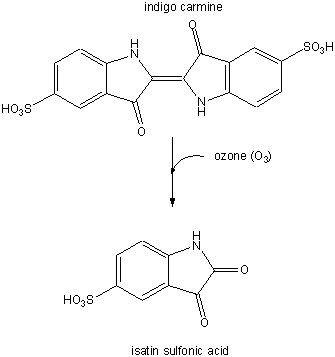
In a subsequent study (Nov. 03) they examined the involvement of ozone (generated from antibodies in the presence of singlet O2 from neutrophils) in the oxidation of lipoproteins, a process which has been associated with atherosclerosis in coronary arteries. First they tested if ozone could be generated by atherosclerotic lesions treated with phorbol myristate acetate. 14/15 plaques produced ozone, as evidenced by the photobleaching of the dye indigo carmen to isatin sulfonic acid. This conversion, known to be specific for ozone, is shown below.
Figure: Conversion of indigo carmen to isatin sulfonic acid
They then looked for the telltale signature of the reaction of ozone with plaque cholesterol (cleavage of the Δ5,6 double bond to form 5,6-secosterol as shown below and found it. This reaction is known not to occur with other oxygen oxidants, including both triplet and singlet oxygen, superoxide, and the hydroxyl free radical. A secondary reaction product aldol condensation product was also produced. They named these new products atheronals.
Figure: Cholesterol/ozone reaction
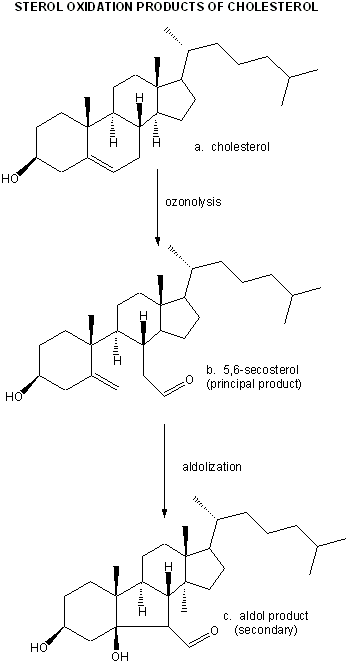
Figure: Ozonolysis mechanism
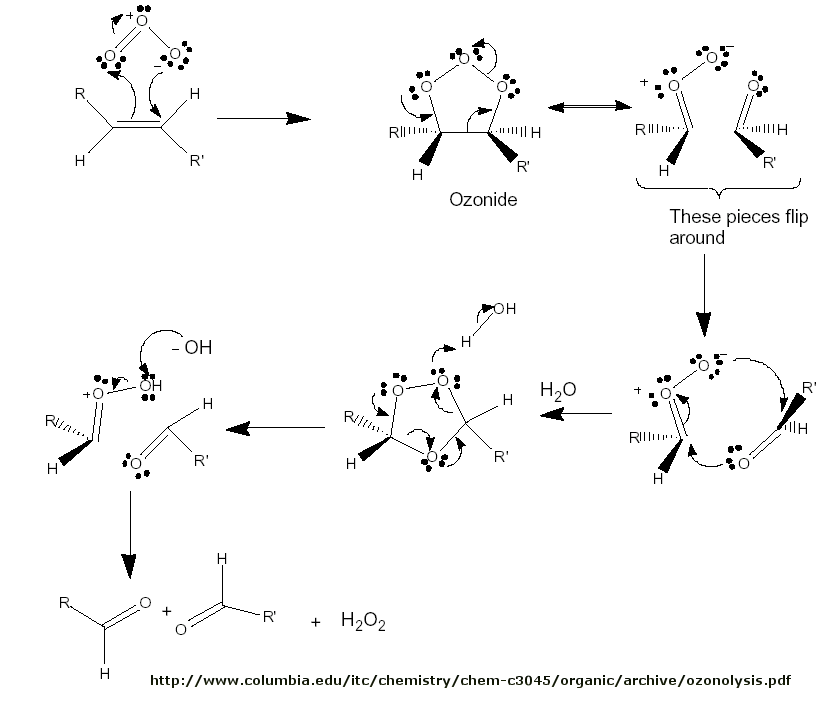
Figure:
Significant increases in secosterol was found in plaques treated with the phorbal ester. 6 of 8 patients with atherosclerosis had elevated plasma levels of the secosterol, while only 1 of 15 of control patients did. They also showed that these ozone products caused normal macrophages to fill with cholesterol (presumably from oxidized LDL), and that the structure of the LDL apoprotein B100 changed. These findings might provide a link between the immune system and cholesterol in the generation of cardiovascular disease.


