C. Cofactors and Electron Pushing: Sources and Sinks
- Page ID
- 4603
Learning Objectives
- explain the general role of cofactors in enzyme-catalyzed reactions;
- identify from their structures thiamine, nicotinamide, and pyridoxal cofactors;
- draw mechanisms to show how thiamine pyrophosphate facilitates decarboxylation of α-ketoacids as a cofactor in enzyme-catalyzed reactions;
- draw mechanisms to show how nicotinamide adenine dinucleotide facilitates oxidation, oxidative decarboxylaitn and oxidative deamination reactions as a cofactor in enzyme-catalyzed reactions;
- draw mechanisms to show how pyridoxal phosphate facilitates cleavage of all bonds to the α-carbon in amino acids as a cofactor in enzyme-catalyzed reactions;
To make and break bonds, electrons have to be moved. In drawing reaction mechanisms, we showed how electrons moved from "sources" to "sinks". In many enzyme-catalyzed reactions, vitamin derivatives are uses as substrates or "cofactors" or "coenzymes" to facilitate the flow of electrons in bond making and breaking. For each of the reactions below, using the analogy of source/sink, write a reasonable mechanism which shows electron flow during the reaction. With the exception of the first reaction, all require a vitamin derivative to facilitate electron flow. Imagine reaction one occurring spontaneously in solution in the absence of an enzyme (although enzymes exists to catalyze this reaction). The rest of the reactions involve vitamin derivatives as part of an enzyme-catalyzed reaction.
Draw plausible mechanisms for each of the reactions below, showing the flow of electrons from a source to a sink. The source might often be a pair of electrons on an anion which was formed by prior removal of a proton from the atom by a general base. A sink could be a carbonyl O which receives a pair of electrons from one of the double C-O bonds of the carbonyl. As a bond is made to the carbonyl, one of the double bonds must break with the electrons going (temporarily if the reaction is a nucleophilic substitution reaction) to the carbonyl O, an excellent sink since it is so electronegative. An even better sink is a positive N of an iminium ion, examples of which are show below.
The cofactors encountered in enzyme catalyzed reactions are often vitamin derivatives. We will study more about some of these later. Just the "business parts" of the cofactors are shown below.
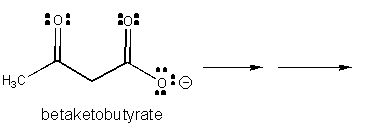
1. RX TYPE - SPONTANEOUS DECARBOXYLATION OF β-KETO ACIDS (No cofactor required, although nucleophilic catalysis by an amine through Schiff Base formation will speed up the reaction.)
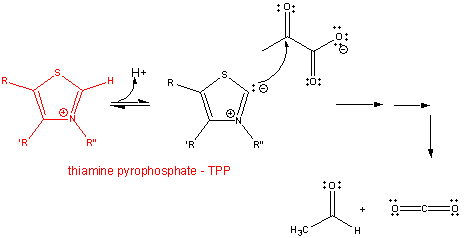
2. RX TYPE - DECARBOXYLATION OF α-KETO ACIDS- requires thiamine pyrophosphate - TPP, a derivative of thiamine - vitamin B1, a deficiency of which causes beriberi. TPP is covalently attached to the enzyme, such as in pyruvate dehydrogenase and alpha-ketoglutarate dehydrogenase. The first part of the reaction. The number of arrows leading to product does not reflect the actual number of step.
![]()
![]() Jmol: THIAMIN DIPHOSPHATE DEPENDENT ENZYME PYRUVATE DECARBOXYLASE (1pyd)
Jmol: THIAMIN DIPHOSPHATE DEPENDENT ENZYME PYRUVATE DECARBOXYLASE (1pyd)
(go to left hand Display Molecule, choose Jmol viewer).
3. RX TYPE - REDOX USING NAD+. Requires the vitamin nicotinic acid, also called niacin (nicotinic acid vitamin. A deficiency causes pellagra. It is chemically modified to form NAD+.
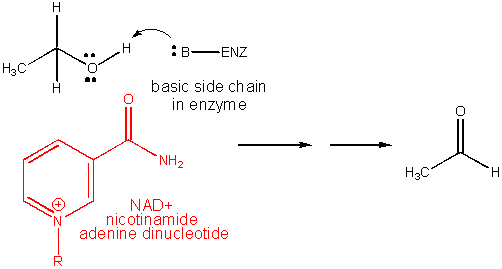
4. RX TYPE - OXIDATIVE DECARBOXYLATION.
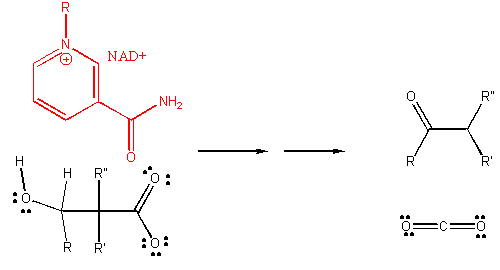
5. RX TYPE - OXIDATIVE DEAMINATION (hint: use NAD before water in the mechanism).
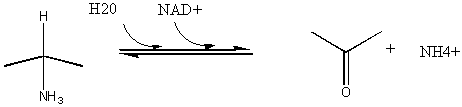
![]() Jmol: Updated Glyceraldehyde-3-phosphate dehydrogenase (NAD) Jmol14 (Java) | JSMol (HTML5)
Jmol: Updated Glyceraldehyde-3-phosphate dehydrogenase (NAD) Jmol14 (Java) | JSMol (HTML5)
PYRIDOXAL PHOSPHATE ENZYMES
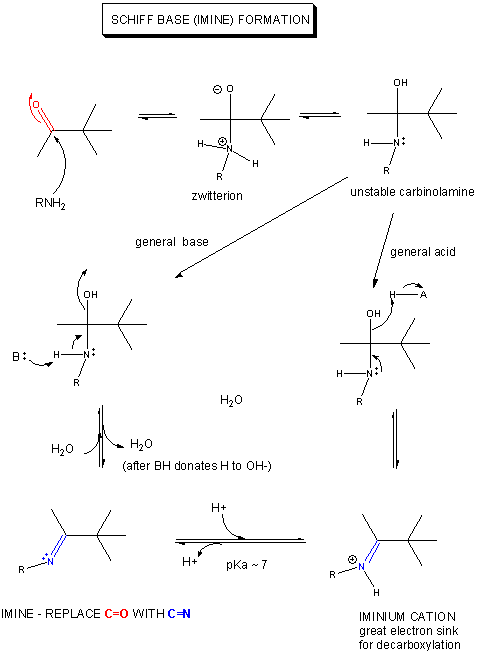
Pyridoxal phosphate (PLP) is a derivative of the vitamin B6 or pyridoxal. Deficiencies cause convulsions, chronic anemia, and neuropathy. It assists in many reactions (catalyzed by PLP-dependent enzymes). The PLP is bound covalently to lysine residues in a Schiff base linkage (aldimine). In this form, it reacts with many free amino acids (as substrates) to replace the Schiff base to Lys of the enzyme with a Schiff base to the amino acid substrate. First a review of Schiff Base formation.
PLP: Structure and Covalent Attachment to Enzyme
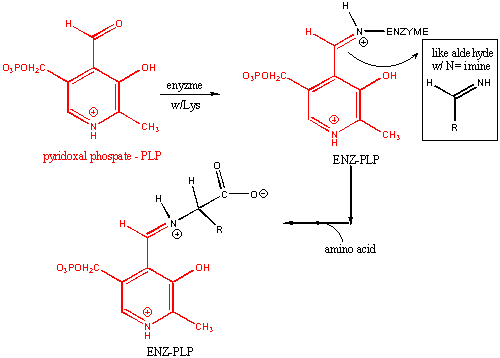
For reactions 6-8, assume that the amino acid substrate is in a Schiff base with PLP.
William Jencks, in his classic text, Catalysis in Chemistry, wrote:
"It has been said that God created an organism especially adapted to help the biologist find an answer to every question about the physiology of living systems; if this is so, it must be concluded that pyridoxal phosphate was created to provide satisfaction and enlightenment to those enzymologists and chemists who enjoy pushing electrons, for no other coenzyme is involved in such a wide variety of reactions, in both enzyme and model systems, which can be reasonably interpreted in terms of the chemical properties of the coenzyme. Most of these reactions are made possible by a common structural feature. That is, electron withdrawal toward the cationic nitrogen atom of the imine and into the electron sink of the pyridoxal ring from the alpha carbon atom of the attached amino acid activates all three of the substitutents of this carbon for reactions which require electron withdrawal from this atom."
![]()
![]() Molecular Modeling: PLP: Tyrosine Aminotransferase Jmol (from PDB)
Molecular Modeling: PLP: Tyrosine Aminotransferase Jmol (from PDB)
6. RX TYPE - α-DECARBOXYLATION OF AMINO ACIDS.
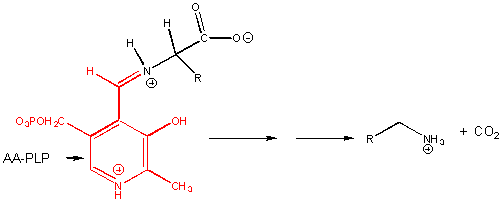
7. RX TYPE - BETA-ELIMINATION FROM SERINE. Example: Serine dehydratase. (hint: remove H on α-C first), then OH)
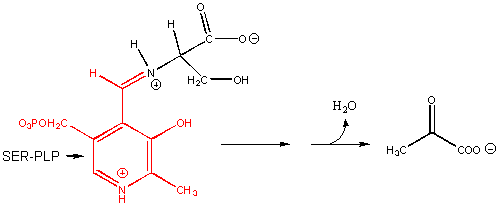
8. RX TYPE - RACEMIZATION OF AMINO ACIDS. (hint: remove H on α-C first)
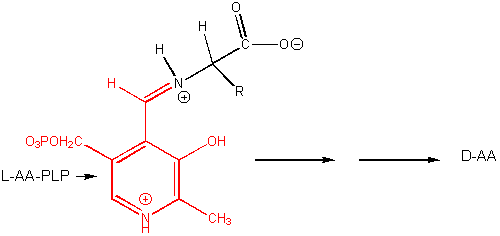
PLP enzymes also catalyze transamination reactions, an example of which is shown below:
Amino Acid 1 + α-keto acid 1 <==> α-keto acid 2 + Amino Acid 2 For example:
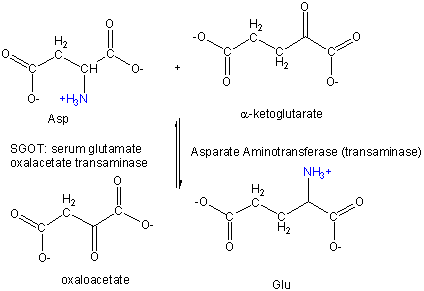
First Asp, bound to PLP through a Schiff base link, loses the α-H , forms a ketimine through a tautomerization reactions, which ultimately hydrolyzes to form the released oxalacetate and pyridoxamine. The pryidoxamine reacts with α-ketoglutarate in the reverse of the first three reactions to from Glu.
9. RX TYPE - ACETYLATION: The "acetic anhydride" of biological acetylation reactions is acetyl-CoA, a derivative of the vitamin pantathenic acid, which contains a free thiol. It is acetylated at the thiol in many metabolic reactions to produce acetylCoA containing a thioester bond, a biological acetylating reagent. This molecule can be cleaved in an exergonic fashion due in part to the weak bond between the acetyl C and the Sk leading to transfer of the acetyl group. We have previously discussed the importance of histone Lys acetylation by histone acetylases in the control of gene expression.
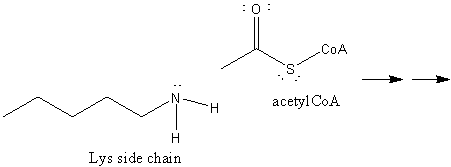
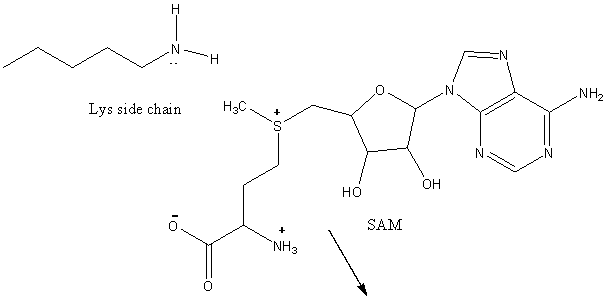
10. RX TYPE - METHYL TRANSFERASES:
An additional post-translational modification of histone proteins, methylation, is yet another method in which gene transcription is control. Nature's methylating agent is S-adenosine methionine, a derivative of methione which is methylated by yet another vitamin derivative, tetrahydrofolate (from folic acid). SAM is a substrate for methylating enzymes that transfer a methyl group to proteins and DNA. The reaction below envisions the methylation of a protein at a Lys side chain.