1.12: Molecular Visualization of an Enzyme, Acetylcholinesterase
- Page ID
- 18915
12.1 Learning Objectives
The goal of this laboratory is to analyze some of the major structural elements of an important enzyme, acetylcholinesterase (AChE). To do this, you will use a common structural visualization program and correlate AChE structural elements with the enzyme mechanism. You will be using Chimera, a state-of-the-art molecular visualization program provided by the National Science Foundation through the University of California, San Francisco. This free program is available at http://www.cgl.ucsf.edu/chimera/.
To provide a quick overview of the program, we will look at a multi-drug resistance efflux pump - the protein that is controlled by the riboswitch you studied. You will then use this program to analyze the enzyme, acetylcholinesterase.
12.2 Introduction and Background
Acetylcholinesterase (AChE) destroys the nerve transmitter, acetylcholine by hydrolysis.

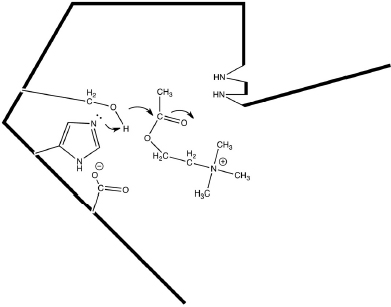
AChE is one of the most efficient enzymes in nature - in some ways, a “perfect” enzyme. Amino acid side chains at the active site are precisely arrayed to force bonding changes in the acetylcholine (Fig. 12.1).
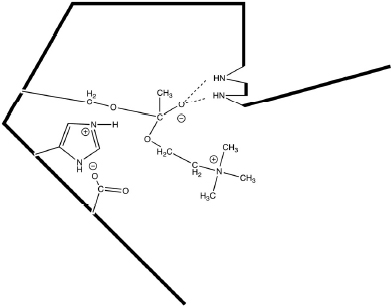
Overall, the electrons are pushed toward the ester carbonyl forming a covalent intermediate between the reactant and the enzyme (Fig. 12.2).
Acetylcholinesterase’s finely tuned mechanism provides a good illustration of some common catalytic processes:
- This week you will examine this enzyme at a molecular level. Next week you will study the enzyme kinetics of acetylcholinesterase.
12.3 Introduction to Molecular Visualization Using the Program Chimera
The visualization program Chimera will be introduced to you by using the structure of the multi-drug resistance (MDR) efflux pump protein. This is the protein that is up-regulated by a riboswitch in response to antibiotics like tetracycline. The protein then pumps the antibiotics out of the bacteria allowing the cells continued growth. Because the MDR efflux pump eliminates many different drugs, the bacteria become multi-drug resistant.
The MDR efflux pump is an integral membrane protein. As such, the protein structure has some common membrane protein structural characteristics. First, transmembrane helices comprise the bulk of the protein. These helices stretch the width of the lipid bilayer and expose hydrophobic amino acids to the lipid bilayer interior. Hydrophilic amino acids are located where the protein meets the lipid bilayer surface, both on the outside of the cell as well as on the inside.
The instructions for using Chimera follow this format:
Italics indicates that you should go to a Menu or a Command line or the Cursor
Bold… tells you which menu to access
…Bold indicates a selection from the accessed menu
Bold indicates a command
- First, download the protein data. Structural data for macromolecules is available from a central data bank, the RCSB Protein Data Bank. Every structure is given its own unique code. For example, the multi-drug resistance efflux pump data set used here is “2GFP.”
Menu: File... Fetch by ID
type in box 2GFP
click Fetch button - The command line can be used for very specific changes. To bring up the command line (or hide it) go to the “Tools” menu, “General Controls” sub-menu.
Command: del :.b
Command: focus - The cursor (mouse) provides for quick changes in the protein view.
Cursor: left mouse button + moving cursor = rotates protein
Cursor: ctrl + left mouse button + moving cursor = selects protein
Cursor: right mouse button + moving cursor = changes size of protein
Cursor: ctrl + right mouse button + moving cursor = translates protein
Place cursor over the protein to identify specific amino acids - The “Presets” menu gives choices for common ways to represent the protein.
Menu: Presets... Interactive 1
Menu: Presets... Interactive 2
Menu: Presets... Interactive 3
Menu: Presets... Interactive 4 - The “Select” menu specifies what part of the protein will be changed by the “Actions” menu. Try the following three examples:
- Example #1: A chain of amino acids linked by peptide bonds (a polypeptide) is selected and any action now will apply to that chain (e.g., changing color).
Menu: Select... Chain... A
Menu: Actions... Color...? - Example #2: All amino acids that carry a negatively charged side chain at neutral pH (i.e., carboxylates) are selected and all the atoms/bonds in the side chains are (a) shown and (b) colored by element.
Menu: Select... Clear Selection
Menu: Select... Residue... amino acid categories... negative
Menu: Actions... Atoms/bonds... show
Menu: Actions... Color... by element - Example #3: All amino acids that carry a positively charged side chain at neutral pH are selected and all the atoms/bonds in the side chains are shown and colored by element. The atoms are then shown as actual size (sphere). Finally, the protein is shown as a solid object with a surface, as it would actually appear.
Menu: Select... Selection Mode (replace)... append
Menu: Select... Residue... amino acid categories... positive
Menu: Actions... Atoms/bonds... show
Menu: Actions... Color... by element
Menu: Actions... Atoms/Bonds... sphere
Menu: Actions... Atoms/Bonds... hide
Menu: Select... Clear Selection
Menu: Select... Chain... A
Menu: Actions... Surface... show
- Example #1: A chain of amino acids linked by peptide bonds (a polypeptide) is selected and any action now will apply to that chain (e.g., changing color).
- The tools menu can provide further information about protein properties.
Menu: Tools... Surface/Binding Analysis... Coulombic Surface
Coloring
click OK button
Menu: Actions...Surface...Hide
Menu: Tools... Depiction... Color Secondary Structure
click OK button
Menu: File... Close Session
12.4 Analysis of Acethylcholinesterase Using the Computer Visualization Program Chimera
The following series of tasks help you learn how to use computer visualization software to better understand how the enzyme acetylcholinesterase works. Follow the instructions below to answer the questions.
1. The overall structure of acetylcholinesterase. Proteins are stabilized by secondary structures, commonly either β-pleated sheets or α-helices.
Menu: File... Fetch by ID
type in box 1AMN
click Fetch button
Menu: Tools... Depiction... Color Secondary Structure
click OK button
Command: del :HOH :SO4
- (2 pts.) How many strands are included in each of the two β-pleated sheets?
- (3 pts.) Identify the two longest α-helices by listing the abbreviations for the amino acids at the beginning and the end of each helix. (Placing your cursor over a spot on the protein will cause the abbreviation to be shown.) How many amino acid residues are in each helix? [Hint: the amino acids are numbered consecutively.]
2. Substrate Analog (NAF) at the Active Site. Active sites are often marked or labeled with a substrate analog. This is a substrate-like molecule that reacts incompletely at the enzyme active site. It remains bound to the enzyme and marks some of the catalytic amino acid side chains. In this case the substrate analog forms a structure like the tetrahedral intermediate shown in the Introduction and Background.
Menu: Select... Residue... NAF
Menu: Actions... Atoms/Bonds... Show
Menu: Select... Invert (all models)
Menu: Actions... Ribbon... Hide
Menu: Actions... Atoms/Bonds... Hide
Command: display: 200
Command: focus
- (4 pts.) Draw the structural formula for NAF connected to an atom of an amino acid side chain (treat the rest of the side chain as an “R” group). This structure contains two ionic charges (one positive and one negative). Can you locate these charges? Mark each with either a “+” or a “-.”
- (4 pts.) NAF forms a covalent intermediate with the enzyme just as the natural substrate, acetylcholine, does. This intermediate is called a “tetrahedral intermediate.” Why do you think it is given that name? Be as specific as possible.
3. Noncovalent Interactions between the Active Site and the Substrate Analog. Van der Waals contacts between the protein and the substrate analog are very common.
Menu: Select... Clear Selection
Menu: Select... Residue... NAF
Menu: Tools... Structure Analysis... Clashes/Contacts
click Designate button
click Contact button
check Select
uncheck Draw pseudobonds of color
check Color
click OK button
a. (3 pts.) How many atoms of NAF are directly in contact with the protein? Out of a total of how many atoms in NAF? [Atoms marked in red are in direct contact with the protein.] Mark each atom that is in contact with an asterisk (*) in your NAF structure in question #3.
4. Visualize the amino acids that immediately surround the substrate analog click on Graphics Window then hit the arrow up key [This will select the complete NAF.]
Menu: Actions... Atoms/Bonds... Show
Menu: Actions... Atoms/Bonds... Sphere
Command: focus
Menu: Select... Clear Selection
Menu: Select... Residue... NAF
Menu: Actions... Color... choose a color that is easy to see
a. (4 pts.) Does the NAF have any “wiggle room” when bound at the active site? How do you think that this might aid enzyme catalysis?
5. Select active site amino acids that help catalyze the reaction.
Menu: Actions... Atoms/Bonds... Stick
Menu: Actions... Atoms/Bonds... Color... by element
Menu: Select...Invert (all models)
Menu: Actions... Atoms/Bonds... Stick
Menu: Actions... Atoms/Bonds... Color... by element
Command: show :118 :119 :200 :327 :440 :NAF
Command: focus
a. (3 pts.) Identify the α-carbon atom for each amino acid. The α-carbon lies between a carboxyl “C=O” and an “N-H” along the protein backbone. The side chain for each amino acid is also connected to the α-carbon. (The side chain for a glycine amino acid is simply a “H” and so it is not shown in the structure.) One side chain is covalently bonded to NAF. Move your cursor to the α-carbon to learn the abbreviated name for this amino acid. Use the list of amino acid structures below to identify this amino acid and draw its side chain structure.
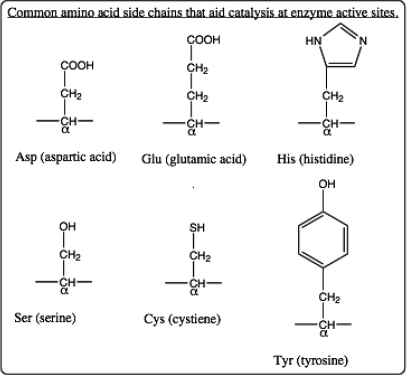
b. (4 pts.) Draw structures for all other amino acid side chains shown at this active site. Identify each amino acid as well. At the catalytic step shown by this structure determined by x-ray crystallography, two of the amino acid side chains carry an ionic charge (one positive and one negative). Identify the charged side chains and give them their appropriate charges.
Menu: Tools...Structural Analysis... FindHbond
click OK button
c. (3 pts.) The blue lines show H-bonds and the H-bonded atoms are close enough together to catalyze reactions.
Two hydrogen bonds that connect to NAF are not from amino acid side chains. Describe or identify the groups from which these H-bonds originate.
Measure the distance between atoms along the blue lines (H-bonds). [Place the cursor over a H-bond. A box will appear detailing the atom within each residue that is involved in H-bonding as well as the distance of the H-bond.] Identify the atoms and give distances.
Ctrl + Left click and drag over all atoms to select
Menu:Actions... Atoms/Bonds... sphere
d. (4 pts.) Note that where H-bonding occurs, the electron clouds are overlapping. What does this mean in terms of bonding?
Menu: Select... Chain... A
Menu: Actions... Ribbons... Show
Menu: Select... Clear Selection
Command: focus
Menu: Actions... Surface... Show
Menu: Select... Residue... NAF
Menu: Actions... Colors... green
a. (4 pts.) Locate NAF. This is a marker for the location of the active site. Describe the active site location relative to the bulk protein. Might this location cause problems for the enzyme? Briefly explain.
7. Protein Surface Charges and Catalysis
Command: del:NAF [This will leave a “hole” in the active site.]
Menu: Tools... Surface/Binding Analysis... Coulombic Surface Coloring
choose 11 for number of colors/values
click OK button
[Close any error messages. This calculation may take a minute.]
a. (3 pts.) Describe the location of the predominately blue region (positively charged region) with respect to the active site. How might this relative location help catalysis?
b. (3 pts.) Describe the location of the predominately red region (negatively charged region) with respect to the active site. How might this relative location help catalysis?
Menu: File... Close Session
Menu: File... Fetch by ID
type in box 2v96
click Fetch button
Menu: Presets... Interactive 1 (ribbons)
Command: del :.b
Menu: File... Fetch by ID
type in box 2va9
click Fetch button
Command: del :.b
Menu: Tools... Structure Comparison... Match Maker
highlight reference structure 2V96
check After superposition, compute structure-based multiple sequence alignment
click Apply button
check Iterate superposition/alignment... in window that opens
check Iterate alignment until convergence
click OK button
Add button
double click on 2V96.pdb (#0) in new window
double click on 2VA9.pdb (#1) in new window
double click on 2V96.pdb (#0) in new window
select Action on Create: hide Conformations
click Create button
Command: focus
MD Movie: Molecular Movement... [This is a new window that opens.]
click Play button
9. Protein Motion at the Active Site
Command: display #2 :200
(Note that this marks the active site using one of the active site amino acids.)
Command: display #2 :70 :74 :84 :121 :279 :290 :330 :331 :334
(Note that these amino acids line the “tunnel” into the active site.)
a. (3 pts.) These motions have been described as “breathing” of the “tunnel.” Why would you expect motions like this in the tunnel that leads to the active site? Briefly explain.
Notes to Instructor
This laboratory allows the students to “try out” the Chimera program - to experiment with different settings and commands. We find that Chimera is relatively user-friendly and forgiving. Students are encouraged to download this free software for their own use. Chimera may be used with at-home assignments based on the lecture portion of a biochemistry course as well. This is a good opportunity to connect with topics covered in lecture as well as the laboratory.
Molecular Visualization of Acethylcholinesterase Prelab
Using any common sources (internet, biochemistry textbook, etc.), answer the following questions concerning the general characteristics of the enzyme, acetylcholinesterase.
1. (3 pts.) What reaction is catalyzed by acetylcholinesterase? Show the structures of reactants and products.
2. (3 pts.) In general terms how is acetylcholinesterase important to neuronal transmission across a synapse?
3. (3 pts.) Where is acetylcholinesterase located within the synaptic space? For example, is acetylcholinesterase found on the presynaptic membrane or on the postsynaptic membrane or as a soluble enzyme in the synaptic cleft or ... ? In general, is this enzyme free to diffuse away from the synapse? Briefly explain.
Acetylcholinesterase Characteristics Worksheet
Using any common sources (internet, biochemistry textbook, etc.), answer the following questions concerning the general characteristics of the enzyme, acetylcholinesterase.
1. (2 pts.) Is the natural form of acetylcholinesterase a single polypeptide chain (a monomer)? Or is acetylcholinesterase found as a polymer? Explain.
2. (2 pts.) Briefly describe the steps acetylcholine takes, starting from being in the presynaptic vesicle and ending when acetylcholine is reacted by acetylcholinesterase.
3. (3 pts.) Many nerve gasses (e.g., sarin (GB), soman (GD)) were designed to impact acetylcholinesterase. Specifically, what do these nerve gasses do to this enzyme? How does the change in acetylcholinesterase (brought about by nerve gasses) affect nerve transmission?
4. (3 pts.) Many common insecticides (e.g., malathion, Sevin) also affect acetylcholinesterase. Also some drugs used to treat diseases such as Alzheimer’s and myasthenia gravis target acetylcholinesterase. Examples of these drugs include physostigmine (eserine), neostigmine, and pyridostigmine. How is the activity of acetylcholinesterase impacted by these insecticides or drugs? How does this change in acetylcholinesterase affect nerve transmission?


