1.7: Synthesizing the ykkCD Mutant Toxin Sensor RNA in vitro
- Page ID
- 18924
7.1 Learning Objective
In the quest toward understanding how the ykkCD toxin sensor recognizes the antibiotic tetracycline you thus far designed mutants to alter the sequence of the sensor, and made the plasmid vectors containing the mutant sensor using PCR amplification. You purified these plasmids from bacterial cells and prepared them to be templates for RNA synthesis. In this lab you will learn how RNA polymerases work. You will synthesize the mutant sensor in vitro using the plasmid DNA template and T7 RNA polymerase.
7.2 Mini Project Flowchart
The bolded block in the flowchart below highlights the role of the current experiment in the mini project.

7.3 How do RNA Polymerases Work?
RNA polymerases are enzymes that synthesize RNA using a DNA template. RNA synthesis using RNA polymerases is also called transcription. During transcription we copy sequence information coded in the genome (DNA) to RNA. When the sequence codes for protein the resulting RNA sequence is used as template for protein synthesis. The chemistry of RNA polymerization is identical to that of DNA synthesis (Chapter 4). The 3'-OH group of the growing polynucleotide chain acts as a nucleophile to attack the phosphorous of the incoming nucleotide triphosphate (NTP) thereby forming the phosphodiester bond. Divalent metal ions (Mg2+) are required for transcription to position the 3'-OH group for nucleophilic attack (Fig. 7.1).
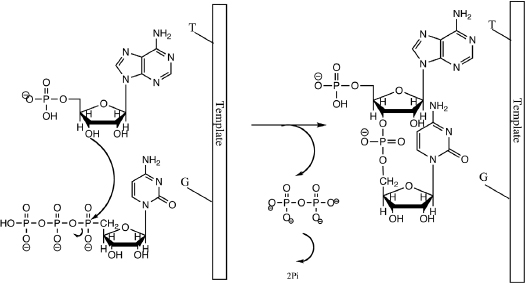
During polymerization, a high-energy bond (the anhydride bond in the incoming NTP) is broken and a high-energy bond (the phosphodiester bond) is formed. What makes polymerization energetically favorable? The breaking of the pyrophosphate (PPi) into two inorganic phosphates is what drives polymerization. Without this step, polymerization would not be energetically favorable. During transcription the polymerase first unwinds the DNA template, then recruits the nucleotide triphosphate (NTP) that is complementary to the template and finally it catalyzes formation of the phosphodiester bond. The polymerase moves from 3' to 5' direction on the template DNA while synthesizing RNA. The schematic depiction of this process is also called the transcription bubble (Fig. 7.2).
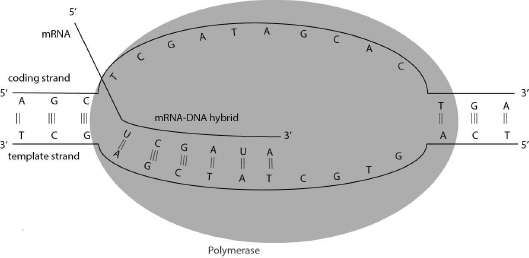
7.4 How Does Transcription Start?
Polymerases are recruited to the DNA template by specific sequences called promoters (Fig 7.3). Each polymerase has its own promoter sequence. Once the polymerase encounters this sequence it binds to the DNA template and initiates RNA synthesis.

Polymerases are oligomer enzymes, meaning each polymerase has a catalytic domain that facilitates the chemistry of polymerization. They also have a domain that recognizes the promoter and helps recruit the polymerase to the DNA template. In E. coli this domain is called the sigma factor (Fig. 7.4).

7.5 How Does Transcription End?
In bacteria transcription ends when the polymerase encounters a stable structure that it cannot unwind. This stable structure is also called the terminator or terminator stem. The polymerase stalls when encountering a stable structure, and together with the nascent RNA, it is released from the DNA template thereby ending transcription (Fig. 7.5).
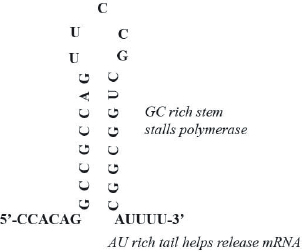
In research laboratories, scientists most commonly use T7 RNA polymerase for in vitro transcription, because it can synthesize milligram amounts of RNA within two hours. The T7 RNA polymerase is from the T7 bacteriophage. The promoter of T7 RNA polymerase is 19-nucleotide long and folds into a hairpin. This sequence has to be present in the cloning vector upstream of the DNA template sequence to enable successful transcription. Since the T7 RNA polymerase is very robust, most terminator stems are not stable enough to stall it. Therefore, to terminate transcription at the end of the desired sequence, the template DNA must be cut (linearized) prior to RNA synthesis to stop the T7 RNA polymerase. Meaning, the polymerase falls off the DNA template at the cut site thereby ending transcription. In case of the ykkCD sensor RNA, the cleavage site for the BamHI restriction endonuclease directly follows the ykkCD sequence in the cloning vector. This site recruits the BamHI restriction enzyme that cleaves (linearizes) the template DNA prior to transcription (Chapter 6.)
7.6 Transcription in Practice
Several kits are commercially available to perform in vitro transcription with high yield. These kits contain T7 RNA polymerase (enzyme), NTPs and a reaction buffer. The reaction buffer has pH~8.0 and contains magnesium ions (usually in the form of MgCl2). Transcription is performed in a 37 °C water bath for 1 or 2 hrs. Inclusion of a ribonuclease (RNase) inhibitor often increases the yield and quality of in vitro synthesized RNA, because it inhibits ribonucleases that might degrade the synthesized RNA.
7.7 What Are We Going To Do Today?
Today we will precipitate the DNA template, check DNA concentration and transcribe the mutant toxin sensor RNA in vitro.
PROTOCOL
Reagents and equipment needs are calculated per six student teams (appropriate excess included).
Equipment/glassware needed
- Three sets of micropipettes 20-100 μl and 2-20 μl.
- UV spectrometer.
- Water bath set to 37 ºC.
- Microcentrifuge.
Solutions needed
- 1 ml ice-cold 75% ethanol.
- 300 μl TE solution (10 mM Tris, 1 mM EDTA pH=8.0); RNase free.
- Kit or polymerase and buffer for in vitro RNA synthesis.
Precipitate plasmid DNA
- Spin plasmid DNA for 30 min at 4 °C at maximum speed. Carefully remove supernatant. This step pellets the DNA.
- Add 50 μl ice-cold 75% ethanol.
- Centrifuge plasmid DNA at 4 °C at maximum speed for 5 minutes.
- Remove supernatant. This step removes residual salt.
- Dry DNA pellet using a speed-vac (about 5 minutes) or on the bench.
- Resuspend pellet in 20 μl TE buffer.
UV spectroscopy (this protocol is for Nanodrop spectrometer; for conventional UV spectrometer larger volumes of DNA need to be used)
- Turn on spectrophotometer and choose nucleic acid assay setup.
- Blank spectrophotometer with 2 μl millipore water.
- Measure the absorbance of a water sample to insure the spectrometer is clean.
- Use 2 μl plasmid DNA sample to measure the concentration of your plasmid DNA.
- Record DNA concentration in g/l.
In vitro transcription
Protocol depends on kit used and will be provided in lab. See “transcription in practice” for a basic overview.
Note to the instructor
The experiment in Chapter 7 is designed to synthesize the ykkCD toxin sensor RNA in vitro using T7 RNA polymerase. The same protocol with minimal modifications can be used to synthesize any RNA sequence. The Ribomax T7 large scale RNA production system was purchased from Promega, but other vendors may be used. RNA production kits are preferable over using home-made T7 RNA polymerase, because they produce more RNA within a shorter period of time. To reduce cost, home-made T7 RNA polymerase may be used in place of an RNA production kit. Since RNA synthesis using a home-made T7 RNA polymerase takes about 1.5 hrs., we recommend starting the laboratory with DNA precipitation and giving pre-laboratory lecture while the RNA is synthesized. Alternatively, the long break may be used for an exam. Even if an RNA production kit is used this laboratory session has two breaks lasting 30 min each. Those breaks may be utilized to complete the restriction endonuclease worksheet.
Prelab Questions for RNA Transcription
Define the following terms.
- RNA polymerase

- Promoter

- Transcription

- Transcription terminator

5. Calculate how to mix a transcription reaction if the final reaction should have (in 20 ml).
2 ml T7 RNA polymerase
1 x transcription bufffer
1 mg linearized DNA

Assume that the transcription buffer stock is 2 x and the stock DNA concentration is 156 ng/ml. Complete volume (to 20ml) using water (if needed). Show your work.
RNA Synthesis
Lab Report Outline and Point Distribution
1. Several sentences defining the goal/purpose of this experiment. Make sure you indicate the role this step plays in the mini project (5 pts.).
2. Report the concentration of the linearized DNA. Calculate linearized DNA yield. Did you get a good yield in your assessment? Explain (8 pts.).
3. Based on the 260/280 absorbance ratio comment on the purity of the linearized DNA (5 pts.).
4. Your colleague had a satellite peak in the absorbance spectrum of his linearized DNA. Explain what might be the source of this contamination and how does this contamination bias his determination of DNA concentration (5 pts.). (Would he overestimate or underestimate the concentration due to the peak?)
5. Restriction endonuclease worksheet (27 pts.).


