Homework Problems - Literature Learning Module: CRISPR/Cas 9 KEY
- Page ID
- 9993
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)esearch Paper: Cut Site Selection by the Two Nuclease Domains of the Cas9 RNA-guided Endonuclease. Hongfan Chen, Jihoon Choi and Scott Bailey.The Journal of Biological Chemistry, 289, 13284-13294 (2014)
Background and Review
A promising tool for genome manipulation and regulation in a wide variety of organisms has recently been identified in the RNA-guided DNA endonuclease activity of the CRISPR-Cas (clustered regularly interspaced short palindromic repeat–CRISPR-associated) system. CRISPR-Cas, an inheritable prokaryotic immune system, protects bacteria and archaea against mobile genetic elements via RNA-guided target silencing. CRISPR-Cas systems consist of an array of short direct repeats interspersed by variable invader-derived sequences (spacers) and a cas operon. During invasion, small fragments of the invading DNA from phage or plasmids (protospacers) are incorporated into host CRISPR loci, transcribed, and processed to generate small CRISPR RNAs (crRNA). The invading nucleic acid is then recognized and silenced by Cas proteins guided by the crRNAs. There are three types of CRISPR-Cas system each characterized by the presence of a signature gene.
Programmed DNA cleavage requires the fewest components in the type II CRISPR-Cas system, requiring only crRNA, a trans-activating crRNA (tracrRNA), and the Cas9 endonuclease, the signature gene of the type II system. The system can be further simplified by fusing the mature crRNA and tracrRNA into a single guide RNA (sgRNA). In addition to its role in target cleavage, tracrRNA also mediates crRNA maturation by forming RNA hybrids with primary crRNA transcripts, leading to co-processing of both RNAs by endogenous RNase III. Cas9 contains two nuclease domains that together generate a double-strand (ds) break in target DNA. The HNH nuclease domain cleaves the complementary strand, and the RuvC-like nuclease domain cleaves the noncomplementary strand.
A short signature sequence, named the protospacer adjacent motif (PAM), is characteristic of the invading DNA targeted by the type I and type II CRISPR-Cas systems. The PAM serves two functions. It has been linked to the acquisition of new spacer sequences, and it is necessary for the subsequent recognition and silencing of target DNA. The sequence, length, and position of the PAM vary depending on the CRISPR-Cas type and organism. PAMs from type II systems are located downstream of the protospacer and contain 2–5 bp of conserved sequence. A variable sequence, of up to 4 bp, separates the conserved sequence of the PAM from the protospacer. This variable region is often included in the definition of the PAM sequence, but for simplicity, we refer to this variable region as the linker and the conserved sequence as the PAM. To date, Cas9 from Streptococcus pyogenes, Cas9 from Streptococcus thermophilus DGCC7710, and Cas9 from Neisseria meningitidis have been employed as tools for genome editing or regulation. For these Cas9 orthologs, the PAMs are GG, GGNG, and GATT, and the linkers are 1, 1, and 4 bp, respectively.
The simplicity of sgRNA design and sequence-specific targeting means the RNA-guided Cas9 machinery has great potential for programmable genome engineering. Cas9 can be employed to generate mutations in cells by introducing dsDNA breaks. The capabilities of Cas9 can be expanded to various genome engineering purposes, such as transcription repression or activation, with its nickase (generated by inactivating one of its two nuclease domains) or nuclease null variants. Another appealing possibility for the Cas9 system is to target different Cas9-mediated activities to multiple target sites, for example transcriptional repression of one gene but activation of another. To achieve this, multiple Cas9 orthologs will need to be employed as a single ortholog cannot concurrently mediate different activities at multiple sites. Therefore to broaden our understanding of Cas9 proteins, we have characterized the Cas9 ortholog from S. thermophilus LMG18311, which we refer to as LMG18311 Cas9.
Questions:
1. Investigators studied the type II CRISPR-Cas system of S. thermophilus LMG18311.
a. Figure A shows a Coomassie Blue-stained SDS-polyacrylamide gel of Cas9, Cas9 D9A, Cas9 D599A, and Cas9 D9A,D599A where D is aspartic acid and A is alanine and D#A reflects the site of the specific mutation where D was changed to A. Why did they perform this experiment?
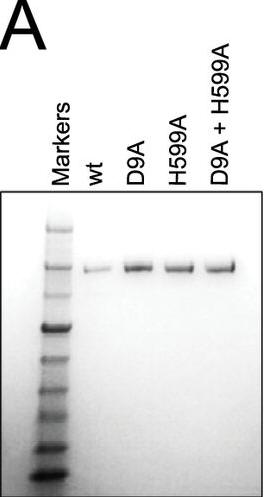
Answer: to verify that the mutants had the same molecular weight and behaved similarly on denaturing gel electrophoresis.
b. To confirm the PAM sequence for LMG18311 Cas9, the authors performed BLAST searches to identify potential protospacers in viral and plasmid genomes that matched any of the 33 spacer sequences from CRISPR-1. This search generated 41 unique target sequences, from the genomes of bacteriophage known to infect S. thermophilus. We then aligned 50-nucleotide segments from the identified target genomes, inclusive of the 30-nucleotide protospacer and 10-nucleotide flanking regions. Figure b shows B, logo plot revealing the PAM for LMG18311 Cas9. The positions of the protospacer, PAM, and linker are indicated.
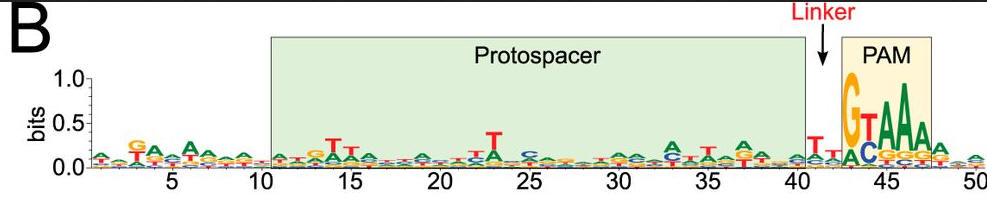
What is the most likely consensus sequence of the PAM sequence? How far is it away from the protospacer:
Answer: GYAAA, invariantly located 2 bp away.
c. Figure C shows a schematic representation of the CRISPR nucleic acid complexes. Label the target DNA, crRNA, tracrRNA,, protospacer, PAM, and linker. Draw in a connecting piece of RNA if you wish to create a single guide RNA containing both the crRNA and tracRNA. What is the specific PAM sequence in this figure.
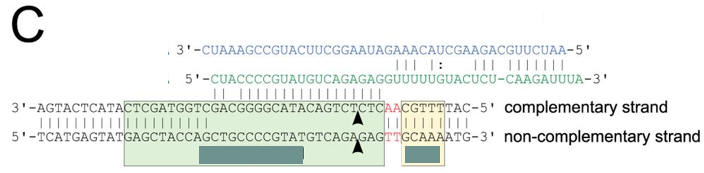
Answer
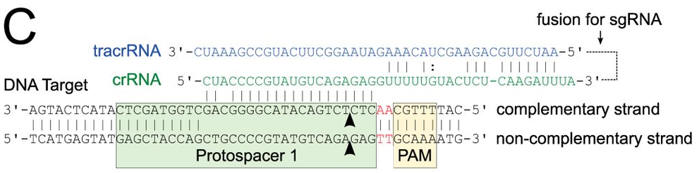
The most commonly observed PAM sequence, found in 7 of the 41 target sequences, was GCAAA.
2. To confirm that the identified PAM was functional, investigators needed a selection system. They used a transformation assay in which E. coli cells containing an exogenous type II CRISPR-Cas system were resistant to plasmid transformation, whereas cells lacking the system are competent for transformation. Explain how this system could be used to determine if specific PAM are functional?
Answer: The E. Coli cells with an added CRISRP-Cas system should be able to cleave any plasmid DNA taken up by the cell as the plasmid DNA would serve as the "target" much like phage DNA. Cells without the exogenous CRISRP-Cas system could not cleave plasmid that were taken up by the cell, which could allow the cells with intact plasmid to be transformed by specific genes brought in and expressed by the plasmid.
a. Figure A shows a schematic representation of transformation assay.
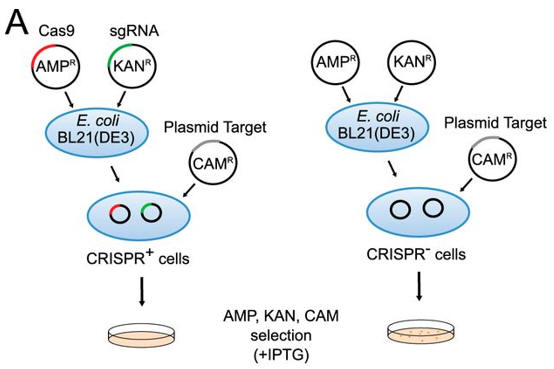
The target plasmid contained protospacer-1 (whose sequence was identical to the first spacer of CRISPR-1), a 2-bp linker, and the identified PAM. The first control plasmid contained only protospacer-1, whereas the second control plasmid lacked both protospacer-1 and PAM. The target and control plasmids were then used to transformed cells. in the presence of IPTG and the appropriate antibiotics.
Why did the investigators add IPTG? What difference would you expect to find in the cells in the left panel compared to the right in their assay?
Answer: Protein expression of the Cas9 protein is necessary for a function CRISPR system. Presumably, the protein is under the control of a lac promoter which is turned out with lactose, allolactose, or IPTG. The cells on the left, which have a functional Cas9 protein and sgRNA will be able to cleave the plasmid that would provoke cell transformation and hence they would not be transformed.
b. Figure 2B shows plasmid transformation by LMG18311 Cas9 and sgRNA in E. coli cells. Transformation efficiency is expressed as cfu per 5 ng of plasmid DNA. Average values from at least three biological replicates are shown, with error bars representing 1 S.D. Interpret and explain results.
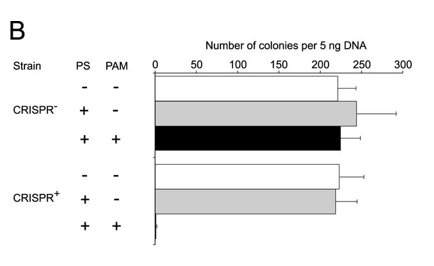
Answer: The control plasmids transformed into both strains with similar efficiency (Figure 2B). The target plasmid failed to transform into the CRISPR+ cells but transformed into the CRISPR−. this shows that a function PAM seqeunce is required for the Cas9 protein to cleave the plasmid and block transformation.
c. For Figure C, the investigators made single nucleotide changes in the PAM sequence of the CRISPR+ plasmid and studied the effect on plasmid transformation efficiency. Interpret he results.
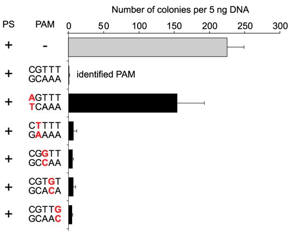
Answer: . Only the plasmid containing a mutation at the position 1 guanosine (that is, the PAM nucleotide closest to the protospacer) was transformed, albeit with a reduced (abouy 66%) transformation efficiency as compared with the intact PAM sequence . Plasmids containing single mutations to any of the other four positions were resistant to transformation. These results indicate that the guanosine at position 1 is important for PAM function but individually the four other positions have little effect on PAM function.
d. For Figure D, the effect of the length of the linker length on plasmid transformation efficiency was studied. PS denotes the protospacer sequence. Interpret the results.
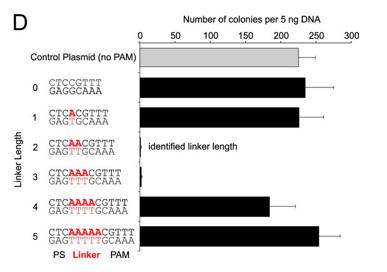
The CRISPR+ cells were equally resistant to transformation by a plasmid target with a linker length of either 2 bp or 3 bp. Plasmids with other linker lengths transformed with efficiencies more similar to the control plasmid, suggesting that plasmids with these linkers were able to escape CRISPR-Cas silencing.
3. The investigators then studied their system in vitro by reconstituting the components. They monitored DNA cleavage by LMG18311 Cas9 as a measure of activity.
a. Figure 3A shows an electrophoresis gel of reaction mixtures containing 5 nm target plasmid, 25 nm Cas9, 25 nm crRNA (42-nucleotide crRNA containing the sequence derived from first spacer of CRISPR-1), 25 nm (42-nucleotide) tracrRNA, and 10 mm Mg2+ that were incubated for 30 min at 37oC. The agarose gels were stained with ethidium bromide. Interpret the results. Why does the linear control migrate less far than the uncut plasmid?
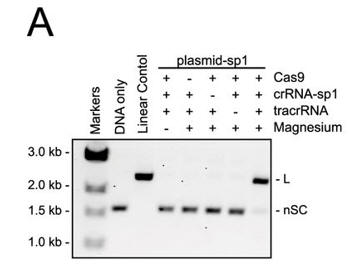
Answer: Cleavage of the plasmid target occurred in the presence of Cas9, tracrRNA, crRNA, and Mg2+. The cut plasmid has a larger hydrodynamic radius than the supercoiled circular plasmid.
b. In Figure B, a cognate sgRNA was substituted for separate crRNA and tracrRNA molecules. Interpret the gel results..
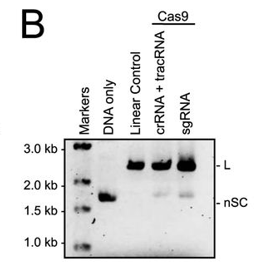
Answer: Cleavage also occurred when an sgRNA was substituted for the tracrRNA and crRNA
c. In Figure C, they studied cleavage in the presence of different sgRNA sequences. Explain the results.
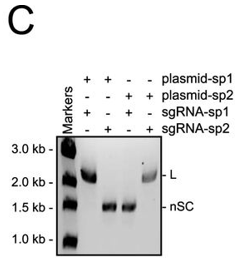
Answer: the cleavage of a plasmid target is dictated by the sgRNA sequence.
d. For figure D, the used different active site mutants of Cas 9. The (D9A) mutant was in the RuvC-like domain while the H599A mutant was in the HNH domain Explain the results. What effect does the double mutant have on the activity?
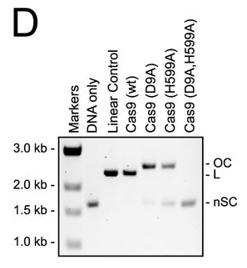
Answer: Cas9 variants with active site mutations in either the RuvC-like domain (D9A) or the HNH domain (H599A) nicked the plasmid targets, whereas a variant with a double mutation (D9A,H599A) displayed no activity
e. In Figure E, the authors studied which domain was involved in the cleavage of each strand (complementary to the guide DNA or the noncomplementary strand) of the target DNA. The dsDNA was radiolabeled at the 5′ end of the complementary strand (left hand gel) or the noncomplementary strand (right hand gel). Reactions were performed as in A, and products were separated by 10% denaturing PAGE. The cleavage sites are indicated with arrows in the schematic diagram (bottom). 50 nt, 50 nucleotides; 37 nt, 37 nucleotides. Interpret the results.
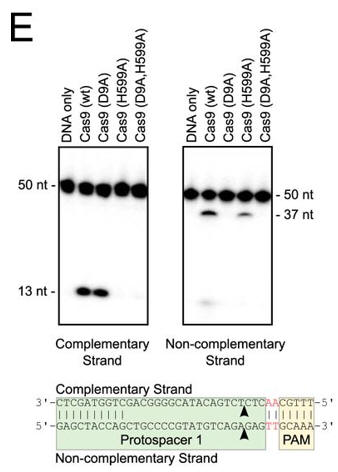
Answer: Cleavage assays using short oligonucleotide substrates confirmed that the HNH domain (H599A) cleaves the strand complementary to the guide RNA, whereas the RuvC-like domain (D9A) cleaves the noncomplementary strand. Mapping the location of the cut sites revealed that, as seen with other Cas9 orthologs, cleavage of both strands occurs within the protospacer, 3 bp from its PAM proximal end, producing a blunt-end dsDNA break.
f. Do mutations in the PAM or changes in linker length have the same effect on DNA interference in vitro as they did in vivo? The investigators monitored cleavage of these variant plasmids by recombinant LMG18311 Cas9. Interpret the results in Figure F.
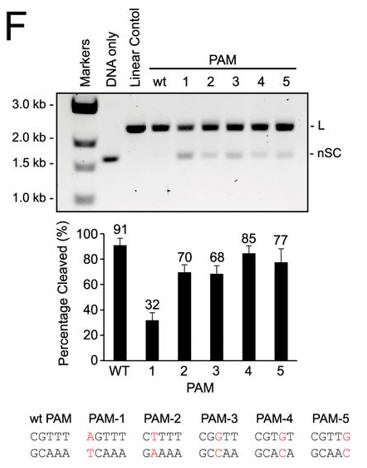
Answer: Mutation of the guanosine at position 1 had the greatest effect, and individual mutations to the other four positions of the PAM had only a modest effect on plasmid cleavage. Cleavage of plasmid targets with different linker lengths was optimal at 2 or 3 bp and then decreased steadily with increasing or decreasing lengths
g. In Figure G, cleavage of plasmid targets containing the indicated linker lengths was studied. In A–C and E–F, the positions of negatively supercoiled (nSC), linear (L) and nicked or open circle (OC) plasmid are indicated. The linear control is a digestion of the plasmid target with the restriction enzyme AgeI
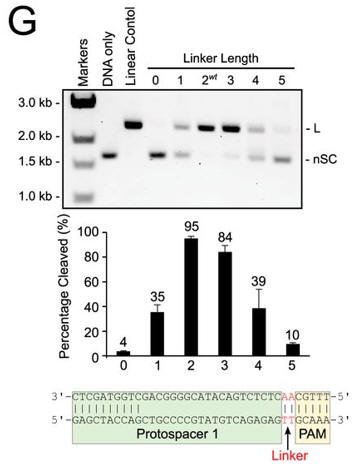
Answer: Cleavage of plasmid targets with different linker lengths was optimal at 2 or 3 bp and then decreased steadily with increasing or decreasing lengths.
h. In the mutational analysis, they substituted asparatic acid for alanine. Draw the structures of the side chains of D and A. Why did they choose A to substitute for D? What role might D have in catalysis in the wild-type protein?
Answer:
The side chain of alanine is not bigger than aspartic acid nor is it large and hydrophobic which could potential interfer with packing of the side chains and hence protein folding which might affect enzyme activity in a general way. In the wild type protein, aspartic acid might be involved in substrate binding through H bond interactions. More than likely is acts as a general acid/base in the actual catalytic mechanism of cleavage of the target strand.
4. To evaluate whether other divalent cations besides Mg2+ can activate DNA cleavage by Cas9, the authors performed plasmid cleavage assays in the presence of one of the following divalent cations: Ca2+, Mn2+, Co2+, Ni2+, and Cu2+.
a. Figure A shows cleavage of a target plasmid by Cas9 with either no metal or 1 mm of the indicated metal ions. All reactions were treated with 0.5 mm EDTA prior to metal addition. Interpret the results.
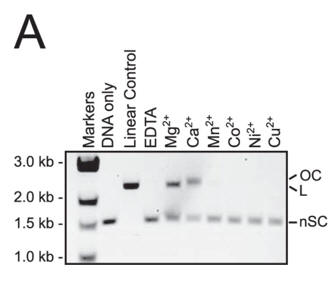
Answer: Reactions containing Ca2+ yielded nicked, instead of linear plasmid, suggesting that Ca2+ activates only one of the Cas9 nuclease domains
b. To identify which domain was activated, we assayed the single active site mutants of Cas9 (D9A or H599A) in a reaction buffer containing Ca2+. In both panels, the positions of negatively supercoiled (nSC), linear (L), and nicked or open circle (OC) plasmid are indicated. The linear control is a digestion of the plasmid target with the restriction enzyme AgeI.
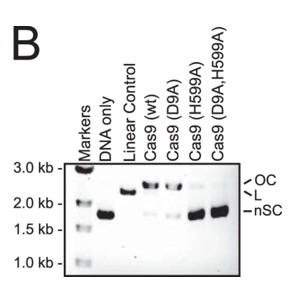
Answer: Little cleavage was with the HNH mutant (H599A) but robust cleavage with the RuvC-like mutant (D9A) , suggesting that the HNH but not the RuvC-like domain was activated by Ca2+ .
5. To determine the effect of PAM sequence and linker length on binding of LMG18311 Cas9 to DNA targets, the authors determined the binding affinity (Kd) of the Cas9-sgRNA complex to 5′ end-labeled dsDNA targets using native gel electrophoresis.
a. Why did they not include SDS in the gel?
Answer: SDS is a powerful denaturant and would have denatured Cas9 and separated the complex.
b. Binding experiments were conducted with the nuclease-deficient mutant of Cas9 (D9A,H599A) in the presence of Mg2+. Fixed concentrations of the dsDNA targets were incubated with increasing concentrations of the Cas9-sgRNA complex. Why did they use the double mutant of Cas9?
Answer: to prevent the enzyme from cleaving the target DNA which would prevent detection of the complex.
c. Figure A below shows a representative gel shift assay for Cas9-sgRNA and the binding curve measured from the assay. From inspection of the graph, what is the approximate Kd for the interaction? The top wedge indicates increasing concentration of the Cas9-sgRNA.
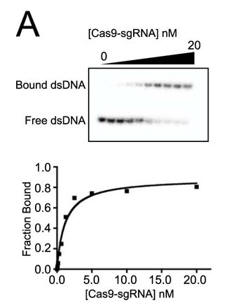
Answer: approximately 2 nM Cas9-sgRNA (the ligand concentration at half maximal binding of the curve is truly a hyperbola.
b. A bar graph plotting Kd values for DNA targets with PAM mutations (labeled red) are shown below (B) . Average values from at least three replicates are shown, with error bars representing 1 S.D. Targets where binding was not observed are shown with Kd values at the lower limit (> 1000 nm). PS denotes the protospacer sequence. Interpret the results.
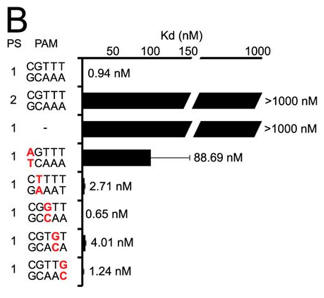
Answer: A target containing a complementary protospacer, a 2-bp linker, and a functional PAM bound to Cas9-sgRNA with an affinity of 0.94 � 0.27 nm. The investigators were unable to detect binding to a target containing a noncomplementary protospacer or to a target that lacked a PAM. Mutation of the guanosine at position 1 of the PAM resulted in an about 100-fold increase in Kd, whereas mutations at positions 2 through 5 did not significantly alter the affinity (all within about 4-fold on the consensus PAM).
c. Next they investigated linker length and binding. Figure C shows a bar graph plotting Kd values for DNA targets with different linker lengths (labeled red) (C). Average values from at least three replicates are shown, with error bars representing 1 S.D. Targets where binding was not observed are shown with Kd values at the lower limit (> 1000 nm). PS denotes the protospacer sequence. Interpret the results.
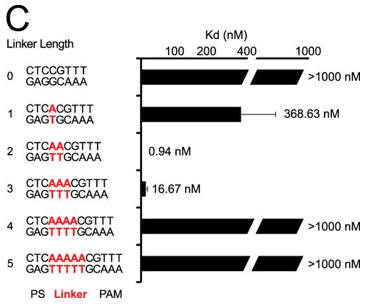
Answer: Changes in linker length had a larger effect on binding affinity. Under the conditions tested, we failed to detect binding to plasmid targets containing linker lengths of 0, 4, or 5 bp (Kd > 1000 nm), whereas linkers of 1 and 3 bp reduced the affinity by about 400- and about 20-fold, respectively.


