12.1: Interacting Phenotypes
- Page ID
- 165830
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Learning Objectives
- Describe the results of crosses involving two genes that affect the same trait as exhibiting complementation, epistasis, or redundancy.
- Predict phenotypes of offspring of crosses involving two genes, given information about biohemical pathways or functions of the genes.
The Big Picture: Studying Biochemical Pathways Using Genetics
One of the most interesting applications of genetics is to the analysis of biochemical pathways. It is possible to learn an enormous amount about interactions at the biochemical level without ever doing any biochemistry! By cleverly designing and interpreting crosses, it is possible to figure out how molecules interact.
A Very Important "rule": "Doing something is dominant to not doing something"
"Rule" is in quotes because, in biology, there are always exceptions. Nonetheless, this "rule" goes a very long way to understanding how to use genetics to dissect biochemical interactions.
This "rule" works most of the time because most of the genes we study encode proteins and many of these proteins are enzymes. Since enzymes are very powerful catalysts, you only need a little to get the job done (most of the time). Consider the following example:
- The A allele encodes fully functional enzyme
- The a allele encodes a non-functional enzyme
Thus:
- AA individuals have 2 copies of the gene encoding fully-functional enzyme. With 2 genes making functional enzyme, this is 100% of the wild-type enzyme level. This is more than enough to carry out whatever reaction this enzyme is required for.
- aa individuals have 2 copies of the gene, neither of which makes any functional enzyme. Thus, their enzyme level is 0%. The reaction the enzyme catalyzes is not carried out - typically resulting in some abnormality.
- So what happens in Aa individuals? They have one copy making functional enzyme and one copy making non-functional enzyme. Thus, their enzyme level is 50%. You might think that this would mean they'd have a mild form of the abnormality (this would be incomplete dominance) but most of the time, 50% enzyme level is more than enough and these individuals are completely normal. Since the phenotype of Aa is the same as that of AA, that means the phenotype of the A allele is simply dominant to that of the a allele.
Especially when we're dealing with biochemical pathways, in general:
- The allele with the dominant phenotype encodes a functional protein.
- The allele with the recessive phenotype encodes a non-functional protein.
Graded Problem \(\PageIndex{1}\)
Some dihybrid crosses produce a phenotypic ratio that differs from 9:3:3:1, such as 9:3:4, 12:3:1, 9:7, or 15:1. Note that each of these modified ratios can be obtained by summing one or more of the 9:3:3:1 classes expected from our original dihybrid cross. In the following sections, we will look at some modified phenotypic ratios obtained from dihybrid crosses and what they might tell us about interactions between genes.
Complementary gene action
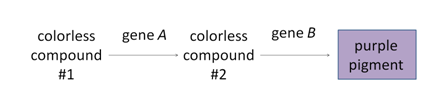
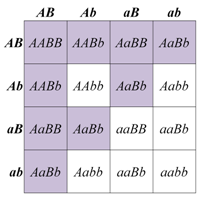
As we have already discussed, in the absence of any phenotypic interaction, there are four phenotypic classes among the progeny of a dihybrid cross. The four phenotypic classes correspond to the genotypes: A_B_, A_bb, aaB_, and aabb. In some cases, the progeny of a dihybrid cross may produce just two phenotypic classes, in an approximately 9:7 ratio. An interpretation of this ratio is that the loss of function of either A or B gene function has the same phenotype as the loss of function of both genes, due to complementary gene action or "complementation" for short (meaning that the functions of both genes work together to produce a final product and the intermediate product is not detectable). For example, consider a simple biochemical pathway in which a colorless substrate is converted by the action of gene A to another colorless product, which is then converted by the action of gene B to a visible pigment. Loss of function of either A or B, or both, will have the same result: no pigment production. Thus A_bb, aaB_, and aabb will all be colorless, while only A_B_ genotypes will produce pigmented product. The modified 9:7 ratio may therefore be obtained when two genes act together in the same biochemical pathway, and when their loss of function phenotypes are indistinguishable from each other or from the loss of both genes.
Graded Problem \(\PageIndex{2}\)
Epistasis
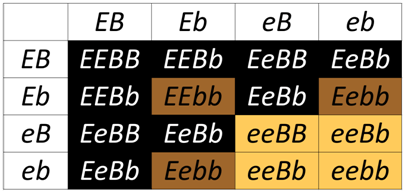
Epistasis (which means “standing upon”) occurs when the phenotype of one locus masks, or prevents, the phenotype of another locus. Thus, following a dihybrid cross fewer than the typical four phenotypic classes will be observed with epistasis. As we have already discussed, in the absence of epistasis, there are four phenotypic classes among the progeny of a dihybrid cross. The four phenotypic classes correspond to the genotypes: A_B_, A_bb, aaB_, and aabb. If either of the singly homozygous recessive genotypes (i.e. A_bb or aaB_) has the same phenotype as the double homozygous recessive (aabb), then a 9:3:4 phenotypic ratio will be obtained. For example, in the Labrador Retriever breed of dogs, the B locus encodes a gene for an important step in the production of melanin. The dominant allele, B is more efficient at pigment production than the recessive b allele, thus B_ hair appears black, and bb hair appears brown. A second locus, which we will call E, controls the deposition of melanin in the hairs. At least one functional E allele is required to deposit any pigment, whether it is black or brown. Thus, all retrievers that are ee fail to deposit any melanin (and so appear pale yellow), regardless of the genotype at the B locus.

The ee genotype is therefore said to be epistatic to both the B and b alleles, since the homozygous ee phenotype masks the phenotype of the B locus. The B/b locus is said to be hypostatic to the ee genotype. Because the masking allele is in this case is recessive, this is called recessive epistasis.
Biochemical Basis for the genetic interaction between E and B in dog coat color
Why is this genetic interaction observed? The B locus encodes the protein TYRP1 (Tyrosinase-Related Protein 1). The enzyme catalyzes a step in the production of melanin within melanocytes. Predicted loss-of-function alleles of TYRP1 were identified in brown dogs, for example a premature stop codon (Schmutz et al. TYRP1 and MC1R genotypes and their effects on coat color in dogs. Mamm Genome. 2002;13(7):380-387). Dogs homozygous or heterozygous for functional TYRP1 produce sufficient melanin to have a black coat.
The E locus encodes the protein MC1R (melanocortin receptor 1), which is a membrane bound receptor for melanocyte-stimulating hormone (MSH) on the surface of the melanocytes. Activation of this receptor is transduced into a transcriptional response to produce proteins needed for eumelanin production. Recessive loss-of-function alleles of MC1R result in reduced or absent receptor activity and therefore loss of the intracellular response of proteins proteins needed for eumelanin production. When MC1R is not activated, cells can still produce the yellow pheomelanin, resulting in a yellow coat.
In the Punnett square above, any genotype that includes a homozygous recessive for MC1R (ee) results in a yellow coat, even when the dog has a dominant allele B that is predicted to produce black fur. Without a functional MC1R receptor, the downstream events needed to produce TYRP1 will not be activated, so the ee genotype "masks" the production of brown of black fur.
Graded Problem \(\PageIndex{3}\)
Redundant genes
When a dihybrid cross produces progeny in two phenotypic classes in a 15:1 ratio, this can be because the two loci’s gene products have the same (redundant) functions within the same biological pathway. Yet another pigmentation pathway, in this case in wheat, provides an example of gene redundancy, sometimes known as duplicate gene action. The biosynthesis of red pigment near the surface of wheat seeds involves many genes, two of which we will label A and B. Normal, red coloration of the wheat seeds is maintained if function of either of these genes is lost in a homozygous mutant (e.g. in either aaB_ or A_bb). Only the doubly recessive mutant (aabb), which lacks function of both genes, shows a phenotype that differs from that produced by any of the other genotypes. A reasonable interpretation of this result is that both genes encode the same biological function, and either one alone is sufficient for the normal activity of that pathway.


Graded Problem \(\PageIndex{4}\)
jsVGL problems for this section:
Click on the links below to access jsVGL problems. Each link will take you to a problem that will let you practice with different genetic models.
- Problems involving Complementation. Starting a problem with this link gives a 50/50 chance of one of these two models:
- Complementation with 2 unlinked autosomal genes; 2 alleles each; simple dominance
- Phenotype 1 ==(Gene A)==> Phenotype 1 ==(Gene B)==> Phenotype 2
- One autosomal gene; 2 alleles; simple dominance
- Complementation with 2 unlinked autosomal genes; 2 alleles each; simple dominance
- Problems involving Epistasis. Starting a problem with this link gives a 50/50 chance of one of these two models:
- Epistasis with 2 unlinked autosomal genes; 2 alleles each; simple dominance
- Phenotype 1 ==(Gene A)==> Phenotype 2 ==(Gene B)==> Phenotype 3
- One autosomal gene; 2 alleles; incomplete dominance
- Epistasis with 2 unlinked autosomal genes; 2 alleles each; simple dominance
Contributors and Attributions
Dr. Todd Nickle and Isabelle Barrette-Ng (Mount Royal University) The content on this page is licensed under CC SA 3.0 licensing guidelines.


