SS1_2019_Lecture_11
- Page ID
- 23939
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
Replication design challenge: proofreading
When the cell begins the task of replicating the DNA, it does so in response to environmental signals that tell the cell it is time to divide. The ideal goal of DNA replication is to produce two identical copies of the double-stranded DNA template and to do it in an amount of time that does not pose an unduly high evolutionarily selective cost. This is a daunting task when you consider that there are ~6,500,000,000 base pairs in the human genome and ~4,500,000 base pairs in the genome of a typical E. coli strain and that Nature has determined that the cells must replicate within 24 hours and 20 minutes, respectively. In either case, many individual biochemical reactions need to take place.
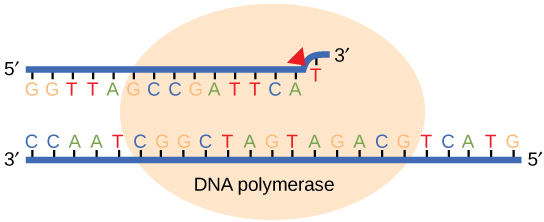
While ideally replication would happen with perfect fidelity, DNA replication, like all other biochemical processes, is imperfect—bases may be left out, extra bases may be added, or bases may be added that do not properly base-pair. In many organisms, many of the mistakes that occur during DNA replication are promptly corrected by DNA polymerase itself via a mechanism known as proofreading. In proofreading, the DNA polymerase "reads" each newly added base via sensing the presence or absence of small structural anomalies before adding the next base to the growing strand. In doing so, a correction can be made.
If the polymerase detects that a newly added base has paired correctly with the base in the template strand, the next nucleotide is added. If, however, a wrong nucleotide is added to the growing polymer, the misshaped double helix will cause the DNA polymerase to stall, and the newly made strand will be ejected from the polymerizing site on the polymerase and will enter into an exonuclease site. In this site, DNA polymerase is able to cleave off the last several nucleotides that were added to the polymer. Once the incorrect nucleotides have been removed, new ones will be added again. This proofreading capability comes with some trade-offs: using an error-correcting/more accurate polymerase requires time (the trade-off is speed of replication) and energy (always an important cost to consider). The slower you go, the more accurate you can be. Going too slow, however, may keep you from replicating as fast as your competition, so figuring out the balance is key.
Errors that are not corrected by proofreading become what are known as mutations.
Suggested discussion
Why would DNA replication need to be fast? Consider the environment the DNA is in, and compare that to the structure of DNA while being replicated.
Suggested discussion
What are the pros and cons of DNA polymerase's proofreading capabilities?
Replication mistakes and DNA repair
Although DNA replication is typically a highly accurate process, and proofreading DNA polymerases helps to keep the error rate low, mistakes still occur. In addition to errors of replication, environmental damage may also occur to the DNA. Such uncorrected errors of replication or environmental DNA damage may lead to serious consequences. Therefore, Nature has evolved several mechanisms for repairing damaged or incorrectly synthesized DNA.
Mismatch repair
Some errors are not corrected during replication but are instead corrected after replication is completed; this type of repair is known as a mismatch repair. Specific enzymes recognize the incorrectly added nucleotide and excise it, replacing it with the correct base. But, how do mismatch repair enzymes recognize which of the two bases is the incorrect one?
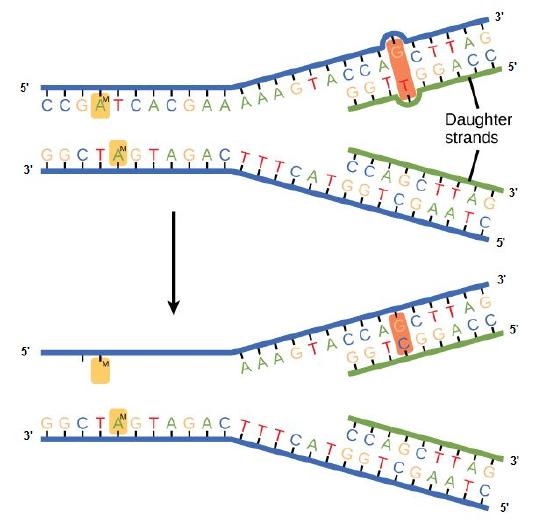
In E. coli, after replication, the nitrogenous base adenine acquires a methyl group; this means that directly after replication the parental DNA strand will have methyl groups, whereas the newly synthesized strand lacks them. Thus, mismatch repair enzymes are able to scan the DNA and remove the wrongly incorporated bases from the newly synthesized, non-methylated strand by using the methylated strand as the "correct" template from which to incorporate a new nucleotide. In eukaryotes, the mechanism is not as well understood, but it is believed to involve recognition of unsealed nicks in the new strand, as well as a short-term, continuing association of some of the replication proteins with the new daughter strand after replication has completed.
Figure 2. In mismatch repair, the incorrectly added base is detected after replication. The mismatch repair proteins detect this base and remove it from the newly synthesized strand by nuclease action. The gap is now filled with the correctly paired base.
Nucleotide excision repair
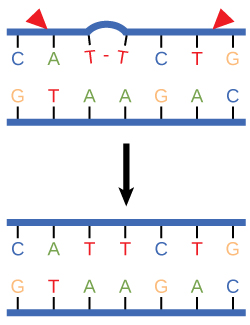
Nucleotide excision repair enzymes replace incorrect bases by making a cut on both the 3' and 5' ends of the incorrect base. The entire segment of DNA is removed and replaced with correctly paired nucleotides by the action of a DNA polymerase. Once the bases are filled in, the remaining gap is sealed with a phosphodiester linkage catalyzed by the enzyme DNA ligase. This repair mechanism is often employed when UV exposure causes the formation of pyrimidine dimers.
Figure 3. Nucleotide excision repairs thymine dimers. When exposed to UV, thymines lying adjacent to each other can form thymine dimers. In normal cells, they are excised and replaced.
Consequences of errors in replication, transcription, and translation
Something key to think about:
Cells have evolved a variety of ways to make sure DNA errors are both detected and corrected. We have already discussed several of them. But why did so many different mechanisms evolve? From proofreading by the various DNA-dependent DNA polymerases, to the complex repair systems. Such mechanisms did not evolve for errors in transcription or translation. If you are familiar with the processes of transcription and/or translation, think about what the consequences would be of an error in transcription. Would such an error affect the offspring? Would it be lethal to the cell? What about errors in translation? Ask the same questions about the process of translation. What would happen if the wrong amino acid is accidentally put into the growing polypeptide during translation? How do these contrast with DNA replication? If you are not familiar with transcription or translation, don't fret. We'll learn those soon and return to this question again.
The flow of genetic information
In bacteria, archaea, and eukaryotes, the primary role of DNA is to store heritable information that encodes the instruction set required for creating the organism in question. While we have gotten much better at quickly reading the chemical composition (the sequence of nucleotides in a genome and some of the chemical modifications that are made to it), we still don't know how to reliably decode all of the information within and all of the mechanisms by which it is read and ultimately expressed.
There are, however, some core principles and mechanisms associated with the reading and expression of the genetic code whose basic steps are understood and that need to be part of the conceptual toolkit for all biologists. Two of these processes are transcription and translation, which are the coping of parts of the genetic code written in DNA into molecules of the related polymer RNA and the reading and encoding of the RNA code into proteins, respectively.
In BIS2A, we focus largely on developing an understanding of the process of transcription (recall that an Energy Story is simply a rubric for describing a process) and its role in the expression of genetic information. We motivate our discussion of transcription by focusing on functional problems (bringing in parts of our problem solving/design challenge rubric) that must be solved the the process to take place. We then go on to describe how the process is used by Nature to create a variety of functional RNA molecules (that may have various structural, catalytic or regulatory roles) including so called messenger RNA (mRNA) molecules that carry the information required to synthesize proteins. Likewise, we focus on challenges and questions associated with the process of translation, the process by which the ribosomes synthesize proteins.
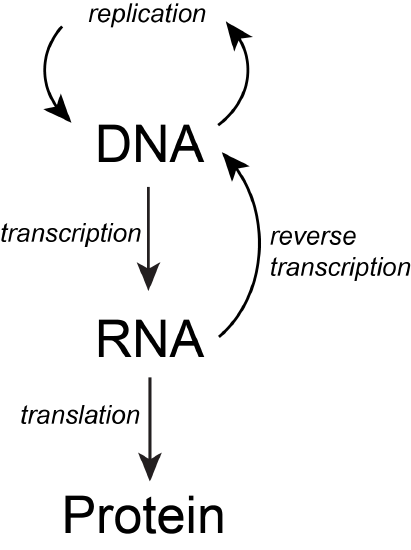
The basic flow of genetic information in biological systems is often depicted in a scheme known as "the central dogma" (see figure below). This scheme states that information encoded in DNA flows into RNA via transcription and ultimately to proteins via translation. Processes like reverse transcription (the creation of DNA from and RNA template) and replication also represent mechanisms for propagating information in different forms. This scheme, however, doesn't say anything per se about how information is encoded or about the mechanisms by which regulatory signals move between the various layers of molecule types depicted in the model. Therefore, while the scheme below is a nearly required part of the lexicon of any biologist, perhaps left over from old tradition, students should also be aware that mechanisms of information flow are more complex (we'll learn about some as we go, and that "the central dogma" only represents some core pathways).
Genotype to phenotype
An important concept in the following sections is the relationship between genetic information, the genotype, and the result of expressing it, the phenotype. These two terms and the mechanisms that link the two will be discussed repeatedly over the next few weeks—start becoming proficient with using this vocabulary.
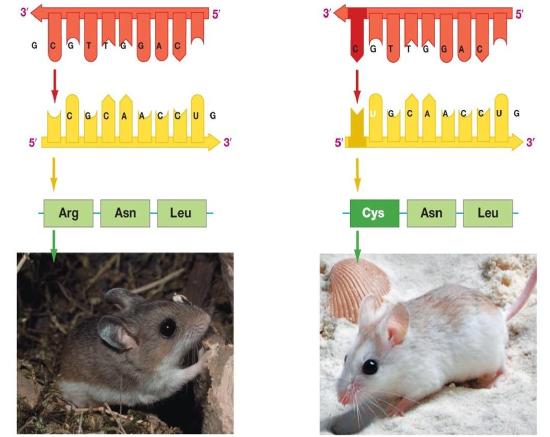
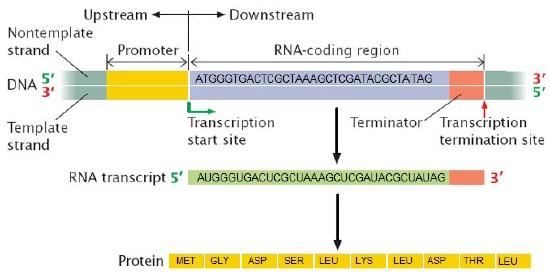
Figure 2. The information stored in DNA is in the sequence of the individual nucleotides when read from 5' to 3' direction. Conversion of the information from DNA into RNA (a process called transcription) produces the second form that information takes in the cell. The mRNA is used as the template for the creation of the amino acid sequence of proteins (in translation). Here, two different sets of information are shown. The DNA sequence is slightly different, resulting in two different mRNAs produced, followed by two different proteins, and ultimately, two different coat colors for the mice.
Genotype refers to the information stored in the DNA of the organism, the sequence of the nucleotides, and the compilation of its genes. Phenotype refers to any physical characteristic that you can measure, such as height, weight, amount of ATP produced, ability to metabolize lactose, response to environmental stimuli, etc. Differences in genotype, even slight, can lead to different phenotypes that are subject to natural selection. The figure above depicts this idea. Also note that, while classic discussions of the genotype and phenotype relationships are talked about in the context of multicellular organisms, this nomenclature and the underlying concepts apply to all organisms, even single-celled organisms like bacteria and archaea.
Note: possible discussion
Can something you can not see "by eye" be considered a phenotype?
Note: possible discussion
Can single-celled organisms have multiple simultaneous phenotypes? If so, can you propose an example? If not, why?
Genes
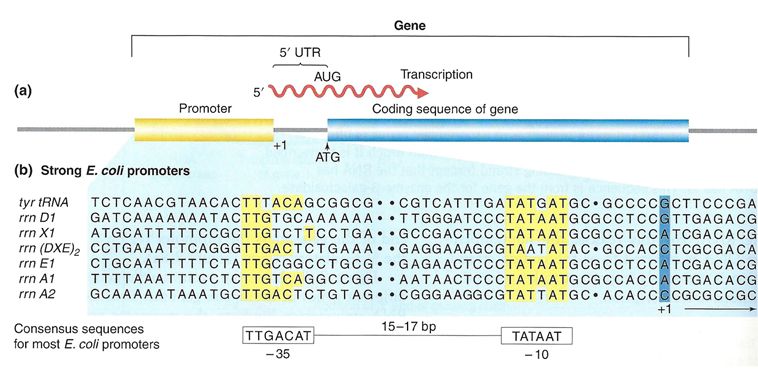
What is a gene? A gene is a segment of DNA in an organism's genome that encodes a functional RNA (such as rRNA, tRNA, etc.) or protein product (enzymes, tubulin, etc.). A generic gene contains elements encoding regulatory regions and a region encoding a transcribed unit.
Genes can acquire mutations—defined as changes in the in the composition and or sequence of the nucleotides—either in the coding or regulatory regions. These mutations can lead to several possible outcomes: (1) nothing measurable happens as a result; (2) the gene is no longer expressed; or (3) the expression or behavior of the gene product(s) are different. In a population of organisms sharing the same gene different variants of the gene are known as alleles. Different alleles can lead to differences in phenotypes of individuals and contribute to the diversity in biology that is under selective pressure.
Start learning these vocabulary terms and associated concepts. You will then be somewhat familiar with them when we start diving into them in more detail over the next lectures.
Elongation
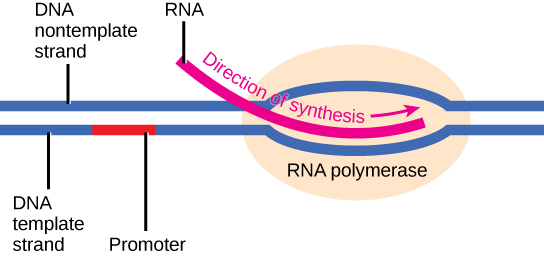
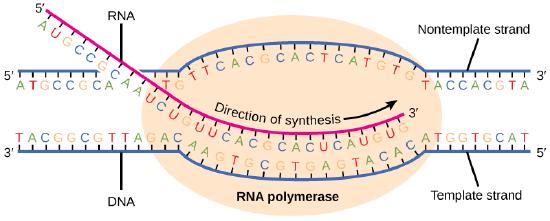
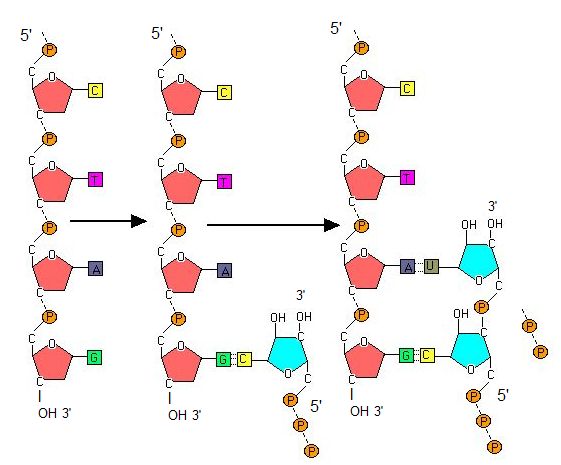
Transcription always proceeds from the template strand, one of the two strands of the double-stranded DNA. The RNA product is complementary to the template strand and is almost identical to the nontemplate strand, called the coding strand, with the exception that RNA contains a uracil (U) in place of the thymine (T) found in DNA. During elongation, an enzyme called RNA polymerase proceeds along the DNA template, adding nucleotides by base pairing with the DNA template in a manner similar to DNA replication, with the difference being an RNA strand that is synthesized does not remain bound to the DNA template. As elongation proceeds, the DNA is continuously unwound ahead of the core enzyme and rewound behind it. Note that the direction of synthesis is identical to that of synthesis in DNA—5' to 3'.
Figure 4. During elongation, RNA polymerase tracks along the DNA template, synthesizing mRNA in the 5' to 3' direction, unwinding and then rewinding the DNA as it is read.
Note: possible discussion
Compare and contrast the energy story for the addition of a nucleotide in DNA replication to the addition of a nucleotide in transcription.
Bacterial vs. eukaryotic elongation
In bacteria, elongation begins with the release of the σ subunit from the polymerase. The dissociation of σ allows the core enzyme to proceed along the DNA template, synthesizing mRNA in the 5' to 3' direction at a rate of approximately 40 nucleotides per second. As elongation proceeds, the DNA is continuously unwound ahead of the core enzyme and rewound behind it. The base pairing between DNA and RNA is not stable enough to maintain the stability of the mRNA synthesis components. Instead, the RNA polymerase acts as a stable linker between the DNA template and the nascent RNA strands to ensure that elongation is not interrupted prematurely.
In eukaryotes, following the formation of the preinitiation complex, the polymerase is released from the other transcription factors, and elongation is allowed to proceed as it does in prokaryotes with the polymerase synthesizing pre-mRNA in the 5' to 3' direction. As discussed previously, RNA polymerase II transcribes the major share of eukaryotic genes, so this section will focus on how this polymerase accomplishes elongation and termination.
Termination
In bacteria
Once a gene is transcribed, the bacterial polymerase needs to be instructed to dissociate from the DNA template and liberate the newly made mRNA. Depending on the gene being transcribed, there are two kinds of termination signals. One is protein-based and the other is RNA-based. Rho-dependent termination is controlled by the rho protein, which tracks along behind the polymerase on the growing mRNA chain. Near the end of the gene, the polymerase encounters a run of G nucleotides on the DNA template and it stalls. As a result, the rho protein collides with the polymerase. The interaction with rho releases the mRNA from the transcription bubble.
Rho-independent termination is controlled by specific sequences in the DNA template strand. As the polymerase nears the end of the gene being transcribed, it encounters a region rich in CG nucleotides. The mRNA folds back on itself, and the complementary CG nucleotides bind together. The result is a stable hairpin that causes the polymerase to stall as soon as it begins to transcribe a region rich in AT nucleotides. The complementary UA region of the mRNA transcript forms only a weak interaction with the template DNA. This, coupled with the stalled polymerase, induces enough instability for the core enzyme to break away and liberate the new mRNA transcript.
In eukaryotes
The termination of transcription is different for the different polymerases. Unlike in prokaryotes, elongation by RNA polymerase II in eukaryotes takes place 1,000–2,000 nucleotides beyond the end of the gene being transcribed. This pre-mRNA tail is subsequently removed by cleavage during mRNA processing. On the other hand, RNA polymerases I and III require termination signals. Genes transcribed by RNA polymerase I contain a specific 18-nucleotide sequence that is recognized by a termination protein. The process of termination in RNA polymerase III involves an mRNA hairpin similar to rho-independent termination of transcription in prokaryotes.
In archaea
Termination of transcription in the archaea is far less studied than in the other two domains of life and is still not well understood. While the functional details are likely to resemble mechanisms that have been seen in the other domains of life, the details are beyond the scope of this course.
Cellular location
In bacteria and archaea
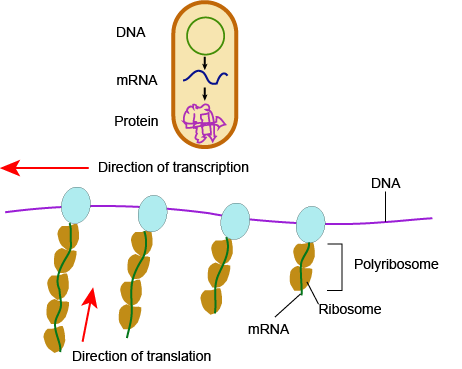
In bacteria and archaea, transcription occurs in the cytoplasm, where the DNA is located. Because the location of the DNA, and thus the process of transcription, are not physically segregated from the rest of the cell, translation often starts before transcription has finished. This means that mRNA in bacteria and archaea is used as the template for a protein before the entire mRNA is produced. The lack of spacial segregation also means that there is very little temporal segregation for these processes. Figure 6 shows the processes of transcription and translation occurring simultaneously.
In eukaryotes....
In eukaryotes, the process of transcription is physically segregated from the rest of the cell, sequestered inside of the nucleus. This results in two things: the mRNA is completed before translation can start, and there is time to "adjust" or "edit" the mRNA before translation starts. The physical separation of these processes gives eukaryotes a chance to alter the mRNA in such a way as to extend the lifespan of the mRNA or even alter the protein product that will be produced from the mRNA.
mRNA processing
5' G-cap and 3' poly-A tail
When a eukaryotic gene is transcribed, the primary transcript is processed in the nucleus in several ways. Eukaryotic mRNAs are modified at the 3' end by the addition of a poly-A tail. This run of A residues is added by an enzyme that does not use genomic DNA as a template. Additionally, the mRNAs have a chemical modification of the 5' end, called a 5'-cap. Data suggests that these modifications both help to increase the lifespan of the mRNA (prevent its premature degradation in the cytoplasm) as well as to help the mRNA initiate translation.
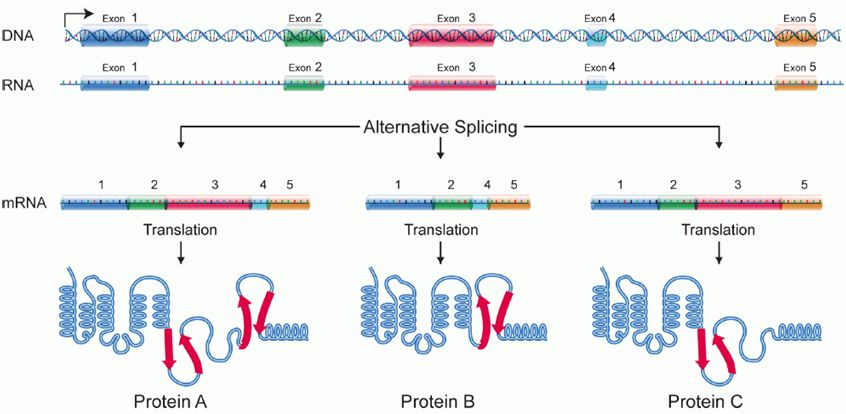
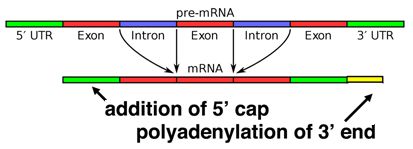
Alternative splicing
Splicing occurs on most eukaryotic mRNAs in which introns are removed from the mRNA sequence and exons are ligated together. This can create a much shorter mRNA than initially transcribed. Splicing allows cells to mix and match which exons are incorporated into the final mRNA product. As shown in the figure below, this can lead to multiple proteins being coded for by a single gene.