6.3: Actin Filaments
- Page ID
- 173593
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Compare and contrast dynamic instability in actin and microtubules.
- Correlate in vivo microfilament polymerization and organization at the cell cortex with its function of deforming the plasma membrane and regulating cell shape.
- Relate how actin-binding proteins influence actin filament polymerization and organization to regulate filament function (e.g., cell motility).
- Interpret results from experiments using drugs that disrupt microfilaments and compare with equivalent experiments that disrupt microtubules.
- Explain the function of myosin motors and their role in contractile bundles as well as in other cellular contexts, such as cytoplasmic streaming or muscle contraction.
The final cytoskeletal element that must be discussed is actin filaments (also known as microfilaments, filamentous actin, f-actin, or just plain “actin”). In animal cells, these filaments are major components of the peripheral cytoskeleton, the cell cortex, and complex contractile systems such as muscle.
RuBisCo is thought to be the most abundant protein on the entire planet, so it beats actin in photosynthetic eukaryotes
Actin filaments can form either stable or dynamic structures, depending on how they are cross-linked to other proteins. As a result, the array of actin-binding proteins (ABPs) that exist are considered extremely important to understanding actin function.
Actin Structure and Filament Formation
Like microtubules, actin filaments are a polymer of actin subunits (sometimes called actin, actin monomers, globular actin, or g-actin). The subunits line up end to end and polymerize in such a way that different sides of the subunit are exposed at either end (Figure 06-18). This means that the filament has a directionality to it, so actin filaments are also referred to as polar molecules (once again, this has nothing to do with charge or other chemical properties per se).

Like microtubules, actin also is capable of dynamic instability even though actin researchers do not commonly discuss actin in those terms. This is, in part, due to the large group of ABPs that are used to control actin monomers and filaments. As a result, the functions of actin filaments and microtubules are very different, as you will see throughout this topic.
Actin and Dynamic Instability
It’s important to note here that in vitro (i.e., in a test tube) actin and tubulin behave very similarly. This means the following:
- They both undergo dynamic instability.
- The plus end has a higher affinity for monomers than the minus end, resulting in
- a lower critical concentration and
- an increased rate of assembly once polymerization starts.
Like tubulin, actin uses chemical energy to drive the process of polymerization (Figure 06-19). However, in this case, ATP is used instead of GTP. In a living cell (i.e., in vivo) there is a high concentration of both ATP and GTP in the cytosol. Thus, it is relatively straightforward to keep the actin monomers in their “active,” ATP-bound state, which favors polymerization. Video 06-08 shows a molecular model of the polymerization of actin.

Actin Function
Actin-Binding Proteins (ABPs)
The cell uses many of the same strategies to control actin filament formation, as we saw in the section on microtubules. Proteins known as ABPs are used extensively to control where and when actin filaments form in the cell. In fact, far more proteins exist to control actin than have been found to control microtubules. This is due, in part, to the number of highly specialized roles that actin plays in different cells. Of course, actin filaments have different functions in the cell compared to microtubules, so the types of binding proteins and the structures formed by actin filaments are distinct from those of microtubules.
Similar to microtubules, actin formation is modulated by the following:
- Controlling where an actin filament forms: Decentralized nucleation sites provide locations to polymerize actin filaments easily. Unlike microtubules, there are no examples of a centralized organizing center for actin. In actin, there are two major types of actin nucleation sites:
- Branching networks—formed when Arp2/3 binds to the side of a preexisting actin filament. We will see this when we discuss cellular locomotion.
- Parallel networks—formed when formin and profilin bind to actin.
- Stabilizing actin filaments using protein caps: Many of the structures formed by actin must be maintained for the long term. ABPs bind to the end of the filament and act as caps to prevent disassembly (Figure 06-20). Examples of these long-lasting, stable actin structures include the microvilli, found in the intestinal tract, and the actin filaments that form the basis for muscle contraction (along with the actin motor protein, myosin).

By controlling where actin forms and when/where it gets stabilized, ABPs and actin can form several different types of actin networks in the cell (known as arrays; Figure 06-21):
- Parallel bundles—filaments are closely spaced and have the same polarity (directionality). For example, parallel bundles are found in filopodia or microvilli. The proteins formin and profilin build these arrays with additional proteins to cross-link.
- Contractile bundles—filaments are arranged antiparallel and are cross-linked by stabilizing proteins. Molecular motors drive the “contraction” of these arrays. Examples include stress fibers in cells and in the contractile ring that splits apart daughter cells during cytokinesis.
- Cross-linked gel—random orientation of fibers linked at the crossing by filamin. Actin gel is a major component of the cell cortex of most cell types and plays a big role in locomotion. Arp2/3 is used to build the filaments in these arrays, and additional proteins are used to cross-link and stabilize.
In association with these internal arrays, actin has also been observed to attach to a variety of plasma membrane proteins, which allows it to help shape the cell and to respond to the cellular environment. Actin is thought to play a key role in cell shape for all cells, but in animal cells, these arrays are the primary driver for most cell shapes.

Myosin Motor Proteins
Just like microtubules, actin filaments also use motor proteins to move cargo along the actin filament. Myosins are a large and diverse class of motor proteins that use actin filaments as their substrate.
Myosin motors are involved in a number of cellular functions (Figure 06-22), including the following:
- vesicle transport (Myosin I is the most important myosin for this, but others also get involved.)
- organelle positioning
- cytokinesis
- cellular locomotion and shape
- phagocytosis
- muscle contraction (Myosin II: We know quite a lot more about this myosin than any of the others, as quite a lot of research is done on muscle function.)
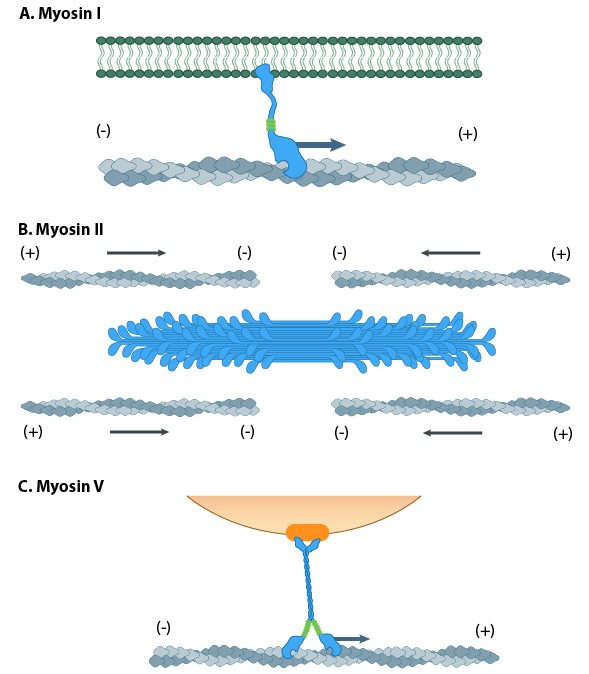
The myosin family of proteins is very large. There are as many as 18 different types of myosin motors found in eukaryotes. Most of these myosins move toward the plus-end of actin filaments; however, there are some exceptions. One subtype, called myosin IV, has been shown to move toward the minus-end. Myosin motors are complexes of several polypeptide chains (Figure 06-22). Some subtypes have one motor head, and some have two. However, the motor head functions the same way regardless of how many there are.

In a nutshell, they bind to the subunits of the actin filament, pull their motor head in, and then release with the help of ATP. Different myosins appear to move slightly differently, so it’s more difficult to find a single video to represent them all. Muscle myosins (myosin II) work through a cycle that includes a “power stroke” that pulls the actin filament in toward the center of the muscle (Video 06-09). Myosin V, on the other hand, appears to use a very different mechanism that is similar to the movement of a Slinky toy (Video 06-10).
Like the microtubule motors, myosin motors require ATP in order to function. The ATP hydrolysis helps prime the motor head to prepare for the next time it binds and pulls on the filament.
Interestingly, no ATP is required for binding to the actin filament, but every other step of the process requires either ATP or ADP. This is the opposite of the microtubule motors, which cannot bind to the microtubule (MT) unless they are bound to ATP.
As a result of this unique situation, myosin motors will freeze in place if the cell runs out of ATP, which happens at the point of death or shortly thereafter. Muscle contraction is dependent on successive rounds of binding and unbinding of myosin, so without ATP, the myosin binds and causes the muscles to become really rigid right after death. This is a well-known phenomenon called rigor mortis. It is commonly referenced in detective shows, especially those with a focus on forensic science.
Example: Cellular Locomotion
Locomotion in animal cells is a good example of how precise control of dynamic instability can be used to the cell’s advantage. Cellular locomotion is a complex interplay of holding on to the substrate, building a network in one area of the cell, and deconstructing the network in another area (Figure 06-24). Every step of this process requires ABPs to make sure that it all goes according to plan.

The act of forming the membrane protrusion at the leading edge requires actin polymerization directly underneath the plasma membrane. Nucleation of new actin filaments at the leading edge is mediated by Arp2/3 complexes. As you may recall, Arp2/3 binds to the side of a preexisting filament so that a branched array is formed (Figure 06-24). The growing actin network is what pushes the leading edge forward, creating membrane waves, or ruffles, called a lamellipodium (plural lamellipodia). At the same time, the polymerized actin at the back end of the leading edge (i.e., on the far side of the branched network from the membrane) is depolymerized so that free actin can be moved forward. This ensures that the cell continues to have access to free actin for polymerization at the leading edge.
As the leading edge is being pushed forward by the lamellipodia, the back of the cell also needs to be pulled forward, so the cell moves forward instead of just flattening out. This contraction is accomplished by a network of actin and myosin motors (sometimes referred to as the actomyosin network) that contract within the cell body, creating tension. As it does this, the proteins holding onto the substratum at the rear release so that the contraction can happen, and the cell moves forward.
Cellular locomotion is well illustrated by Video 06-11, filmed in the 1950s using a light microscope attached to a film camera. In it we see a white blood cell chasing, capturing, and engulfing a bacterium.
Studying Cells: Drugs That Influence Actin and Microtubule Function
Before we leave our discussion of the cytoskeleton, we must once again explore how scientists study the function of the cytoskeleton. As we’ve seen before, a commonly used tactic in biology is to disrupt a biological system and observe the results. You may not be surprised to know that both actin and microtubules are essential cellular structures. So mutational analysis is not a very good option for experimental study.
However, since the cytoskeleton is so important, especially as a key player in cell division and muscle contraction, it has become the target of many natural toxins produced by a variety of plants, animals, and fungi in order to protect themselves from other organisms in their environment. As scientists, we are able to exploit these compounds to our advantage and use them to temporarily disrupt the cytoskeleton and then observe the impact on cellular function. Table 06-01 shows the most commonly used compounds and how they impact the cytoskeleton when applied.
| Microtubule-specific | ||
|---|---|---|
| Compound | Action | Source |
| Paclitaxel | Binds to microtubule and prevents disassembly | Bark of Pacific yew (Taxus brevifolia) |
| Colchicine | Binds to unpolymerized tubulin and prevents assembly | Autumn crocus (Colchicum autumnale) |
| Vinblastine | Binds to unpolymerized tubulin and prevents assembly | Madagascar periwinkle (Catharanthus roseus) |
| Oryzalin | In plants only; binds to unpolymerized tubulin and prevents assembly | Synthetic herbicide in the dinitroaniline family |
| Actin-specific | ||
| Compound | Action | Source |
| Phalloidin | Binds to actin filament and prevents disassembly | Death cap mushroom (Amanita phalloides) |
| Latrunculin | Binds to unpolymerized actin and prevents assembly | Sea sponges in the Latrunculia and Negombata families |
| Cytochalasin B or D | Caps the plus end of actin filaments | Fungal plant pathogen of the Helminthosporum genus |
What we hope you will notice in the table is how varied the effects of the drugs are. While some of them interact directly with the polymerized microtubule, others work by disrupting the chemical equilibrium through interactions with the monomer. This shows how essential dynamic instability is to the proper function of both actin and microtubules.
One final note: There is often confusion between the difference in “capping” and “stabilizing,” especially with respect to the function of these compounds. A capped filament is also stabilized, as the polymer cannot fall apart. But it is possible to stabilize without capping. A perfect illustration of this is the difference between paclitaxel, which stabilizes microtubules without capping so that they may still grow (but not shrink), and something like cytochalasin, which acts as a protein cap at the plus end of the actin filament. Cytochalasin stops the microtubule from growing or shrinking, whereas paclitaxel only inhibits shrinking but not growth.


