iCn3D Skill: Analysis of Noncovalent Interactions
- Page ID
- 114616
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Structure
- PDB ID: 3K83
- Protein: Inhibitor (below, abbreviated F278458) bound to a humanized variant of Fatty Acid Amide Hydrolase (FAH)

- Activity: FAH Catalyzes the hydrolysis of endogenous amidated lipids like the sleep-inducing lipid oleamide, the endocannabinoid anandamide, and other fatty amides, regulating the signaling functions of these molecules
- Description: The functional unit is a dimer, each with an active site. A chloride binds between the two subunits
In this activity you will see key noncovalent interactions between the inhibitor and (FAH). You will pick one inhibitor and view its noncovalent interactions with one subunit
Modeling Instructions
- Open a new iCn3D window, 3K83 at: https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html
- Select, Select on 3D to ensure default is Residue; With mouse, zoom and center to Alt-Click the inhibitor (F278458) in the magenta subunit (option-click on a Mac).
- Select, Save Selection, name it Drug
- Analysis, Interactions.
- Under 1. Choose interaction types and their thresholds, Check only the noncovalent interactions shown below. Make sure the Contacts/Interactions is unchecked as it shows all interactions between contacting Van der Waals surfaces including many hydrophobic interactions, so it is quite cluttered.
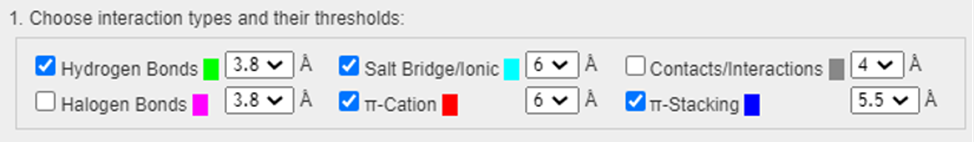
- Select the Drug as the first set (under 2. Select the first set); choose 3K83 (under 3. Select the second set)
- Click on the box: 4. 3D Display Interactions. This will display interacting side chains. Drag the HBonds/Interactions window to the bottom right away from the molecular display. Note the coloring of the dotted lines:
Green - hydrogen bonds
Red - π-cation
Blue - π stacking
- Select, Save Selection, name it Interactions
- Style, Sidechains, Stick (so the next step will color side chains)
- Color, atom (Note: this drug is covalently bound to Serine241)
- In defined sets Ctrl click Drug and Interaction to highlight all
- View, View Selection (to only see Drug and Interactions)
- Select, Toggle Highlight (to remove highlights if necessary)
Note: Once you run interactions, iCn3D adds many new additions to the Defined sets window. Explore these to learn different ways to examine the interactions.
Optional (if you’d like to add labels, recolor the background, and obtain the share link as in the previous activity)
- Analysis, Label, Per Residue & No
- Analysis, Label Scale, pick a number that works for you
- Style, Background, Transparent
- File, Share Link, copy short link
Pre-Rendered Model Link
To check your work (or if you got stuck during any of the steps above) view the model using this link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?pDr5EBZmo3TyAbTP6
Note: You can alter interactions on the 3D model, or examine 2D displays of them. Try:
● Click 5. Reset at the bottom of the HBonds/Interactions window. Add Contacts/ Interactions (mostly nonpolar) by clicking on the box. Choose Drug as the first set. Choose 3K83 as the second set. Then click 4. 3D Display Interactions to show.
● Next, try 4. 2D Interaction Network. In the popup that opens, click a colored line in the 2D window to highlight a specific interaction.
If using the Pre-Rendered model: From the top menu, select: Analysis → Interactions. In the popup choose 5. Reset, select 3K83 (full protein) as set 1 and drug as set 2, then 4. 2D Interaction Network. Click colored lines in the 2D window.

