BioMolViz Theme: Macromolecular Building Blocks (MB) ✔
- Page ID
- 114275
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Learning themes, goals, and objectives to promote biomolecular visualization literacy are articulated in a Biomolecular Visualization Framework created by BioMolViz. Here we present models that illustrate key learning goals (MB1, MB2) and objectives for the theme Macromolecular Building Blocks (MB)
Goal MB1. Students can identify individual building blocks of biological polymers.
MB1.01 Given a rendered structure of a biological polymer, students will be able to identify the ends of a biological polymer. (Novice, Amateur, Expert)
MB1.02 Given a rendered structure, students can divide the polymer into its individual building blocks. (Novice)
MB1.03 Given a rendered structure, students can identify the individual building blocks. (Novice)
Click the images below to see pre-rendered examples to illustrate the MB theme.
Proteins
Example: 1L2Y - Trp-Cage Miniprotein Construct TC5b, 20 amino acids artificial protein construct
| iCn3D GUI commands | iCn3D structure (click on image) |
|
To duplicate the image, open iCn3D, load 1L2Y and perform these commands
|
The protein backbone in CPK colors |
|
Continue with this link
|
The backbone in sequential rainbow colors |
|
Continue with the previous render
|
The full protein (backbone and sidechains). Each amino acid is colored differently, starting with the N-terminal in red, and continuing with the colors of the spectrum. |
|
Continue with the previous render
|
The protein backbone atoms are shown in CPK colors and the sidechains are shown with sequential rainbow colors |
|
Continue with the previous render Analysis, Color, Atom Analysis, Label, Per Residue and Number Analysis, Label Scale, 4 |
All atoms are in CPK colors with the sequential amino acids labeled |
Nucleic Acid
Example: dsDNA - 1BNA
|
To duplicate the image, open iCn3D, load 1BNA and perform these commands
|
The backbone (sugar, phosphate) of each chain in CPK colors |
|
Continue with above model
|
The backbone (sugar, phosphate) in sequential rainbow colors |
|
The deoxynucleotides (sugar, phosphate, base) in each chain in sequential rainbow colors |
|
The backbone of each chain in CPK colors, with the bases in rainbow colors |
Example: Part of a stem-loop from a messenger RNA (4QOZ)
| PDB ID, iCn3D GUI commands | iCn3D structure (click on image) |
|
4QOZ- some iCn3D commands For the second render, start with this link
|
Left: nucleotide backbone (sugar, phosphate) in CPK colors. Right: Full RNA with the backbone in CPK and bases in rainbow colors. |
Glycans
| PDB ID, iCn3D GUI commands | iCn3D structure (click on image) |
|
3LL2 - some iCn3D commands
|
Left: glycan in CPK colors; Middle: glycan CPK colors with SNFG cartoons for monomers; Right: glycan monomers in sequential rainbow colors |
|
1CAP
|
Left: glycan in CPK colors; Middle: glycan with rainbow colors; Right: Glycan monomers in sequential rainbow colors and SNFG colors |
Mixed Polymer
Use this iCn3D structure and answer the following questions: https://structure.ncbi.nlm.nih.gov/i...155e9ovENSGa67
- Identify the polymer type(s)
- Determine the sequences
- Color code the structure to show the building blocks (monomers)
- Answer
-
- There are two polymers, a protein (short peptide) and RNA (short segment)
- Protein: Ala(472)-Arg-Ser-Cys(475)-SS-Cys(98)-Arg97. The protein consists of two peptides fragments connected by a disulfide bond.
- Here is the link - https://structure.ncbi.nlm.nih.gov/i...3iRYoHriLKidv8. The protein fragments are labeled and the RNA is shown in rainbow colors
Use this iCn3D structure and answer the following questions: https://structure.ncbi.nlm.nih.gov/i...9BmRTG7iPAWPs7
- Identify the polymer type(s)
- Determine the sequences
- Color code the structure to show the building blocks (monomers)
- Answer
-
- There are two polymers, a protein (short peptide) and glycan (short branched segment)
- Protein: 60 YKLFKNGTLKI 70; Glycan: contains NAG (N-acetylglucosamine, blue squares) and BMA (β-MAN)and MAN (α-mannose), both in green circles.

- https://structure.ncbi.nlm.nih.gov/i...uQXx5pAwmAZNi8
Goal MB2: Students can describe the contributions different individual building blocks make in determining the 3D shape of the polymer.
MB2.01 Students can describe the physical/chemical properties of an individual building block/functional group in a rendered structure of a polymer. (Amateur)
MB2.02 Students can describe the significance of the location of individual building blocks within the 3D structure of a polymer (e.g. protein, carbohydrate or nucleic acid). (Novice, Amateur, Expert)
MB2.03 Students can identify physical/chemical properties of individual building blocks/functional groups in different local environments. (Amateur)
MB2.04 Using a visualized structure, students can identify stereochemistry (e.g. in carbohydrate, lipid, and protein structures). (Amateur)
MB2.05 Students can modify/mutate a building block to change the 3D structure of a polymer (e.g. protein, carbohydrate or nucleic acid). (Amateur, Expert)
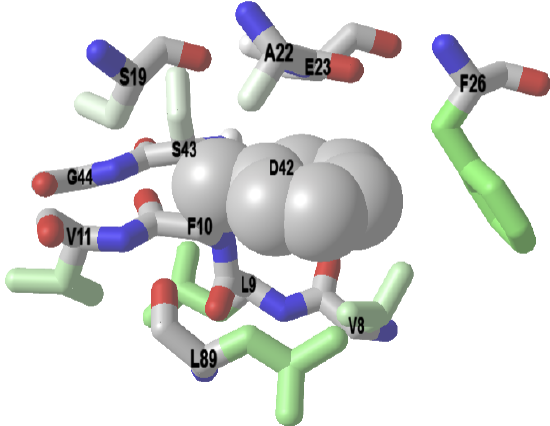
Example: Bovine low molecular weight protein tyrosine phosphatase - LMW-PTP (1xww)
|
1xww Follow these instructions to reproduce the image to the right.
|
A significant fraction of nonpolar amino acid side chains are buried in protein. Surrounding them are typically other nonpolar side chains. This maximizes nonpolar interactions and removes them from the aqueous exterior, a process that is entropically favored. The iCn3D below shows a buried Phe 10 (F10) surrounded mostly by nonpolar side chains. Burial of nonpolar side chains into a localized nonpolar environment occurs early and seeds protein folding. Amino acids within 4 Angstroms of Phe 10 (F10) in LMW-PTP. The F10 sidechain is shown in spheres.
|
Example: human frataxin (1EKG)
|
Follow these instructions to reproduce the image to the right.
|
The wild-type human protein has a Leu 106. A missense mutation caused by T to C DNA change at position 317 in exon 3 causes a L106S mutation in the protein, which leads to protein misfolding and disease (Friedreich’s Ataxia). This is a neurodegenerative disease that causes loss of muscle control (ataxia), heart dysfunction (cardiomyopathy), and diabetes. Click on the image. Press the "a" key to toggle back and forth between the wild-type and mutant protein |