1.4: Genetic Foundations
- Page ID
- 14916
Introduction
The development of complex biological organisms on our planet has arisen through the evolutionary mechanism of natural selection. The British naturalist, Charles Darwin proposed the theory of biological evolution by natural selection in his book, ‘On the Origins of Species’ that was published in 1859. Darwin defined evolution as “descent with modification,” the idea that species change over time, give rise to new species, and share a common ancestor. The mechanism that Darwin proposed for evolution is natural selection. Because resources are limited in nature, organisms with heritable traits that favor survival and reproduction will tend to leave more offspring than their peers, causing the traits to increase in frequency within a population over generations. Thus, natural selection causes populations to become adapted, or increasingly well-suited, to their environments over time. Natural selection depends on the environment and requires existing heritable variation in a group.
Natural selection acts on an organism’s phenotype, or physical characteristics. Phenotype is determined by an organism’s genetic make-up (genotype) and the environment in which the organism lives. When different organisms in a population possess different versions of a gene for a certain trait, each of these versions is known as an allele. It is primarily this genetic variation that underlies differences in phenotype. Some traits are governed by only a single gene, but most traits are influenced by the interactions of many genes. A variation in one of the many genes that contribute to a trait may have only a small effect on the phenotype; together, these genes can produce a continuum of possible phenotypic values.
For example, interactions between different equine coat color genes determine a horse’s coat color. Many colors are possible, but all variations are produced by changes in only a few genes. Extension and agouti are particularly well-known genes with dramatic effects. For example, differences at the agouti gene can help determine whether a horse is bay or black in coloration, and a change to the extension gene can in turn make a horse chestnut-colored instead (Figure 1.30). Yet other gene variants are responsible for the myriad of other coat color possibilities, including palomino, buckskin, and cremello horses.

Thus, the primary molecular mechanism that drives natural selection is controlled by the heritability and mutability of genetic traits housed in the major macromolecule, deoxyribonucleic acid (DNA). In chapter 4, you will learn about the structural characteristics of DNA, whereas chapter 9 focuses on the biochemical mechanisms involved with DNA replication and also details the importance of DNA repair process and molecular mechanisms of evolution at the genetic level.
Genetic Code
Notably, the phenotypic traits determined by the genetic makeup of an organism are not controlled directly by the genetic material, DNA, but by the proteins that are produced from the information housed within the gene. In 1945, geneticist George Beadle proposed the one gene-one enzyme hypothesis suggesting that genes are highly specific when they encode for a protein sequence. However, it would take 16 more years before the biochemical nature of this process was deduced. Efforts to understand how proteins are encoded began after DNA’s structure was discovered in 1953. George Gamow postulated that sets of three bases must be employed to encode the 20 standard amino acids used by living cells to build proteins, which would allow a maximum of 43 = 64 amino acids.
The Crick, Brenner, Barnett and Watts-Tobin experiment first demonstrated that codons consist of three DNA bases (Figure 1.31). Marshall Nirenberg and Heinrich J. Matthaei were the first to reveal the nature of a codon in 1961.
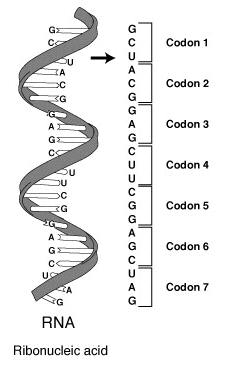
They used a cell-free system to translate a poly-uracil RNA sequence (i.e., UUUUU…) and discovered that the polypeptide that they had synthesized consisted of only the amino acid phenylalanine. They thereby deduced that the codon UUU specified the amino acid phenylalanine.
This was followed by experiments in Severo Ochoa‘s laboratory that demonstrated that the poly-adenine RNA sequence (AAAAA…) coded for the polypeptide poly-lysine and that the poly-cytosine RNA sequence (CCCCC…) coded for the polypeptide poly-proline. Therefore, the codon AAA specified the amino acid lysine, and the codon CCC specified the amino acid proline. Using various copolymers most of the remaining codons were then determined.
Subsequent work by Har Gobind Khorana identified the rest of the genetic code. Shortly thereafter, Robert W. Holley determined the structure of transfer RNA (tRNA), the adapter molecule that facilitates the process of translating RNA into protein. This work was based upon Ochoa’s earlier studies, yielding the latter the Nobel Prize in Physiology or Medicine in 1959 for work on the enzymology of RNA synthesis.
Extending this work, Nirenberg and Philip Leder revealed the code’s triplet nature and deciphered its codons (Figure 1.32). In these experiments, various combinations of mRNA were passed through a filter that contained ribosomes, the components of cells that translate RNA into protein. Unique triplets promoted the binding of specific tRNAs to the ribosome. Leder and Nirenberg were able to determine the sequences of 54 out of 64 codons in their experiments. Khorana, Holley and Nirenberg received the 1968 Nobel for their work.
The three stop codons were named by discoverers Richard Epstein and Charles Steinberg. “Amber” was named after their friend Harris Bernstein, whose last name means “amber” in German. The other two stop codons were named “ochre” and “opal” in order to keep the “color names” theme.

Each gene contains a reading frame is defined by the initial triplet of nucleotides from which translation starts. It sets the frame for a run of successive, non-overlapping codons, which is known as an open reading frame (ORF). For example, the string 5′-AAATGAACG-3′, if read from the first position, contains the codons AAA, TGA, and ACG ; if read from the second position, it contains the codons AAT and GAA ; and if read from the third position, it contains the codons ATG and AAC. Every sequence can, thus, be read in its 5′ → 3′ direction in three reading frames, each producing a possibly distinct amino acid sequence: in the given example, Lys (K)-Trp (W)-Thr (T), Asn (N)-Glu (E), or Met (M)-Asn (N), respectively. When DNA is double-stranded, six possible reading frames are defined, three in the forward orientation on one strand and three reverse on the opposite strand. Protein-coding frames are defined by a start codon, usually the first AUG (ATG) codon in the RNA (DNA) sequence.
To terminate the translation process, there are three stop codons: UAG is amber, UGA is opal (sometimes also called umber), and UAA is ochre. Stop codons are also called “termination” or “nonsense” codons. They signal the release of the nascent polypeptide from the ribosome.
During the process of DNA replication, errors occasionally occur in the polymerization of the second strand. These errors, called mutations, can affect an organism’s phenotype, especially if they occur within the protein-coding sequence of a gene. Error rates are typically 1 error in every 10–100 million bases—due to the “proofreading” ability of DNA polymerases.
Missense mutations and nonsense mutations are examples of point mutations that can cause genetic diseases such as sickle-cell disease and thalassemia respectively. Clinically important missense mutations generally change the properties of the coded amino acid residue among basic, acidic, polar or non-polar states, whereas nonsense mutations result in a stop codon.
Mutations that disrupt the reading frame sequence by indels (insertions or deletions) of a non-multiple of 3 nucleotide bases are known as frameshift mutations. These mutations usually result in a completely different translation than from the original RNA, and likely cause a stop codon to be read, which truncates the protein. These mutations may impair the protein’s function and are thus rare in in vivo protein-coding sequences. One reason inheritance of frameshift mutations is rare is that, if the protein being translated is essential for growth under the selective pressures the organism faces, the absence of a functional protein may cause death before the organism becomes viable. Frameshift mutations may result in severe genetic diseases such as Tay–Sachs disease.
Although most mutations that change protein sequences are harmful or neutral, some mutations have benefits. These mutations may enable the mutant organism to withstand particular environmental stresses better than wild-type organisms, or reproduce more quickly. In these cases, a mutation will tend to become more common in a population through natural selection. Different sequence variations of the same gene or protein within a single organism, within a population, or between different species are known as sequence polymorphisms. Larger-scale gene duplication events can also lead to evolutionary events.
Similar Proteins
The evolution of proteins is studied by comparing the sequences and structures of proteins from many organisms representing distinct evolutionary clades. If the sequences/structures of two proteins are similar indicating that the proteins diverged from a common origin, these proteins are called homologous proteins. More specifically, homologous proteins that exist in two distinct species are called as orthologs. In contrast, homologous proteins encoded by the genome of a single species are called paralogs. Unrelated genes that have separate evolutionary origins, but that each encode proteins that have similar functions, are termed analogs (Figure 1.33).

DNA sequencing techniques have rapidly improved over the last 15 to 20 years making it possible to sequence the entire genomes of organisms and thus, predict the entire proteome of an organism, based on the translation of the sequenced genome followed by the annotation of predicted ORFs using phylogenetic comparison of similar genes/proteins from other known organisms. This has given rise to the field of Bioinformatics which uses computer science, mathematics, and statistical analysis to analyze the large quantities of biological data created in genome sequencing projects. The phylogenetic relationships, and hence ancestral relationships, of various genes, proteins, and ultimately organisms can be established through the statistical analysis of sequence alignments. Such phylogenetic trees have established that the sequence similarities among proteins reflect closely the evolutionary relationships among organisms.
Protein evolution describes the changes over time in protein shape, function, and composition. Through quantitative analysis and experimentation, scientists have strived to understand the rate and causes of protein evolution. Using the amino acid sequences of hemoglobin and cytochrome c from multiple species, scientists were able to derive estimations of protein evolution rates. What they found was that the rates were not the same among proteins. Each protein has its own rate, and that rate is constant across phylogenies (i.e., hemoglobin does not evolve at the same rate as cytochrome c, but hemoglobins from humans, mice, etc. do have comparable rates of evolution.). Not all regions within a protein mutate at the same rate; functionally important areas mutate more slowly and amino acid substitutions involving similar amino acids occurs more often than dissimilar substitutions. Overall, the level of polymorphisms in proteins seems to be fairly constant. Several species (including humans, fruit flies, and mice) have similar levels of protein polymorphism.
Gene duplication events followed by mutation can also give rise to paralogs, with unique and different functions within an organism. This can make the annotation of genomes based on sequence alone a difficult task, as homologous protein sequences may not have similar functions in vivo. It is estimated that approximately 10-25% of annotations made on sequence homology are incorrect and require experimental validation. For example, human pancreatic ribonuclease is a digestive enzyme utilized to breakdown nucleic acids. The angiogenin protein is a paralog of pancreatic ribonuclease and shares high sequence homology and 3-D shape (Figure 1.34). However, the functions of these proteins are quite different. Angiogenin induces vascularization by activating transcriptional processes in endothelial cells. However, if the function of only one of these homologs was known, it would be easy to mistakenly hypothesize that the homologous protein would be similar in function. Thus, care must be taken when using bioinformatic tools to not overestimate the predictive ability of sequence alignments.

The control of gene expression is critical in all processes of life, allowing for the differentiation of cells to form different body structures and organs, as well as smaller more reversible changes that allow an organism to respond to different environmental situations and stimuli. In chapter 12, you will explore the major biochemical mechanisms used to control gene expression within cells. This will include the discussion of a fairly new and exciting field of study known as epigenetics. In addition to the heritability of traits through the passage of genetic information, it is fast becoming clear that the environmental factors that an organism is exposed to throughout its life can affect gene expression without physically altering the DNA sequence, and that these changes in expression patterns can be long-lasting and can even be inherited in the following generations. The term epigenetics literally means ‘on top of’ or ‘in addition to’ genetics and focuses on the heritable gene expression patterns that are induced by the exposure or experience of an organism within its environment.
For example in human populations, stressful events such as starvation can have lasting imprints on children that are born under these conditions. These children have higher risks of obesity and metabolic disorders as adults, including the development of type II diabetes. In fact, these predispositions can be carried not only to the children born during starvation but also to their future children indicating that environmental events can affect gene expression patterns through multiple generations. In more controlled laboratory experiments using rats, it has been demonstrated that the more a mother rat licks and nurtures its offspring, the calmer and more relaxed the offspring will be as an adult. Mother rats that are less nurturing and ignore their young, have offspring that will grow up displaying higher levels of anxiety. These changes are not caused by genetic differences between the offspring, but rather by differences in gene expression patterns. In fact, calm and relaxed mice can be altered to show high anxiety by exposing them to agents that alter gene expression patterns. Mechanisms controlling such heritable alterations in gene expression patterns will be covered in a future chapter.
Central Dogma of Biology
DNA encodes the genetic material. It must be replicated on cell division. Its information is decoded into an RNA in a process called transcription. That information is decoded to form a protein sequence. Collectively these processes are referred to as the Central Dogma of Biology. A variant occurs when RNA is decoded into DNA, a process called reverse transcription. These processes are described briefly below and in great depth in subsequent chapters.
Replication
DNA must be duplicated in a process called replication before a cell divides. The replication of DNA allows each daughter cell to contain a full complement of chromosomes.
Transcription and Splicing
For a given gene, only one strand of the DNA serves as the template for transcription. An example is shown below. The bottom (blue) strand in this example is the template strand, which is also called the minus (-) strand, or the sense strand. It is this strand that serves as a template for mRNA synthesis. The enzyme RNA polymerase synthesizes an mRNA in the 5' to 3' direction complementary to this template strand. The opposite DNA strand (red) is called the coding strand, the nontemplate strand, the plus (+) strand, or the antisense strand.
The easiest way to find the corresponding mRNA sequence (shown in green below) is to read the coding, nontemplate, plus (+), or antisense strand directly in the 5' to 3' direction substituting U for T.
5' T G A C C T T C G A A C G G G A T G G A A A G G 3' 3' A C T G G A A G C T T G C C C T A C C T T T C C 5'
5' U G A C C U U C G A A C G G G A U G G A A A G G 3'
As we've learned more about the structure of DNA, RNA, and proteins, it become clear that transcription and translation differ in eukaryotes and prokaryotes. Specifically, eukaryotes have intervening sequences of DNA (introns) within a given gene that separate coding fragments of DNA (exons). A primary transcript is made from the DNA, and the introns are sliced out and exons joined in a contiguous stretch to form messenger RNA which leaves the nucleus. Translation occurs in the cytoplasm. Remember, prokaryotes do not have a nucleus.
Translation
Information in a mRNA sequence is decoded to form a protein. In this process, a triplet of nucleotides (a codon) in the RNA has the information of a single amino acid. Translation occurs on a large RNA-protein complex called the ribosome. An intermediary transfer RNA (tRNA) molecule becomes covalently linked to a single amino acid by the enzyme tRNA-acyl synthetase. This "charged" tRNA binds through a complementary anticodon region to the triplet codon in the tRNA. The ribosome/tRNA complex ratchets down the mRNA allowing a new "charged" tRNA complex to bind at an adjacent site. The two adjacent amino acids form a peptide bond in a process driven by ATP cleavage. This process repeats until a "stop" codon appears in the mRNA sequence. The genetic code shows the relationship between the triplet mRNA codon and the amino acid which corresponds to it in the growing peptide chain.
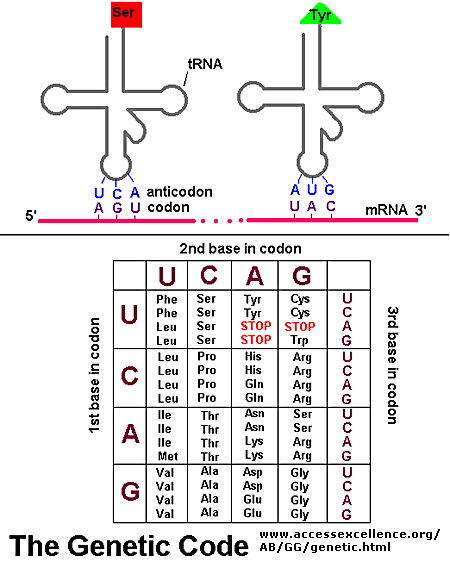
As was mentioned in the Protein Chapter (amino acid section) two other amino acids occasionally appear in proteins (excluding amino acids altered through post-translational modification. One is selenocysteine, which is found in Arachea, eubacteria, and animals. The other is just recently found is pyrrolysine, found on Arachea. These new amino acids derive from modifications of Ser-tRNA and probably Lys-tRNA after the tRNA is charged with the normal amino acid, to produce selenocys-tRNA and pyrrolys-tRNA, respectively. The pyrrolysine-tRNA recognizes the mRNA codon UAG which is usually a stop codon, while selenocys-tRNA recognizes UGA, also a stop codon. Clearly, only a small fraction of stop codons in mRNA sequences would be recognized by this usual tRNA complex. What determines that recognition is unclear.
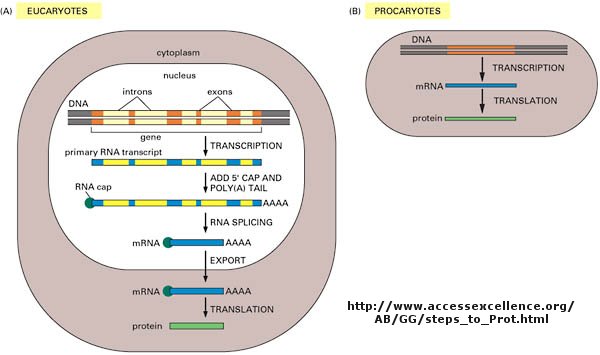
What is a gene?
The definition of a gene can differ depending on whom you ask. The world gene has literally become a cultural icon of our day. Can our genes explain what it is to be human? The definition of a gene has changed with time. Eukaryotic genes contain exons (coding regions) and introns (intervening sequences) that are transcribed to produce a primary transcript. In a post-transcriptional process, introns are spliced out by the spliceosome, to produce a messenger RNA, mRNA, which is translated into a protein sequence. (See diagram above).
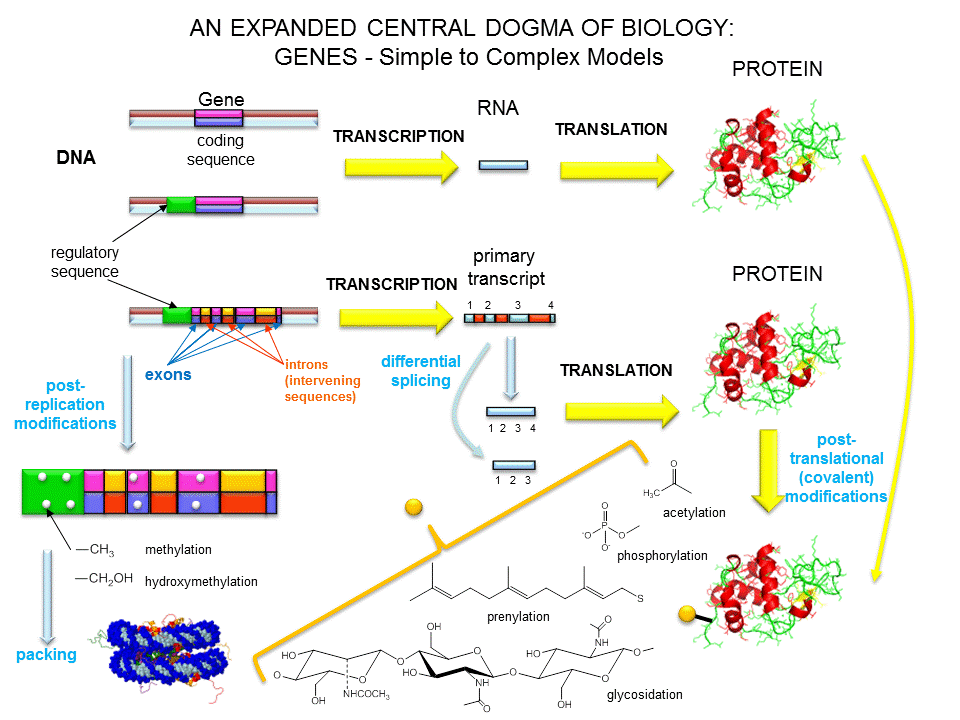
Over the last 100 years, as our understanding of biochemistry has increased, the definition of a gene has evolved from
- the basis of inheritable traits
- certain regions of chromosomes
- a segment of a chromosome that produces one enzyme
- a segment of a chromosome that produces one protein
- a segment of a chromosome that produces a functional product
The last definition was necessary since some gene products that have functions (structural and catalytic) are RNA molecules. The last definition also includes regulatory regions of the chromosome involved in transcriptional control. Snyder and Gerstein have developed five criteria that can be used in gene identification which is important as the complete genomes of organisms are analyzed for genes.
- identification of an open reading frame (ORF) - this would include a series of codons bounded by start and stop codons. This gets progressively harder to do if the gene has a large number of exons embedded in long introns.
- specific DNA features within genes - these would include a bias towards certain codons found in genes or splice sites (to remove intron RNA)
- comparing putative gene sequences for homology with known genes from different organisms, but avoiding sequences that might be conserved regulatory regions.
- identification of RNA transcripts or expressed protein (which does not require DNA sequence analysis as the top three steps do) -
- inactivating (chemically or through specific mutagenesis) a gene product (RNA or protein).
New findings make it even more complicated to define a gene, especially if the transcripts of a "gene region" are studied. Cheng et al studied all transcripts from 10 different human chromosomes and 8 different cell lines. They found a large number of different transcripts, many of which overlapped. Splicing often occurs between nonadjacent introns. Transcripts were found from both strands and were from regions containing introns and exons. Other studies found up to 5% of transcripts continued through the end of "gene" into other genes. 63% of the entire mouse genome, which is comprised of only 2% exons, is transcribed.
The Language of DNA
In this short chapter, you will briefly learn how modern molecular biologists manipulate DNA, the blueprint for all of life. The details will be found in subsequence chapters. The four-letter alphabet (A, G, C, and T) that makes up DNA represents a language that when transcribed and translated leads to the myriad of proteins that make us who we are as a species and as individuals. Let's continue with the metaphor that DNA is a language. To master that language, as with any other language, we need to be able to read, write, copy, and edit that language. If you were using a word processor to find one line in a hundred-page document or one article from one book out of the Library of Congress, you would also need a way to search the large print base available. You might want to compare two different copies of files to see if they differ from each other. From the lab and this online discussion and problem set, you will learn how modern scientists read, write, copy, edit, search, and compare the language of the genome. These abilities, acquired over the last twenty years, have revolutionized our understanding of life and have given us the potential to alter, for good or evil, life itself.
DNA in human chromosomes exists as one long double-stranded molecule. It is too long to physically study and manipulate in the lab. Using a battery of enzymes, the DNA of chromosomes can be chemically cleaved into smaller fragments, which are more readily manipulable. (Similar techniques are used to sequence proteins, which require overlapping polypeptide fragments to be made.) After the fragments have been made, they must be separated from each other in order to study them. DNA fragments can be separated on the basis of some structural feature that differentiates the fragments from each other. Polarity can not be used since all DNA fragments have negatively charged phosphates in the sugar-phosphate backbone of the molecule. Although each fragment would have a unique sequence, it would be hard to separate all the different fragments, by, for instance, attaching some molecule that binds to a unique sequence in the major groove of a given fragment to a big bead and using that bead to separate out that one unique fragment. You would need a different bead for each unique fragment! The best way to separate the fragments from each other is to base the separation on the actual size of the fragment by using electrophoresis on an agarose or polyacrylamide gel.
A carbohydrate extract called agarose is made from algae. Water is added to the extract, which is then heated. The carbohydrate extract dissolves in the water to form a viscous solution. The agarose solution is poured into a mold (like warm jello) and is allowed to solidify. A plastic comb with wide teeth was placed in the agarose when it was still liquid. When the agarose is solid, the comb can be removed, leaving in its place little wells. A solution of DNA fragments can be placed in the wells. The agarose slab with the sample is covered with a buffer solution and electrodes are placed at each end of the slab. The negative electrode is placed near the well-end of the agarose slab while the positive electrode is placed at the other end. If a voltage is applied across the agarose slab, the negatively charged DNA fragments will move through the agarose gel toward the positive electrode. This migration of charged molecules in solution toward an oppositely charged electrode is called electrophoresis. Pretend you are one of the fragments. To you, the gel looks like a tangled cobweb. You sneak your way through the openings in the web as you move straight forward to the positive electrode. The larger the fragment, the slower you move because it is hard to get through the tangled web. Conversely, the shorter the fragment, the faster you move. Using this technique and its many modifications, oligonucleotides differing by just one nucleotide can be separated from each other. In the electrophoresis of DNA fragments, a fluorescent, uncharged dye, ethidium bromide, is added to the buffer solution. This dye literally intercalates -between the base pairs of DNA, which imparts a fluorescent yellow-green color to the DNA when UV light is shown on the agarose gel.
Reading DNA
We will discuss one method of reading the sequence of DNA. This method, developed by Sanger won him a second Nobel prize. To sequence a single-stranded piece of DNA, the complementary strand is synthesized. Four different reaction mixtures are set up. Each contains all 4 radioactive deoxynucleotides (dATP, dCTP, dGTP, dTTP) required for the reaction and DNA polymerase. In addition, dideoxyATP (ddATP) is added to one reaction tube The dATP and ddATP attach randomly to the growing 3' end of the complementary stranded. If ddATP is added no further nucleotides can be added after since its 3' end has an H and not a OH. That's why they call it dideoxy. The new chain is terminated. If dATP is added, the chain will continue to grow until another A needs to be added. Hence a whole series of discreet fragments of DNA chains will be made, all terminated when ddATP was added. The same scenario occurs for the other 3 tubes, which contain dCTP and ddCTP, dTTP and ddTTP, and dGTP and ddGTP respectively. All the fragments made in each tube will be placed in separate lanes for electrophoresis, where the fragments will separate by size.
Didexoynucleotides
Figure: Didexoynucleotides
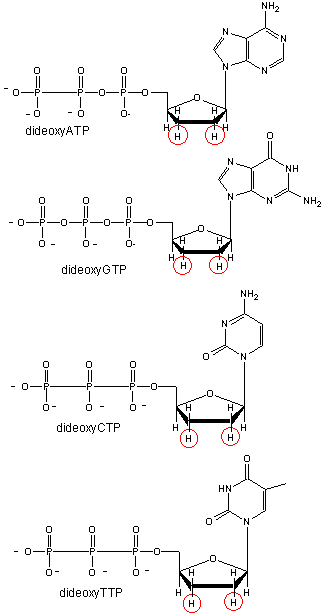
PROBLEM: You will pretend to sequence a single-stranded piece of DNA as shown below. The new nucleotides are added by the enzyme DNA polymerase to the primer, GACT, in the 5' to 3' direction. You will set up 4 reaction tubes, Each tube contains all the dXTP's. In addition, add ddATP to tube 1, ddTTP to tube 2, ddCTP to tube 3, and ddGTP to tube 4. For each separate reaction mixture, determine all the possible sequences made by writing the possible sequences on one of the unfinished complementary sequences below. Cut the completed sequences from the page, determine the size of the polynucleotide sequences made, and place them as they would migrate (based on size) in the appropriate lane of an imaginary gel, which you have drawn on a piece of paper. Lane 1 will contain the nucleotides made in tube 1, etc. Then draw lines under the positions of the cutout nucleotides to represent DNA bands in the gel. Read the sequence of the complementary DNA synthesized. Then write the sequence of the ssDNA that was to be sequenced.
5' T C A A C G A T C T G A 3' (STAND TO SEQUENCE)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
3' G A C T 5' (primer)
Since the DNA fragments have no detectable color, they can not be directly visualized in the gel. Alternative methods are used. In the one described above, radiolabeled ddXTP's were used. Once the sequencing gel is run, it can be dried and the bands visualized by radioautography (also called autoradiography). A place of x-ray film is placed over the dried gel in a dark environment. The radiolabeled bands will emit radiation which will expose the x-ray film directly over the bands. The film can be developed to detect the bands. In a newer technique, the primer can be labeled with a fluorescent dye. If a different dye is used for each reaction mixture, all the reaction mixtures can be run in one lane of a gel. (Actually, only one reaction mix containing all the ddXTP's together is performed.) The gel can then be scanned by a laser, which detects fluorescence from the dyes, each at a different wavelength.
Figure: DNA sequencing using different fluorescent primers for each ddXTP reaction
One recent advance in sequencing allows for real-time determination of a sequence. The four deoxynucleotides are each labeled with a different fluorophore on the 5' phosphate (not the base as above). A tethered DNA polymerase elongates the DNA on a template, releasing the fluorophore into solution (i.e. the fluorophore is not incorporated into the DNA chain). The reaction takes place in a visualization chamber called a zero mode waveguide which is a cylindrical metallic chamber with a width of 70 nm and a volume of 20 zeptoliters (20 x 10-21 L). It sits on a glass support through which laser illumination of the sample is achieved. Given the small volume, non-incorporated fluorescently tagged deoxynucleotides diffuse in and out in the microsecond timescale. When a deoxynucleotide is incorporated into the DNA, its residence time is in the millisecond time scale. This allows for prolonged detection of fluorescence which gives a high signal-to-noise ratio. Newer technology in which sequence is done by moving DNA through pores in membranes could bring sequencing down to $1000/genome or less.
Writing DNA
Oligonucleotides can be synthesized on a solid bead. By adding one nucleotide at a time, the sequence and length of the oligonucleotide can be controlled.
Copying DNA
Several methods exist for copying a sequence of DNA millions of times. Most methods make use of plasmids (which are found in bacteria) and viruses (which can infect any cell). The DNA of the plasmid or virus is engineered to contain a copy of a specific DNA sequence of interest. The plasmid or virus is then reintroduced into the cell where amplification occurs.
Initially, a DNA containing a gene or regulatory sequence of interest is cut at specific places with an enzyme called a restriction endonuclease, or restriction enzyme for short. The enzyme doesn't cleave DNA anywhere, but rather at "restricted" places in the sequence, much as an endoprotease cleaves a protein after a given amino acid within a protein chain. Instead of cleaving one strand, as in proteins, the restriction endonuclease must cleave both strands of dsDNA. It can cut the strands cleanly to leave blunt ends, or in a staggered fashion, to leave small tails of ssDNA. Multiple such sites exist at random in the genome. The gene of interest must be flanked on either side by such a sequence. The same enzyme is used to cleave the plasmid or virus DNA.
Figure: Cleaving DNA with the Restriction Enzyme EcoR1
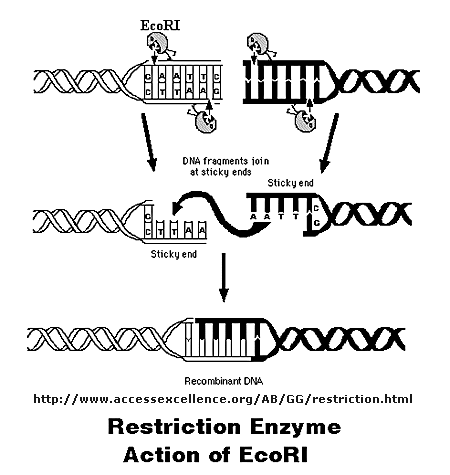
The foreign fragment of DNA can then be added to the plasmid or viral DNA as shown to make a recombinant DNA molecule. This technique of DNA cloning is the basis for the entire field of recombinant DNA technology.
Figure: Cloning a Restriction Fragment into a Plasmid

The plasmid can be added to bacteria, which take it up in a process called transformation. The plasmid can be replicated in the bacteria which will copy the DNA fragment of interest. Typically the plasmid carries a gene that can make the bacteria resistant to an antibiotic. Only bacteria that carry the plasmid (and presumably the insert) will grow. To isolate the desired fragment, the plasmids are isolated from bacteria, and cleaved with the same restriction enzyme to remove the desired fragment, after which it can be purified. In addition, the bacteria can be induced to express the protein from the foreign gene. In lab 4, we will transform bacteria with a plasmid containing the gene for human adipoctye acid phosphatase beta, HAAP-B, and induce expression of the gene.
A similar method can be used to copy DNA in which the foreign fragment is recombined with the DNA of bacteriophage, a virus, which infects bacteria like E. Coli. The recombinant DNA can be packaged into actual viruses, as shown below. When the virus infects the bacteria, it instructs the cells to make millions of new viruses, hence copying the foreign fragment of interest.
Sometimes, "cloning" or copying a fragment of DNA is not what an investigator really wants. If the genomic DNA comes from a human cell, for instance, the gene will contain introns. If you put this DNA into a plasmid or bacteriophage, the introns go with it. Bacteria can replicate this DNA, but often one wants not to just copy (amplify) the DNA but also transcribe it into RNA and then translate it into protein. Bacteria, however, can not splice out the intron RNA, so mature mRNA can not be made. If one could clone into the bacteria's DNA without the introns, this problem would not exist. One such possible method exists in which you start with the actual mRNA for a protein of interest. In this technique, a dsDNA copy is made from a ss-mRNA molecule. Such dsDNA is called cDNA, for complementary or copy DNA. This can then be cloned into a plasmid or bacteriophage vector and amplified as described above.
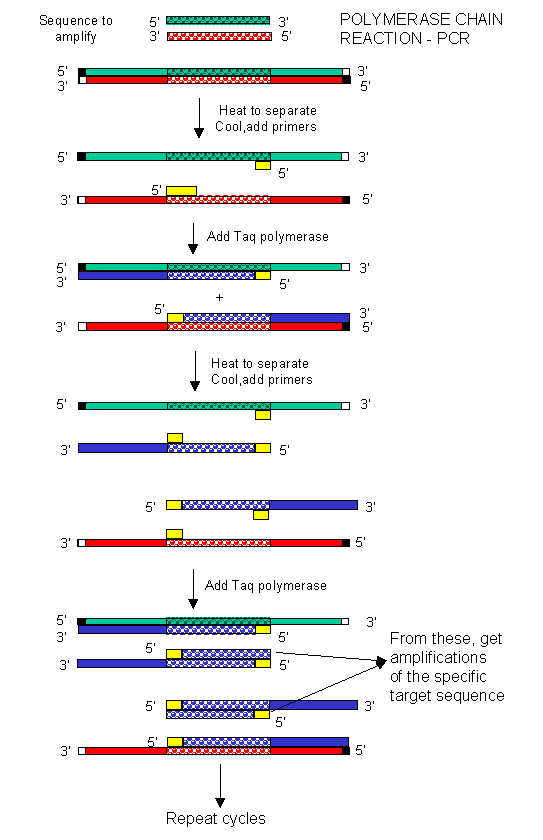
In the mid '80s a new method was developed to copy (amplify) DNA in a test tube. It doesn't require a plasmid or a virus. It just requires a DNA fragment, some primers (small oligonucleotides complementary to sections of DNA on each strand and straddling the section of DNA to be amplified. Just add to this mixture dATP, dCTP, dGTP, dTTP, and a heat-stable DNA polymerase from the organism Thermophilus aquaticus (which lives in hot springs), and off you go. The mixture is first heated to a temperature, which causes the DsDNA strands to separate. The temperature is cooled allowing a large stoichiometric excess of the primers to anneal to the ssDNA. The heat-stable Taq polymerase (from Thermophilus aquaticus) polymerizes DNA from the primers. The temperature is raised again, allowing dsDNA strand separation. On cooling the primers anneal again to the original and newly synthesized DNA from the last cycle and synthesis of DNA occurs again. This cycle is repeated as shown in the diagram. This chain reaction is called the polymerase chain reaction (PCR). The target DNA synthesized is amplified a million times in 20 cycles, or a billion times in 30 cycles, which can be done in a few hours.
Editing DNA
We will spend much time discussing how specific amino acids could be covalently modified to either identify the presence of a specific amino acid or to modify the activity of the protein. Now it is routine to use recombinant DNA technology, to alter one or more nucleotides, to either change the amino acid or add or delete one or more amino acids. This technique, called site-specific mutagenesis, is used extensively by protein chemists to determine the importance of a given amino acid in the folding, structure, and activity of a protein. The techniques are described in the diagram below;

Searching DNA
Where on a chromosome is the gene that codes for a given protein? One way to find the gene is to synthesize a small oligonucleotide "probe" which is complementary to part of the actual DNA sequence of the gene (determined from previous experiments). Attach a fluorescent molecule to the DNA probe. Then take a cell preparation in which the chromosomes can be seen under the microscope. Base is added which unwinds the double-stranded DNA helix. A fluorescent probe is added that will bind to the chromosome at the site of the gene to which the DNA is complementary. Hybridization is the process whereby a single-stranded nucleotide sequence (the target) binds through H-bonds to another complementary nucleotide sequence (the probe).
What if you don't know the nucleotide sequence of the gene, but you know the amino acid sequence of the protein, as in the example shown below? From the genetic code table, you could predict the possible sequence of all possible RNA molecules that are complementary to the DNA in the gene. Since some of the amino acids have more than one codon, there are many possible sequences of DNA that could code for the protein fragment. The link below shows all possible corresponding mRNA sequences that could code for a short amino acid sequence. The 20 mer sequence of minimal degeneracy in the nucleotide sequence should be used as genomic probes.
Comparing DNA
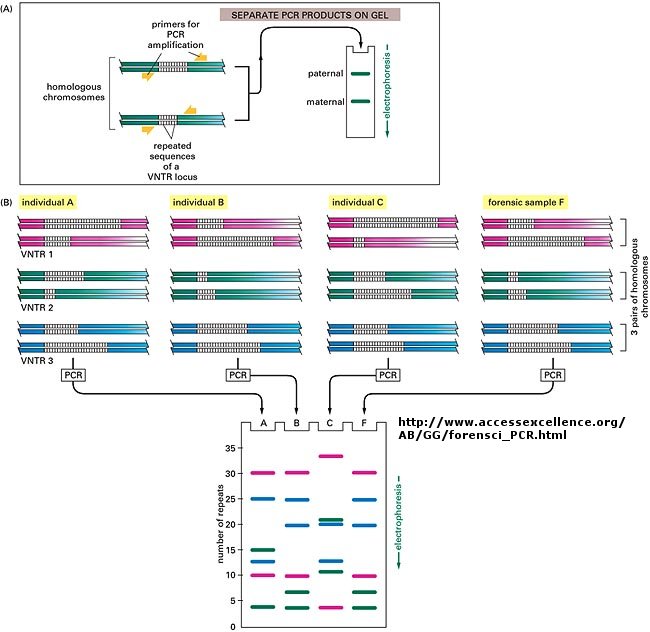
The DNA sequence of each individual must be different from every other individual in the world (with the exception of identical twins). The difference must be less than the differences between a human and a chimp, which are 98.5 % identical. Let us say that each of us have DNA sequences that are 99.9 % identical as compared to some "a normal humans". Given that we have about 4 billion base pairs of DNA, that means we are all different in about 0.001 x 4,000,000,000 which is about 4 million base pairs different. This means that on average we have one nucleotide difference for each 1000 base pairs of DNA. Some of these are in genes, but most are probably in between DNA, and many have been shown to be clustered in areas of highly repetitive DNA at the ends of chromosomes (called the telomeres) and in the middle (called the centromeres).
Now, remember that there are restriction enzyme sites interspersed randomly along the DNA as well. If some of the differences in the DNA among individuals occur within the sequences where the DNA is cleaved by restriction enzymes, then in some individuals a particular enzyme won't cleave at the usual site, but at a more distal site. Hence, the size of the restriction enzyme fragments should differ for each person. Each person's DNA, when cut by a battery of restriction enzymes, should give rise to a unique set of DNA fragments of sizes unique to that individual. Each person's DNA has a unique Restriction Fragment Length Polymorphism (RFLP). How could you detect such polymorphism?
You already know how to cut sample DNA with restriction enzymes, and then separate the fragments on an agarose gel. An additional step is required, however, since thousands of fragments could appear on the gel, which would be observed as one large continuous smear. If however, each fragment could be reacted with a set of small, radioactive DNA probes which are complementary to certain highly polymorphic sections of DNA (like telomeric DNA) and then visualized, only a few sets of discrete bands would be observed in the agarose gel. These discrete bands would be different from the DNA bands seen in another individual's gene treated the same way. This technique is called Southern Blotting and works as shown below. DNA fragments are electrophoresed in an agarose gel. The ds DNA fragments are unwound by heating, and then a piece of nitrocellulose filter paper is placed on top of the gel. The DNA from the gel transfers to the filter paper. Then a small radioactive oligonucleotide probe, complementary to a polymorphic site on the DNA, is added to the paper. It binds only to the fragment containing DNA complementary to the probe. The filter paper is dried, and a piece of x-ray film is placed over the sheet. A set of radioactive fragments (which are not complementary to the probe) are run as well. They serve as a set of markers to ensure that the gel electrophoresis and transfer to the filter paper were correct. This technique is shown on the next page, along with a RFLP analysis from a particular family.
When this technique is used in forensic cases or in paternity cases, it is called DNA fingerprinting. With present techniques, investigators can state unequivocally that the odds of a particular pattern not belonging to a suspect are in the range of one million to one. The x-ray film shown below is a copy of real forensic evidence obtained from a rape case. Shown are the Southern blot results from suspect 1, suspect 2, the victim, and the forensic evidence. Analyze the data.