8.4: Chromosomes and Chromatin
- Page ID
- 14964
Chromatin
When stained and viewed in a microscope, eukaryotic nuclear DNA in nondividing cells is observed in two different states, heterochromatin (dark areas) and euchromatin (light areas), as shown in Figure \(\PageIndex{1}\).
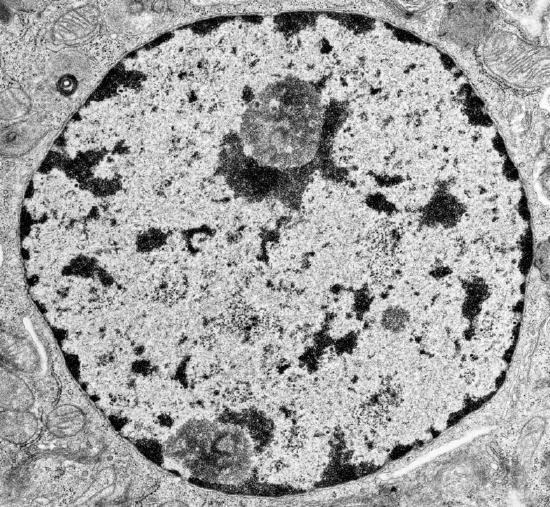
The heterochromatin is darkly stained and found along the inner side of the nuclear envelope as well as inside the nucleus. Euchromatin doesn't stain well since it is more dispersed, and is found throughout the nucleus. The image above is of a cell in interphase, a part of the cell cycle in which the cell is in-between cell divisions. Here are some differences between heterochromatin and euchromatin:
Heterochromatin:
- contains genes that are transcriptionally inactive and not expressed;
- is not found in prokaryotic cells;
- in humans, one of the X chromosomes is silenced in heterochromatin while the other is transcribable and in euchromatin;
- protects the tightly packed in it from nucleases.
Euchromatin:
- contains about 90% of genome DNA;
- contains chromosomes that are unfolded into nucleosomal DNA resembling beads on a string and is characterized by looser association with packaging histone proteins;
- is the only form of DNA in bacteria;
- is transcriptionally active so it is open to the binding of RNA polymerase and transcription factors;
- has some genes which can be transcriptionally silenced by moving them into heterochromatin.
Look as long as you wish but you won't see the iconic pictures of chromosomes in Figure \(\PageIndex{1}\). They are there but dispersed. They are only discretely visible at certain times in the cell cycle when cells are preparing to divide.
For those with a more chemistry-centric background, a short review of the cell cycle would be helpful. Somatic (non-germ cells) spend only part of their lives preparing for and dividing into two cells in a process called mitosis. Figure \(\PageIndex{2}\) shows a simple model for the cell cycle.
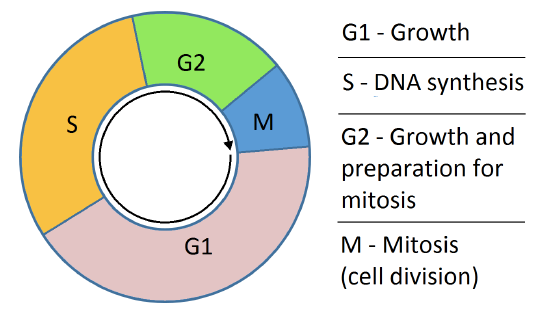
Before a cell gets ready to divide, it is in interphase (between mitotic phases), which encompasses the G1, S, and G2 phases of the cell cycle. In interphase, cells grow and replicate their DNA Mitosis or cell division occurs after G2 and consists of a series of new discrete phases, including prophase, metaphase, anaphase, and telophase, after which the cell divides in a process called cytokinesis. Table \(\PageIndex{1}\) below shows the stages of mitosis and cytokinesis which occurs after interphase.
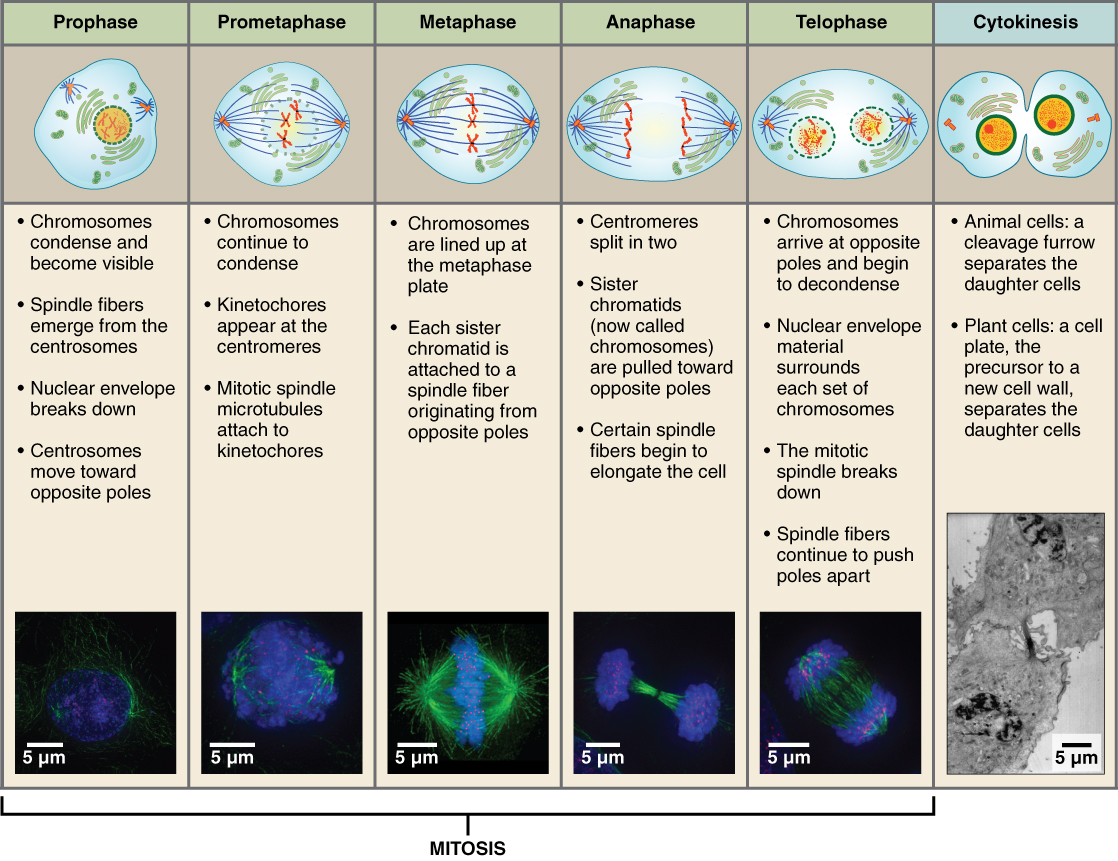
Let's start with a discussion of the structure of chromosomes as classically observed in mitotic cells. Then we will look more closely at the seemingly amorphous and more complicated structures of chromatin.
Chromosomes
Within eukaryotic cells, DNA is organized into long linear structures called chromosomes. A chromosome is a "thread-like" structure in the nucleus of animal and plant that consists of a single but long molecule of double-stranded DNA (so it's two ss-DNAs) with a myriad of bound proteins. The proteins bind to and condense the DNA molecule to prevent it from becoming an unmanageable tangle. Before typical cell division (mitosis), these chromosomes are replicated by DNA polymerase to make two identical chromosomes, one for each future daughter cell. The two identical chromosomes, called sister chromatids, bind to each other at a common structure called the centromere. A replicated chromosome (sister chromatids) bound to each other at the centromere, is shown in Figure \(\PageIndex{3}\). Each chromosome in the sister chromatid structure represents one chromatid.

In humans, each cell normally contains 23 pairs of chromosomes, for a total of 46. In twenty-two of these pairs, called autosomes, each member of the pair is similar. The 23rd pair, the sex chromosomes, differ between males and females. Females have two copies of the X chromosome, while males have one X and one Y chromosome. Figure \(\PageIndex{4}\) shows a DNA karyotype with 22 pairs of autosomes and one pair of sex chromosomes. Karyotypes are prepared using standardized staining procedures that reveal characteristic structural features for each chromosome, usually from white blood cells. The karyotype of human cells shown in Figure 4 below shows an extra copy of chromosome 21 (trisomy 21), which causes Down's Syndrome.
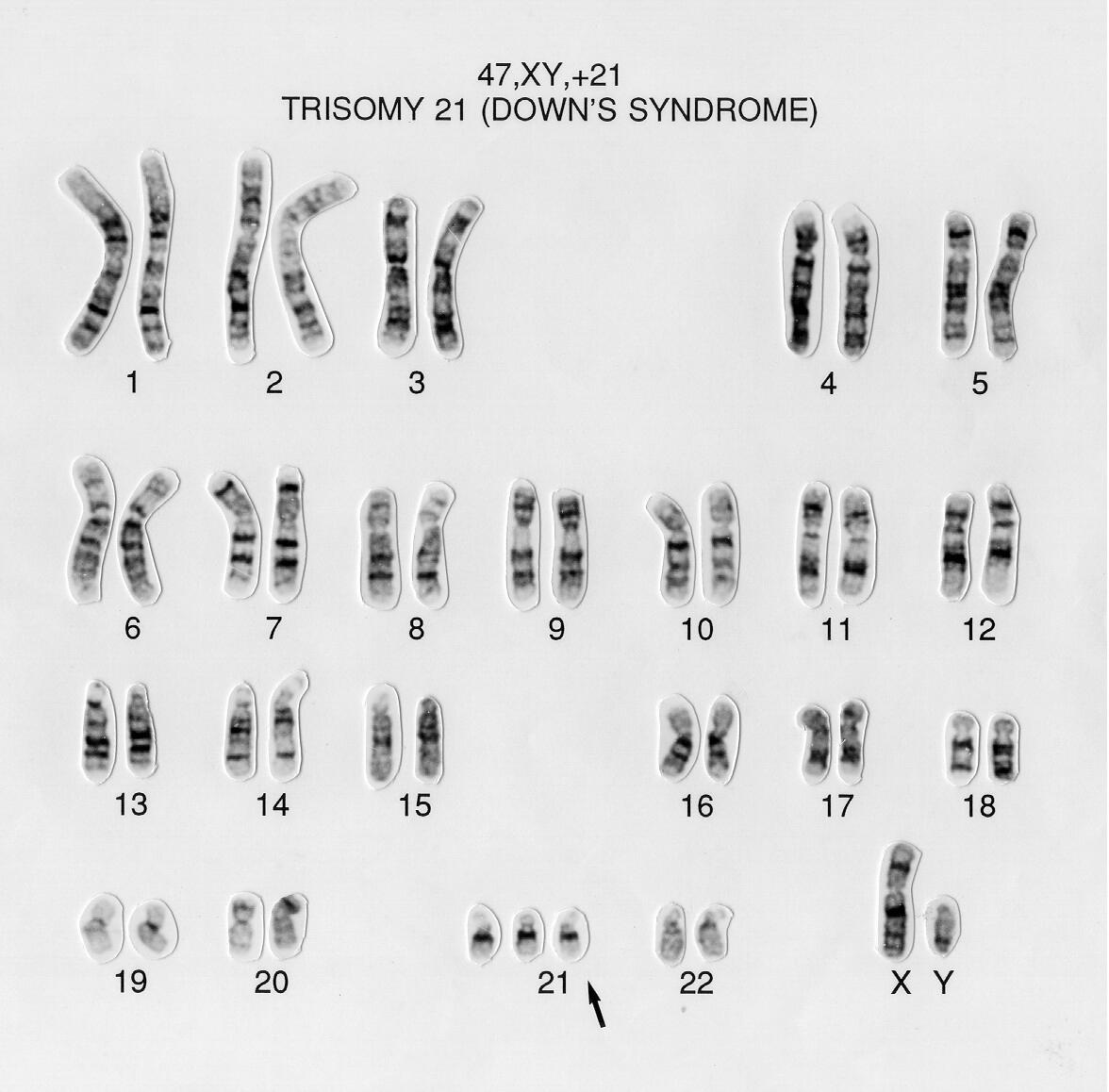
Each species has a unique chromosomal complement. For example, chickens have 39 pairs of chromosomes (78 total) with 38 autosomal pairs and one pair of sex chromosomes (Z and W). In this species, ZW chickens are female and ZZ chickens are male. The total length of the human genome is over 3 billion base pairs. The total length of the human genome is over 3 billion base pairs. The genome also includes mitochondrial DNA.
Eukaryotic organisms (animals, plants, fungi, and protists) store most of their DNA inside the cell nucleus as linear nuclear DNA. However, DNA in the mitochondria and chloroplast are circular (with no ends). Bacteria and Archaea cells do not have organelle structures and thus, store their DNA only in a region of the cytoplasm known as the nucleoid region. Prokaryotic chromosomes consist of double–stranded circular DNA.
Packing of DNA: Supercoiling
The genome of a cell is often significantly larger than the cell itself. For example, if the DNA from a human cell containing 46 chromosomes were stretched out in a line, it would extend more than 6 feet (2 meters)! How is it possible that the genetic information not only fits into the cell but fits into the cell nucleus? Eukaryotes solve this problem by a combination of supercoiling and packaging DNA around the histone family of proteins (described below). Prokaryotes do not contain histones (with a few exceptions). Prokaryotes tend to compress their DNA using nucleoid-associated-proteins (NAPs) and supercoiling, as shown in Figure \(\PageIndex{5}\).
DNA supercoiling refers to the over- or under-winding of a DNA strand and is an expression of the strain on that strand. Supercoiling is important in a number of biological processes, such as compacting DNA and regulating access to the genetic code. DNA supercoiling strongly affects DNA metabolism and possibly gene expression. Additionally, certain enzymes such as topoisomerases can change DNA topology to facilitate functions such as DNA replication or transcription.
In a “relaxed” double-helical segment of B-DNA, the two strands twist around the helical axis once every 10.4–10.5 base pairs of sequence. Adding or subtracting twists, as some enzymes can do, impose strain. If a DNA segment under twist strain were closed into a circle by joining its two ends and then allowed to move freely, the circular DNA would contort into a new shape, such as a simple figure-eight, as shown in Figure \(\PageIndex{5}\). Such a contortion is a supercoil. The noun form “supercoil” is often used in the context of DNA topology.
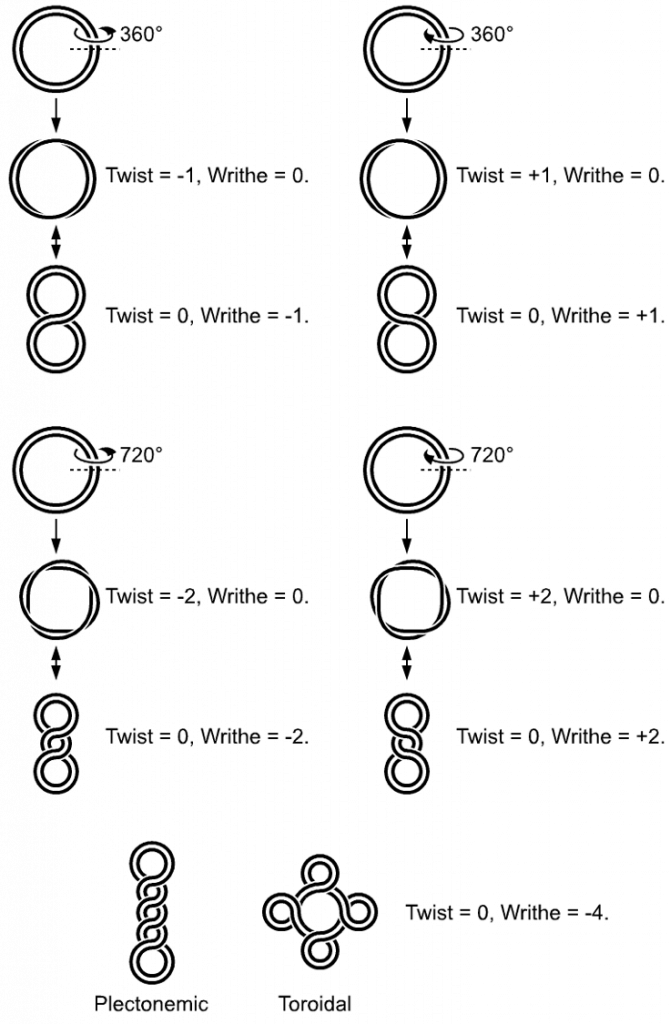
Positively supercoiled (overwound) DNA is transiently generated during DNA replication and transcription, and, if not promptly relaxed, inhibits (regulates) these processes. The simple figure eight is the simplest supercoil and is the shape a circular DNA assumes to accommodate one too many or one too few helical twists. The two lobes of the figure eight will appear rotated either clockwise or counterclockwise with respect to one another, depending on whether the helix is over- or underwound. For each additional helical twist being accommodated, the lobes will show one more rotation about their axis. As a general rule, the DNA of most organisms is negatively supercoiled.
Lobal contortions of a circular DNA, such as the rotation of the figure-eight lobes above, are referred to as writhe. The above example illustrates that twisting and writhing are interconvertible. Supercoiling can be represented mathematically by the sum of twist and writhe, The twist is the number of helical turns in the DNA and the writhe is the number of times the double helix crosses over on itself (these are the supercoils). Extra helical twists are positive and lead to positive supercoiling, while subtractive twisting causes negative supercoiling. Many topoisomerase enzymes sense supercoiling and either generate or dissipate it as they change DNA topology.
In part, because chromosomes may be very large, segments in the middle may act as if their ends are anchored. As a result, they may be unable to distribute excess twist to the rest of the chromosome or to absorb twist to recover from underwinding—the segments may become supercoiled, in other words. In response to supercoiling, they will assume an amount of writhe, just as if their ends were joined.
Supercoiled circular DNA forms two major structures; a plectoneme or a toroid, or a combination of both. A negatively supercoiled DNA molecule will produce either a one-start left-handed helix, the toroid, or a two-start right-handed helix with terminal loops, the plectoneme. Plectonemes are typically more common in nature, and this is the shape most bacterial plasmids will take. For larger molecules, it is common for hybrid structures to form – a loop on a toroid can extend into a plectoneme as shown in Figure \(\PageIndex{6}\). DNA supercoiling is important for DNA packaging within all cells and seems to also play a role in gene expression.
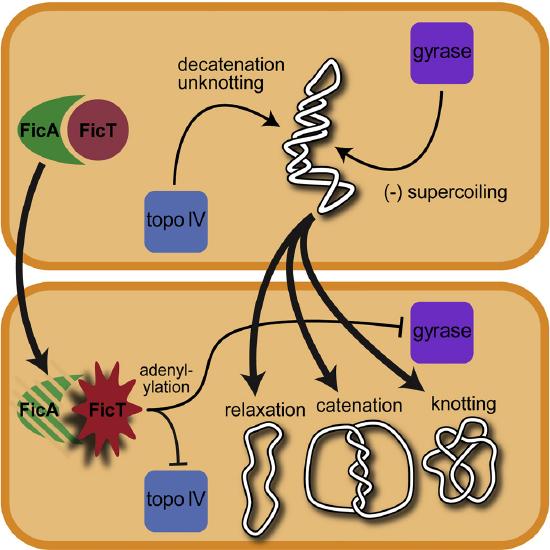
In addition to forming supercoiled structures, circular chromosomes from bacteria have been shown to undergo the processes of catenation and knotting upon the inhibition of topoisomerase enzymes. Catenation is the process by which two circular DNA strands are linked together like chain links, whereas DNA knotting is the interlooping structures occurring within a single circular DNA structure. These are illustrated in Figure \(\PageIndex{7}\). In vivo, the action of topoisomerase enzymes is critical to keep knots and catenoids from tangling the DNA structure.
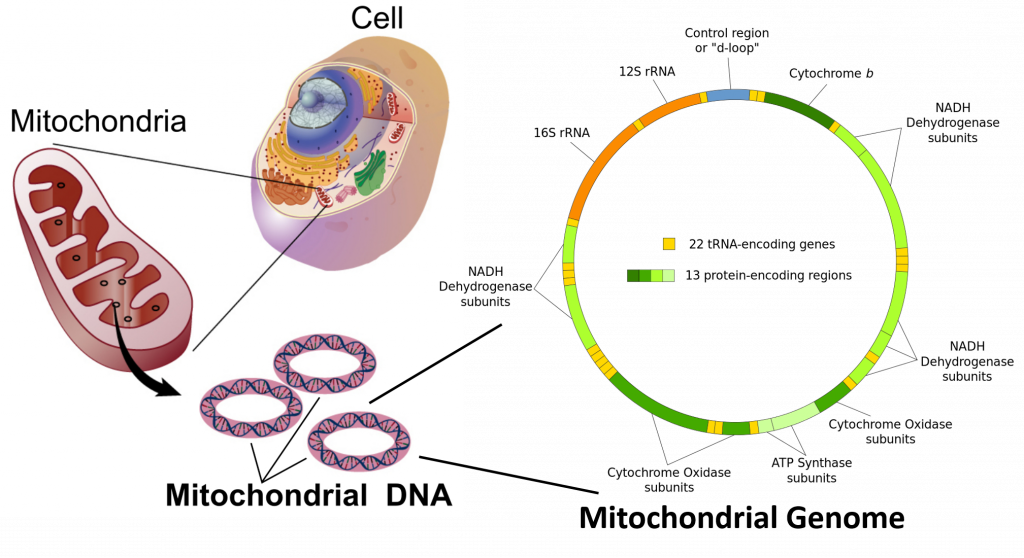
Mitochondrial and Chloroplast DNA are circular suggesting a bacterial origin for both of these organelle structures. Sequence alignments further lend support for the endosymbiotic theory, which proposes that bacteria were engulfed by early eukaryotic organisms and subsequently became symbiotic to their eukaryotic counterpart, rather than being digested.
In the cells of eukaryotic organisms, the vast majority of the proteins present in the mitochondria (numbering approximately 1500 different types in mammals) are coded for by nuclear DNA. However, sequencing of the human mitochondrial genome has revealed 16,569 base pairs encoding 13 proteins, as shown in Figure \(\PageIndex{8}\). Many of the mitochondrially produced proteins are required for electron transport during the production of ATP.
Histones and Nucleosomes
Within eukaryotic chromosomes, chromatin proteins, known as histones, compact and organize DNA. These compacting structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed.
Histones are highly basic proteins found in eukaryotic cell nuclei that package and order the DNA into structural units called nucleosomes. They are the chief protein components of chromatin, acting as spools around which DNA winds, and playing a role in gene regulation. Without histones, the unwound DNA in chromosomes would be very long (a length-to-width ratio of more than 10 million to 1 in human DNA). For example, each human diploid cell (containing 23 pairs of chromosomes) has about 1.8 meters of DNA wound on the histones, and the diploid cell has about 90 micrometers (0.09 mm) of chromatin.
There are five major families of histones, H1/H5, H2A, H2B, H3, and H4. Histones H2A, H2B, H3, and H4 are known as the core histones, while histones H1/H5 are known as the linker histones.
The core histones all exist as dimers, which are similar in that they all possess the histone fold domain: three alpha helices linked by two loops (Figure 4.13). It is this helical structure that allows for interaction between distinct dimers, particularly in a head-tail fashion (also called the handshake motif). The resulting four distinct dimers then come together to form one octameric nucleosome core, approximately 63 Angstroms in diameter. Around 146 base pairs (bp) of DNA wrap around this core particle 1.65 times in a left-handed super-helical turn to give a particle of around 100 Angstroms across, called a nucleosome as illustrated in Figure \(\PageIndex{9}\).
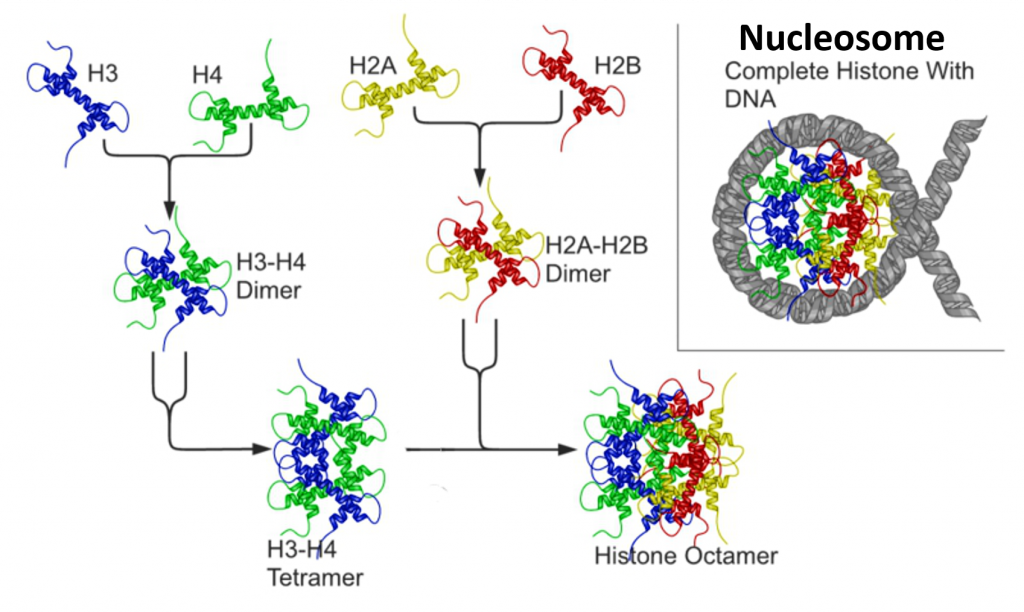
The linker histone H1 binds the nucleosome at the entry and exit sites of the DNA, thus locking the DNA into place and allowing the formation of higher order structure (Figure 4.14). The most basic such formation is the 10 nm fiber or beads on a string conformation. This involves the wrapping of DNA around nucleosomes with approximately 50 base pairs of DNA separating each pair of nucleosomes (also referred to as linker DNA).
The nucleosome contains over 120 direct protein-DNA interactions and several hundred water-mediated ones. Direct protein – DNA interactions are not spread evenly about the octamer surface but rather located at discrete sites. These are due to the formation of two types of DNA binding sites within the octamer; the α1α1 site, which uses the α1 helix from two adjacent histones, and the L1L2 site formed by the L1 and L2 loops. Salt links and hydrogen bonding between both side-chain basic and hydroxyl groups and main-chain amides with the DNA backbone phosphates form the bulk of interactions with the DNA. This is important, given that the ubiquitous distribution of nucleosomes along genomes requires it to be a non-sequence-specific DNA-binding factor. Although nucleosomes tend to prefer some DNA sequences over others, they are capable of binding practically to any sequence, which is thought to be due to the flexibility in the formation of these water-mediated interactions. In addition, non-polar interactions are made between protein side-chains and the deoxyribose groups, and an arginine side-chain intercalates into the DNA minor groove at all 14 sites where it faces the octamer surface. The spatial distribution and strength of DNA-binding sites about the octamer surface distort the DNA within the nucleosome core. The DNA is non-uniformly bent and also contains twist defects. The twist of free B-form DNA in solution is 10.5 bp per turn. However, the overall twist of nucleosomal DNA is only 10.2 bp per turn, varying from a value of 9.4 to 10.9 bp per turn.
The histone tail extensions constitute up to 30% by mass of histones, but are not visible in the crystal structures of nucleosomes due to their high intrinsic flexibility, and have been thought to be largely unstructured (Figure 4.14). The N-terminal tails of histones H3 and H2B pass through a channel formed by the minor grooves of the two DNA strands, protruding from the DNA every 20 bp. The N-terminal tail of histone H4, on the other hand, has a region of highly basic amino acids (16-25), which, in the crystal structure, forms an interaction with the highly acidic surface region of an H2A-H2B dimer of another nucleosome, being potentially relevant for the higher-order structure of nucleosomes. This interaction is thought to occur under physiological conditions also and suggests that acetylation of the H4 tail distorts the higher-order structure of chromatin.
Figure \(\PageIndex{10}\) shows an interactive iCn3D model of the human nucleosome (3afa). One member of each pair of histones is shown in cartoon rendering, while the other member of the pair is shown in the same color but in spacefill rendering. The structure of a human nucleosome (3afa) is shown below (H2A is shown in cyan, H2B in blue, H3 in magenta, and H4 in purple). Each strand of DNA is shown in a different shade of gray.
Figure \(\PageIndex{10}\): Human nucleosome (3afa). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...B2SwQHYDLj4BJ6
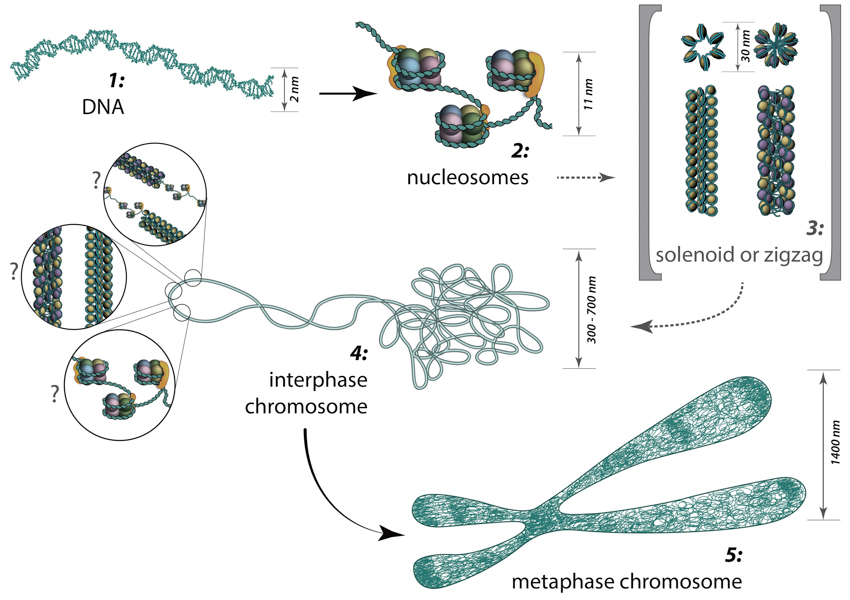
The packing of DNA from dsDNA to the metaphase chromosomes is schematically shown in Figure \(\PageIndex{11}\). The formation of the DNA double helix represents the first-order packaging of the chromosome structure. The formation of nucleosomes represents the second level of packaging for eukaryotic chromosomes. In vitro data suggests that nucleosomes are then arranged into either a solenoid structure which consists of 6 nucleosomes linked together by the Histone H1 linker proteins or a zigzag structure that is similar to the solenoid construct. Both the solenoid and zigzag structures are approximately 30 nm in diameter. The solenoid and zigzag structures reported from in vitro data have not yet been confirmed to occur in vivo.
Telomeres
At the ends of the linear eukaryotic chromosomes are specialized regions of DNA called telomeres. The main function of these regions is to allow the cell to replicate chromosome ends using the enzyme telomerase, as the enzymes that normally replicate DNA cannot copy the extreme 3′ ends of chromosomes. A cartoon showing telomeres and their extension is shown in Figure \(\PageIndex{12}\).
.jpg?revision=1)
These specialized chromosome caps also help protect the DNA ends, and stop the DNA repair systems in the cell from treating them as damage to be corrected.
In human cells, telomeres contain 300-8000 repeats of a simple TTAGGG sequence. The repetitive TTAGGG sequences in telomeric DNA can form unique higher-order structures called quadruplexes. Figure \(\PageIndex{13}\) shows an interactive iCn3D model of parallel quadruplexes from human telomeric DNA (1KF1). The structure contains a single DNA strand (5'-AGGGTTAGGGTTAGGGTTAGGG-3') which contains four TTAGGG repeats.
Figure \(\PageIndex{13}\): A buried phenylalanine in low molecular weight protein tyrosyl phosphatase (1xww) (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...y5joFHDgWJQsQ6
Rotate the model to see 3 parallel layers of quadruplexes. In each layer, 4 noncontiguous guanine bases interact with a K+ ion. Hover over the guanine bases in one layer and you will find that one layer consists of guanines 4, 10, 16, and 22, which derive from the last G in each of the repeats in the sequence of the oligomer used (5'-AGGGTTAGGGTTAGGGTTAGGG-3'). These quadruplexes certainly serve as recognition and binding sites for telomerase proteins. The guanine-rich telomere sequences which can form quadruplex may also function to stabilize chromosome ends
During DNA replication, the double-stranded DNA is unwound and DNA polymerase synthesizes new strands. However, as DNA polymerase moves in a unidirectional manner (from 5’ to 3’), only the leading strand can be replicated continuously. For the complementary lagging strand, DNA replication is discontinuous. In humans, small RNA primers attach to the lagging strand DNA, and the DNA is synthesized in small 5'-3' stretches of about 100-200 nucleotides, which are termed Okazaki fragments. The RNA primers are removed and replaced with DNA and the Okazaki DNA fragments are ligated together. At the end of the lagging strand, it is impossible to attach an RNA primer, meaning that there will be a small amount of DNA lost each time the cell divides. This ‘end replication problem’ has serious consequences for the cell as it means the DNA sequence cannot be replicated correctly, with the loss of genetic information. Hence most telomeres have 3´ overhangs. Bacteria DNA, which is circular, does not have the problem.
To prevent this, telomeres are repeated hundreds to thousands of times at the end of the chromosomes. Each time cell division occurs, a small section of telomeric sequences is lost to the end replication problem, thereby protecting the genetic information. At some point, the telomeres become critically short. This decreases leads to cell senescence, where the cell is unable to divide, or apoptotic cell death. Telomeres are the basis for the Hayflick limit, the number of times a cell can divide before reaching senescence.
Telomeres can be restored by the enzyme telomerase, which extends the telomere's length. Telomerase activity is found in cells that undergo regular division, such as stem cells and lymphocyte cells of the immune system. The enzyme has two major subunits. One is the catalytic enzyme named telomerase reverse transcriptase (TERT), which is an RNA-dependent DNA polymerase. Almost all DNA polymerases use DNA as a template for replication. Some RNA viruses that use RNA as their genetic information like HIV encode their own reverse transcriptase, which directs the polymerization of a DNA copy of the viral RNA genome as part of its life cycle. )The virus that causes the Covid-19 pandemic is called severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). It is also an RNA virus but in contrast to HIV it uses and encodes an RNA-dependent RNA polymerase.)
The second subunit of telomerase is the telomerase RNA (TR), which contains a template from which the new telomer is made. The enzyme makes many DNA copies from this to create a multitude of DNA repeat in the telomeres. Figure \(\PageIndex{14}\) shows the structure of the telomerase RNA used to build new telomers. Ovals show proteins that form part of the complex. 'For this discussion, the most important is TERT, telomerase reverse transcriptase, the RNA-dependent DNA polymerase which synthesizes new telomeric DNA from the template sequence of the RNA.
Trace the black single-stranded telomeric DNA strand from the 5' end (bottom left) to its 3' end containing the terminal sequence 5'GCTGG3'. Now start tracing the telomerase RNA starting with its 5' end, which is between the CEH and SM binding site in Figure \(\PageIndex{14}\). It bends sharply near TBE, and continues to the right through the Ku binding site, where it forms stem-loops. It then continues into the template region containing a template sequence (3'CACCGACC5'), from which new telomeric DNA is made (5'GTGGCTGG3' - a slightly different repeat than the human repeat), and an annealing sequence (3'CGACC5'), which is complementary to the 3' end of the existing telomeric DNA (5'GCTGG3'). The RNA continues in the 3' direction through the Est1 bind site region, through the pseudoknot, and ends at the 3' end past the SM binding site.
Telomeres can also be extended through the Alternative Lengthening of Telomeres (ALT) pathway. In this case, rather than being extended, telomeres are switched between chromosomes by homologous recombination. As a result of the telomere swap, one set of daughter cells will have shorter telomeres, and the other set will have longer telomeres
A downside to telomere extension is the potential for uncontrolled cell division and cancer. Abnormally high telomerase activity has been found in the majority of cancer cells, and non-telomerase tumors often exhibit ALT pathway activation. As well as the potential for losing genetic information, cells with short telomeres are at a higher risk for improper chromosome recombination, which can lead to genetic instability and aneuploidy (an abnormal number of chromosomes).
Chromatin Structure
During interphase, distinct chromosomes as shown in Figure \(\PageIndex{4}\) are not observed. Rather each chromosome occupies a spatially limited, roughly elliptical domain which is known as a chromosome territory (CT). Each chromosome territory is comprised of higher-order chromatin units of ~1 Mb each. These units are likely built up from smaller loop domains that contain the solenoid/zigzag structural motifs. On the other hand, 1Mb domains can themselves serve as smaller units in higher-order chromatin structures. With the development of high-throughput biochemical techniques, such as 3C (chromosome conformation capture) and 4C (chromosome conformation capture-on-chip and circular chromosome conformation capture), numerous spatial interactions between neighboring chromatin territories have been described as shown in Figure \(\PageIndex{15}\).
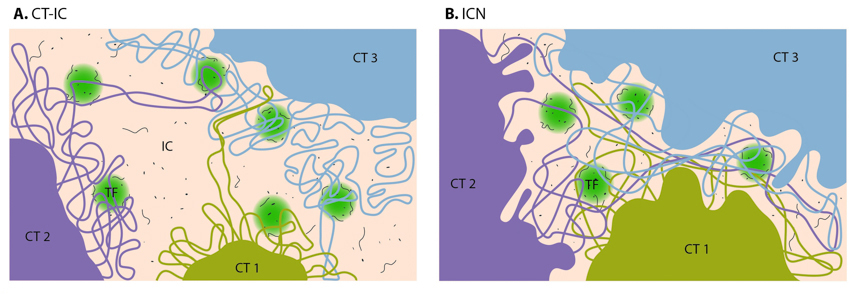
Chromosome territories are known to be arranged radially around the nucleus. This arrangement is both cell and tissue-type specific and is also evolutionarily conserved. The radial organization of chromosome territories was shown to correlate with their gene density and size. In this case, the gene-rich chromosomes occupy interior positions, whereas larger, gene-poor chromosomes, tend to be located around the periphery. Chromosome territories are also dynamic structures, with genes able to relocate from the periphery towards the interior once they have been ‘switched on’. In other cases, genes may move in the opposite direction or simply maintain their position.
Figure \(\PageIndex{16}\) shows nano- to more micro-folding structures of chromosomes in the nucleus. The top left shows the most zoomed-in view where DNA is wound around histone complexes (nucleosomes), which condense to form 10 nm fibers. These condense further into Topologically Associated Domains (TAD) which further separate into Compartments A and B. These then pack into discrete territories. Individual chromosomes occupy their own chromosome territories in the nucleus.
This may seem very complex and it is, but it is somewhat analogous to protein folding, which starts with a linear primary sequence and moves into more complicated secondary structures, secondary structure motifs, domains, tertiary structures, and quaternary structures, which display varying degrees of symmetry.
Because these different types of folding and compaction of chromosomes are difficult to visualize, we will present several different representations of these nano- to microstructures to help your understanding. Figure \(\PageIndex{17}\) offers one that shows scaling factors and differential structures of chromosomes and chromatin.
The large-scale A compartment is gene-rich and actively transcribed so it best represents euchromatin. In contrast, B compartments are gene-poor and best represent heterochromatin. At the subscale level, boundaries between TADs are transcriptionally rich and can be separated from other TADS by heterochromatin "islands".
Another representation of chromatin organization that emphasizes TADs is shown in Figure \(\PageIndex{18}\).
The TAD boundaries in Figure \(\PageIndex{18}\) show a blue CTCF in-between compartments, with each TAD consisting of many interacting loops. A more detailed view is shown in Figure \(\PageIndex{19}\).
A protein complex containing cohesin (which forms a ring) and CTCF is found at sites where loops of relationally transcribable DNA are extruded through the ring. The extruded loops interact with others to form a cluster of regulatory loops containing genes with similar potential for transcription (activated if they end up in Compartment A or repressed if in Compartment B). Cohesin is a functional complex that forms a ring that traps sister chromatids. During anaphase, the complex is cleaved and the sister chromatids separate. CTCF is a chromatin-binding factor with many associated activities.
"Mechanism of chromatin loop formation. TADs contain varying numbers of chromatin loops generated through loop extrusion by CTCF/cohesin
complexes. (Right panel) In the presence of NIPBL and MAU2, the cohesin complex loaded onto the DNA. Then, cohesin extrudes chromatin until a pair of convergent CTCF binding sites is reached. (Right panel) The N-terminus of CTCF and convergent positioning of the CTCF-DNA complex stabilizes cohesin binding and stall chromatin extrusion leading to the establishment of higher-order chromatin organization. The intervening DNA between two convergent CTCF sites leads to the formation of a loop domain, which adopts a variety of complex shapes comprised of multiple regulatory loops. The internal structure of the loop domain is likely determined by polymer chromatin-chromatin self-interactions, which may be further stabilized by phase separation. The contacts within the loop domains facilitate the targeting of enhancers to specific genes (104). The black arrow depicts the direction of loop extrusion."
Special elements in the DNA called enhancers and silencers of gene transcription have long been known to influence gene transcription. These are sequences that are cis (i.e. on the same molecule of DNA, not trans factors like separate proteins that bind to promoters, for example) and can be quite distant from the proximal promoter sequence which controls the transcription of a target gene. How might they work from such a large distance from the promoter? One obvious answer is the DNA folds in 3D space so the enhancers and silencers are close to each in 3D space. Chromatin loop formation facilitates such promoter and enhancer/silencer interactions. Evidence suggests that specific enhancers and silencers are housed in loops in specific TADs so their effects are limited to a subset of genes. Enhancers and promoters seem to interact only within TADs, which suggests that TADS are a fundamental folding "domain" based on gene regulation. If boundaries between different TADS were removed, then promoters and enhancers in one TAD might affect transcription in other TADS, leading to aberrant gene expression. Chromatin loop formation facilitates interactions between promoter and enhancer/silencer elements. Figure \(\PageIndex{20}\) shows how enhancers and silencers can be brought close in space to promoters and their genes through TAD formation.
TADs appear highly conserved in mammals and comprise most (90%) of the genome. The median size is about 880 Kb. The Boundaries between TADs have CCCTC-binding factor (CTCF) and the structural maintenance of chromosomes (SMC) cohesin complex. In Drosophila, TADs are organized by epigenetic state (methylation, condensation). For example, some TADS are transcriptionally active with epigenetic histone modifications (for example trimethylation of histone H3 lysines 4 and 36 or H3Kme3 and H3K36me3) that activate transcription). Others are transcriptionally repressed (enriched in H3K27me3 and containing Polycomb group (PcG) proteins) and some are more classically representative of heterochromatin.
Mechanisms of Separation of Euchromatin and Heterochromatin
From a chemistry focus, what are the interactions which stabilize TADs, compartments, and ultimately heterochromatin and euchromatin? It appears that the easiest way to conceptualize their separation in the nucleus is to use the idea of phase separations. Figure \(\PageIndex{21}\) shows a cartoon representation of the separation of heterochromatin and euchromatin in the zebrafish embryo. The euchromatin is shown as more centrally located and dispersed with red dots indicating transcriptionally active sites. In the late blastula stage, gene expression is dramatically increased and the nucleolus and heterochromatin are not seen.
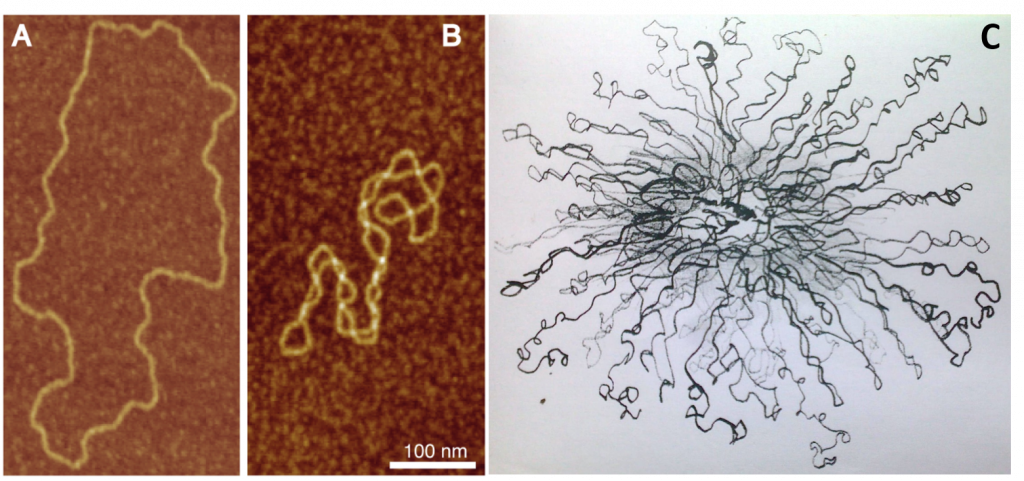
Analyses show that transcription forms regions enriched in RNA, RNA binding proteins, and accordingly transcriptionally active chromatin that are separated from transcriptionally inactive, heterochromatin. Given the dispersion of the DNA required for transcription, the regions enriched in RNA are also depleted in "stainable" DNA. Micrographs showing the depletion of DNA in the actively transcribed regions are shown in Figure \(\PageIndex{22}\). Note the clear lack of DNA in the white rectangles, which contain high levels of RNA and RNA polymerase II (as a proxy for protein).
It appears that the presence of a high concentration of RNA drives "phase" separation of heterochromatin and euchromatin. Hence euchromatin might act as an "oil in water" type microemulsion, with the euchromatin core stabilized by "tethered" RNA acting as an amphiphile. Ribonucleoproteins (RNPs) can be modeled as phase-separated droplets or condensates Figure \(\PageIndex{23}\) shows how increasing RNA leads to the formation of euchromatin "domains" which stay dispersed in the presence of continuing RNA formation.
A final summary cartoon that describes the formation and separation of chromosomal DNA into euchromatin and heterochromatin in Zebra fish is shown in Figure \(\PageIndex{24}\).




.png?revision=1&size=bestfit&width=285)
.png?revision=1&size=bestfit&height=315)