4.7: Fibrillar Proteins
- Page ID
- 62324
Introduction
Most proteins have a roughly spherical or "globular" tertiary structure. However, there are many proteins that form elongated fibrils with properties like elasticity, which allows deformation on the application of a force and subsequent return to the original state. Elastic molecules must store energy (go to a higher energy state) when the elongating force is applied, and the energy must be released on return to the equilibrium resting structure. Structures that can store energy and release it when subjected to a force have resiliency. Proteins that stretch with an applied force include elastin (in blood vessels, lungs and skins where elasticity is required), resilin in insects (which stretches on wing beating), silk (found in spider web and whose structure we showed in 4.2) and fibrillin (found in most connective tissues and cartilage). Some proteins have high resiliency (90% in elastin and resilin), while others are only partially resilient (35% in silk, which has a tensile strength approaching that of stainless steel).
In contrast to rubber, which has an amorphous structure, which imparts elasticity, these proteins, although they have a dissimilar amino acid sequence, seem to have a common structure inferred from their DNA sequences. In some (like fibrillin), the protein has a folded beta sheet domain, which unfolds like a stretched accordion. Others, like elastin and spider silk, have a beta sheet domain and other secondary structures (alpha-helices and beta turns) along with Pro and Ala repetitions. Scientists are studying these structures to help in the synthesis of new elastic and resilient products.
Fibrous Proteins are characterized by elongated protein structures. These types of proteins often aggregate into filaments or bundles forming structural scaffolds in biological systems. Within animals, the two most abundant fibrous protein families are collagen and α-keratin. Let's start our exploration of fibrillar proteins with these.
Collagen
Collagen is the most abundant protein in mammals, making 25% to 35% of the whole-body protein content. It is found predominantly in the extracellular space within various connective tissues in the body. Collagen contains a unique quaternary structure of three protein strands wound together to form a triple helix. It is mostly found in fibrous tissues such as tendons, ligaments, and skin.
Depending upon the degree of mineralization, collagen tissues may be rigid (bone), compliant (tendon), or have a gradient from rigid to compliant (cartilage). It is also abundant in corneas, blood vessels, the gut, intervertebral discs, and dentin in teeth. In muscle tissue, it serves as a major component of the endomysium. Collagen constitutes one to two percent of muscle tissue and accounts for 6% of the weight of strong, tendinous, muscles. The fibroblast is the most common cell that creates collagen. Gelatin, which is used in food and industry, is collagen that has been irreversibly hydrolyzed. In addition, partially and fully hydrolyzed collagen powders are used as dietary supplements. Collagen also participates in many binding interactions with target proteins in addition to its role in the structure.
The name collagen comes from the Greek (kólla), meaning "glue", and the suffix -gen, denoting "producing". This refers to the compound's early use in the process of boiling the skin and tendons of horses and other animals to obtain glue.
Over 90% of the collagen in the human body is type I. However, as of 2011, 28 types of collagen have been identified, described, and divided into several groups according to the structure they form. The five most common types are:
- Type I: skin, tendon, vasculature, organs, bone (the main component of the organic part of bone)
- Type II: cartilage (the main collagenous component of cartilage)
- Type III: reticulate (the main component of reticular fibers), commonly found alongside type I
- Type IV: forms basal lamina, the epithelium-secreted layer of the basement membrane
- Type V: cell surfaces, hair, and placenta
Let's focus on Type I collagen, which has unusual amino acid composition and sequence:
- Glycine is found at almost every third residue.
- Proline makes up about 17% of collagen.
- Collagen contains many hydroxyproline and hydroxylysine which are formed on post-translational modifications by different enzymes, both of which require vitamin C as a cofactor.
- Some hydroxylysines are glycosylated, mostly with disaccharides.
Figure \(\PageIndex{1}\) shows the post-translational hydroxylations of lysine and proline.
Vitamin C deficiency causes scurvy, a serious and painful disease in which defective collagen prevents the formation of strong connective tissue. Gums deteriorate and bleed, with loss of teeth; skin discolors, and wounds do not heal. Prior to the 18th century, this condition was notorious among long-duration military, particularly naval, expeditions during which participants were deprived of foods containing vitamin C. Many bacteria and viruses secrete virulence factors, such as the enzyme collagenase, which destroys collagen or interferes with its production.
Collagen has many (GXY)n repeats, where G is glycine (Gly), and X and Y are frequently proline (Pro) and hydroxyproline (Hyp). Three strands of collagen self-associate to for a triple-stranded helix with 10 GXY triplets in 3 complete turns of the helix. Others suggest that there are seven triplet units in 2 turns of the stands. Note that the helix of each strand in the triple helix is not an alpha helix and has different phi/psi angles. Each strand is "frameshifted" by one amino acid, resulting in a staggered arrangements of the individual stands and helices. The glycines are buried along the central axis so there is no essential hydrophobic core. The X, Y amino acids are solvent-exposed. All the other side chains, both hydrophobic and hydrophilic, are likewise exposed to solvent. Hydrogen bonding occurs between the amide hydrogen of the peptide bond of Gly and the carbonyl O of an X amino acid in another chain.
Figure \(\PageIndex{2}\) shows an interactive iCn3D model of a triple helical collagen-like peptide (4Z1R). The main chain atoms, shown in CPK colors, are shown forming hydrogren bonds with neighboring chains. The side chains are color based on the three chains (blue, brown and magenta). Two sets of Pro-HPro-Gly repeats are labeled.
Figure \(\PageIndex{2}\): Triple helical collagen-like peptide (4Z1R). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?TCrji1wPhekypJtc6
α-Keratin
α-keratin is the key structural element making up hair, nails, horns, claws, hooves, and the outer layer of skin. Due to its tightly wound structure, it can function as one of the strongest biological materials and has various uses in mammals, from predatory claws to hair for warmth.
The first sequences of α-keratins were determined by Hanukoglu and Fuchs. These sequences revealed that there are two distinct but homologous keratin families which were named Type I keratin and Type II keratins. There are 54 keratin genes in humans, 28 of which code for type I, and 26 for type II. Type I proteins are enriched in Asp and Glu amino acids, while type II proteins contain more basic amino acids, such as lysine. This differentiation is especially important in α-keratins because in the formation of a keratin dimer, the coiled coil, one protein coil must be type I, while the other must be type II. Even within type I and II, there are acidic and basic keratins that are particularly complementary within each organism. For example, in human skin, K5, a type II α-keratin, pairs primarily with K14, a type I α-keratin, to form the α-keratin complex of the epidermis layer of cells in the skin.
Coiled-coil dimers then assemble into a tetramer of two staggered coiled-coil dimers. Two tetramers can then pack together to form an elongated protofilament, a very stable, left-handed superhelical structure as shown in the figure below. The keratin filaments stay associated through hydrophobic interactions between apolar residues along the keratin's helical segments. This is illustrated in Figure \(\PageIndex{3}\).
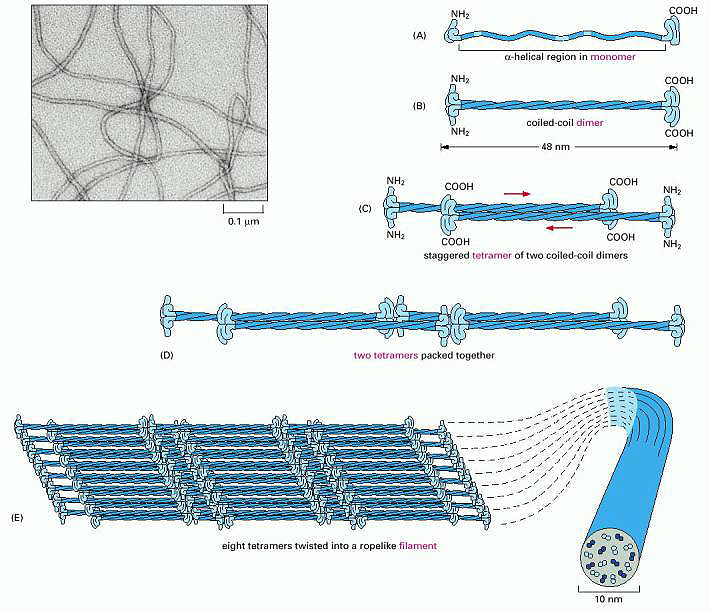
Figure \(\PageIndex{3}\): Assembly of Keratin Fibers. Wiki lectures. https://www.wikilectures.eu/w/Indivi...e_and_function
Initially, two keratin monomers (A) form a coiled coil dimer structure (B) Two coiled coil dimers join to form a staggered tetramer (C), next, the tetramers start to join together (D), ultimately forming a sheet of eight tetramers (E). The sheet of eight tetramers is then twisted into a lefthanded helix forming the final intermediate filament (E) An electron micrograph of the intermediate filament is shown in the upper left hand corner.
Figure \(\PageIndex{4}\) shows an interactive iCn3D model of a dimer of Type I alpha-keratin (magenta backbone) and Type II (blue backbone) (6JFV). Acidic (red) side chains (Asp and Glu) and basic (blue) side chains (Lys) can be seen projecting away from the dimer. Both A and B chains have negative and positive side chains. The A (more acidic) chain in this structure has 5 Lys , 7 Arg, 1 Asp and 16 Glu side chains for a net charge of +12 -17 = -5. The B (more basic) chain in this structure has 8 Lys , 9 Arg, 5 Asp and 12 Glu side chains for a net charge of +17 -17 = 0. It is clearly more basic with 17 Lys and Arg side chains, compared to the A chain with 12. Depending on their 3D orientation, they could present a positive face to the more negative monomer in the dimer.
Figure \(\PageIndex{4}\): Dimer of a Type I and Type II alpha-keratin backbones (6JFV). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...g3L3sQh9dj5pw8
Acidic (red) side chains (Asp and Glu) and basic (blue) side chains (Lys, Arg) can be seen projecting away from the dimer. Both A and B chains have negative and positive side chains. The A (more acidic) chain in this structure has 5 Lys , 7 Arg, 1 Asp and 16 Glu side chains for a net charge of +12 -17 = -5. The B (more basic) chain in this structure has 8 Lys , 9 Arg, 5 Asp and 12 Glu side chains for a net charge of +17 -17 = 0. It is clearly more basic with 17 Lys and Arg side chains, compared to the A chain with 12. Depending on their 3D orientation, they could present a positive face to the more negative monomer in the dimer.
Figure \(\PageIndex{5}\) shows an interactive iCn3D model of a spacefill model of the dimer. Note that a significant fraction of the nonpolar side chains is pointed inward between the two monomers and are much less exposed to solvent.
Figure \(\PageIndex{5}\): Dimer of a Type I and Type II alpha-keratin backbones in spacefill (6JFV). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...uyJrw28mZ6x5h7
Elastin
As its name implies, the protein confers elasticity in target structures such as connective tissue and blood vessels. It has low-complexity hydrophobic domains and the protein is cross-linked to form larger structures. It contains repeating hydrophobic amino acid sequences mostly of valine, proline, glycine and alanine, and mimetics of the repeating structure (LGGVG)6 have been studied. This protein also displays significant disorder.
Resilin
The following description of resilin is taken directly from an article under Creative Commons Attribution 4.0 International License available at http://creativecommons.org/licenses/by/4.0/. Balu, R., Dutta, N.K., Dutta, A.K. et al. Resilin-mimetics as a smart biomaterial platform for biomedical applications. Nat Commun 12, 149 (2021). https://doi.org/10.1038/s41467-020-20375-x
Native elastomeric proteins are biomaterials that have been perfected over billions of years by natural selection to act as molecular springs in a wide range of biological systems to drive unique functions. Among native proteins, resilin is purported to be one of the most efficient elastic proteins known. It is essentially a structural protein, which exists mainly in insect exoskeleton structures and exhibits outstanding resilience and fatigue life. The first description of resilin was made in 1960s as a rubber-like protein observed in locust-wing hinge and dragonfly tendon. Early studies on the composition and structure of resilin revealed the protein to contain about 66% hydrophobic residues (much lower than elastin) with about 45% proline and glycine residues combined. In native state, resilins exist as di- and trityrosine crosslinked hydrogels, and exhibit highly amorphous structures when examined using X-ray diffraction and electron microscopy. During biosynthesis, pro-resilins (uncrosslinked) are secreted from the apical surface of the epidermal cells into the subcuticular space, where they are crosslinked by an enzyme-mediated process to form hydrogels. Over the course of next three decades, resilin was also identified in many other insects and arthropods, including copepods, reduviidae and moth. In arthropods, resilin is largely involved in a number of different functions, including the flexibility and deformability of membranous cuticle and joint systems, the storage of elastic energy in locomotion (jumping, flying, etc.) and catapulting systems, the adaptability to surface topography by multiple contact attachment, and prey catching systems and the reduction of fatigue and damage in feeding and traumatic reproductive system.
The amino acid sequence of resilin was first identified in early 2000s from the CG15920 gene segment of the fruit fly Drosophila melanogaster, which opened up new opportunities for synthesis and development of biomimetic resilins. The CG15920 gene comprises N-terminal (exon-1), C-terminal (exon-3) and the middle chitin-binding (exon-2) domains, where exon-1 and exon-3 consist of 18 and 11 copies of consensus amino acid sequences: GGRPSDSYGAPGGGN and GYSGGRPGGQDLG, respectively. The first recombinant pro-resilin or resilin-like polypeptide (RLP), namely Rec1-resilin (encoded from the exon-1 of CG15920 gene) was synthesized in mid-2000s as a water soluble polypeptide expressed in the bacteria Escherichia coli. The synthesized pro-resilin was photo-crosslinked (dityrosine) using a ruthenium-persulfate crosslinking system to form hydrogels, which exhibited 97% resilience, outperforming native resilin dissected from dragonfly tendon (92%), natural elastin (90%) and synthetic polybutadiene rubber (80%)
The synthesized RLPs have several advantages over other elastomeric polypeptides, such as elastin-like polypeptides (ELPs), silk-like polypeptides (SLPs) and collagen-like polypeptides (CLPs). These include:
- Unique sequences rich in uncharged, polar amino acids and devoid of canonical hydrophobic residues, and contain high proportions of glycine- and proline-rich segments.
- Average negative hydropathy index.
- Intrinsically disordered protein structure with rapidly-interchangeable conformational ensemble in physiological conditions.
- Multi-stimuli (pH, temperature, ions, mechanical stress, other molecules, etc.) responsiveness, including dual-phase transition behavior (existence of both upper critical solution temperature, UCST and lower critical solution temperature, LCST).
- Low stiffness, high extensibility, outstanding resilience and excellent fatigue life.
- No inflammatory response.
Figure \(\PageIndex{6}\) shows possible transitions in the resilin protein which demonstrate such resiliency.
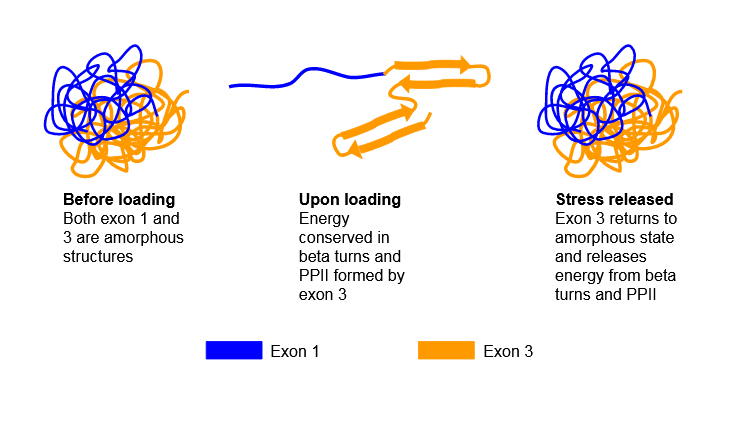
Given its disordered nature, there are no available PDB structures for resilin.
Fibrinogen
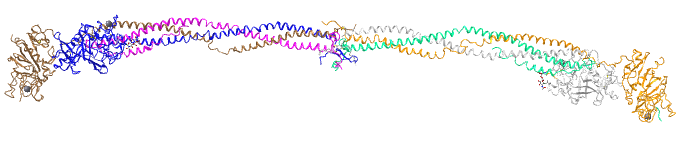
This very large molecule is a hexamer of three monomers (Aα, Bβ and γ) each present in two copies (α2β2γ2). Disulfide bond connect one structural unit (αβγ) with another. When two fibrinopeptides (FpA and FpB) are cleaved from the amino ends of the α and β chains by the clotting enzyme thrombin, small structural "knobs" form that bind to "holes" on another fibrinogen, causing the formation of large fibrils of fibrin clots. Figure \(\PageIndex{7}\) shows an interactive iCn3D model of human fibrinogen (3ghg). The "floppy" parts of the alpha chain (αC region) and FpA and FpB peptides are not shown as they were not resolved (due to their disorder) in the crystal structure.
Figure \(\PageIndex{7}\): Human fibrinogen (3ghg). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...S3dXzzsrkujKZ8
It is a very long, flexible molecule. The two alpha chains are shown in magenta and light green, the beta chains as dark blue and gray, and the gamma chains as brown and orange. Note the helical chains are actually alpha-helical for this molecule.
Figure \(\PageIndex{8}\) shows the domain structure and hints at the flexibility of this long molecule, which is required to form a fibrous mesh clot as well as to be accessible to an enzyme, plasmin, which cleaves fibrin clots, facilitating their removal. The Aα, Bβ and γ are shown in blue, red and green, respectively. Carbohydrates are shown in orange in the spacefilling model (b).
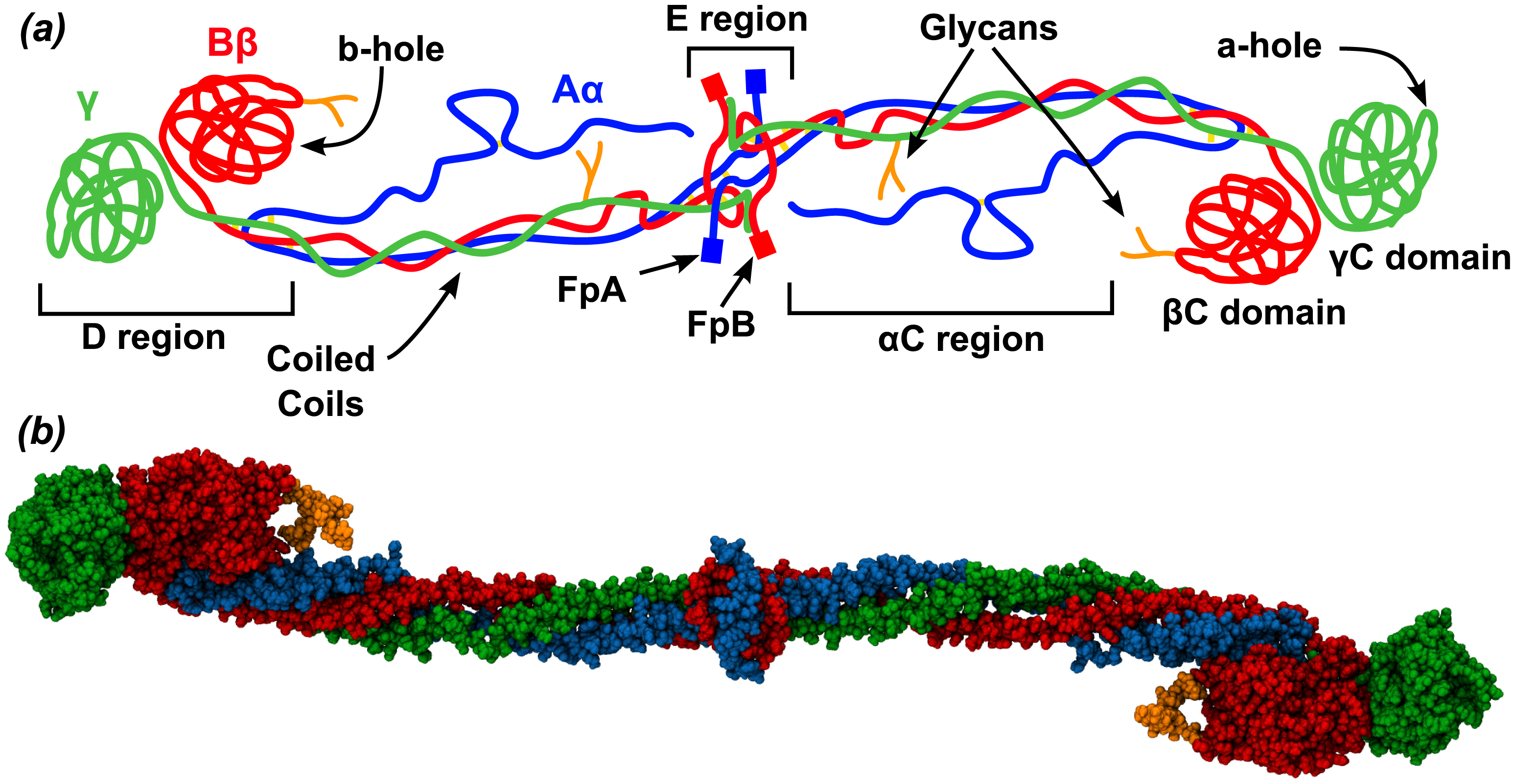
The coiled coils contain mostly alpha helical structures. The central E region is where the N-terminal ends of all of the fibrinogen chains are located and where the fibrinopeptides A (FpA, 16 residues) and B (FpB, 14 residues) are located, where they will be cleaved by thrombin in clot formation. The C-terminals of the chains are located at the distal D region which houses two C domains. After cleavage of the fibrinopeptides, conformational changes occur in the central E region to produce the "knobs" A (with starting sequence Gly-Pro-Arg) and B (Gly-His-Arg). These knobs bind through noncovalent interactions corresponding "holes" a and b at the distal D regions of another fibrinogen to form a dimer and subsequently a fibrin clot.
Myosin Heavy Chain
Myosin is a chief component of muscles and in complex with actin and other proteins allows muscle contraction. It has 2 clear domains and an elongated rod-like tail sequence, which has 28-residue repeats of 4 heptapeptides, characteristic of alpha-helical proteins that form the coiled-coils quaternary structure. Figure \(\PageIndex{9}\) shows the domain structure of human myosin heavy chain 1. The orange represents the motor domain of the protein, which binds and hydrolyzes ATP, providing the free energy that powers muscle contraction.

Figure \(\PageIndex{10}\) shows a cartoon of myosin heavy chains (blue) associated with myosin light chains, and how they interact with actin in the actinomyosin complex in striated muscle cells. The motor domain also binds actin filaments. In a simplistic way, the myosin thick filaments can be considered to slide back and forth in muscle contraction and extension.
Myosin can exist in two major conformations. One is the "6S" (extended tail) form that assembles into myosin filaments, which interacts with actin as shown in Figure \(\PageIndex{10}\) to transduce the chemical energy from ATP hydrolysis into mechanical forces and filament sliding. The other is the "10S" conformation which is folded on itself. The heads interact with each other and the tail. In this compact form, all necessary steps (ATP cleavage, filament assembly, actin activation) required for actin/myosin- mediated contraction are inhibited.
Figure \(\PageIndex{11}\) shows an interactive iCn3D model of an inactive (nonextended) conformation of myosin heavy chain (6xe9) from smooth muscle. The long cyan and green chains are the myosin (II) heavy chains from smooth muscle.
Figure \(\PageIndex{11}\): Inactive (nonextended) conformation of myosin heavy chain (6xe9). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...thwKbyQQ5ezB96
References
OpenStax, Proteins. OpenStax CNX. Sep 30, 2016 http://cnx.org/contents/bf17f4df-605c-4388-88c2-25b0f000b0ed@2.
File:Chirality with hands.jpg. (2017, September 16). Wikimedia Commons, the free media repository. Retrieved 17:34, July 10, 2019 from commons.wikimedia.org/w/index.php?title=File:Chirality_with_hands.jpg&oldid=258750003.
Wikipedia contributors. (2019, July 6). Zwitterion. In Wikipedia, The Free Encyclopedia. Retrieved 21:48, July 10, 2019, from en.Wikipedia.org/w/index.php?title=Zwitterion&oldid=905089721
Wikipedia contributors. (2019, July 8). Absolute configuration. In Wikipedia, The Free Encyclopedia. Retrieved 15:28, July 14, 2019, from en.Wikipedia.org/w/index.php?title=Absolute_configuration&oldid=905412423
Structural Biochemistry/Enzyme/Active Site. (2019, July 1). Wikibooks, The Free Textbook Project. Retrieved 16:55, July 16, 2019 from en.wikibooks.org/w/index.php?title=Structural_Biochemistry/Enzyme/Active_Site&oldid=3555410.
Structural Biochemistry/Proteins. (2019, March 24). Wikibooks, The Free Textbook Project. Retrieved 19:16, July 18, 2019 from en.wikibooks.org/w/index.php?title=Structural_Biochemistry/Proteins&oldid=3529061.
Fujiwara, K., Toda, H., and Ikeguchi, M. (2012) Dependence of a α-helical and β-sheet amino acid propensities on teh overall protein fold type. BMC Structural Biology 12:18. Available at: https://bmcstructbiol.biomedcentral.com/track/pdf/10.1186/1472-6807-12-18
Wikipedia contributors. (2019, July 16). Keratin. In Wikipedia, The Free Encyclopedia. Retrieved 17:50, July 19, 2019, from en.Wikipedia.org/w/index.php?title=Keratin&oldid=906578340
Wikipedia contributors. (2019, July 13). Alpha-keratin. In Wikipedia, The Free Encyclopedia. Retrieved 18:17, July 19, 2019, from en.Wikipedia.org/w/index.php?title=Alpha-keratin&oldid=906117410
Open Learning Initiative. (2019) Integumentary Levels of Organization. Carnegie Mellon University. In Anatomy & Physiology. Available at: https://oli.cmu.edu/jcourse/webui/syllabus/module.do?context=4348901580020ca6010f804da8baf7ba.
Wikipedia contributors. (2019, July 16). Collagen. In Wikipedia, The Free Encyclopedia. Retrieved 03:42, July 20, 2019, from en.Wikipedia.org/w/index.php?title=Collagen&oldid=906509954
Wikipedia contributors. (2019, July 2). Rossmann fold. In Wikipedia, The Free Encyclopedia. Retrieved 16:01, July 20, 2019, from https://en.Wikipedia.org/w/index.php?title=Rossmann_fold&oldid=904468788
Wikipedia contributors. (2019, May 30). TIM barrel. In Wikipedia, The Free Encyclopedia. Retrieved 16:46, July 20, 2019, from en.Wikipedia.org/w/index.php?title=TIM_barrel&oldid=899459569
Wikipedia contributors. (2019, July 16). Protein folding. In Wikipedia, The Free Encyclopedia. Retrieved 18:30, July 20, 2019, from https://en.Wikipedia.org/w/index.php?title=Protein_folding&oldid=906604145
Wikipedia contributors. (2019, June 11). Globular protein. In Wikipedia, The Free Encyclopedia. Retrieved 18:49, July 20, 2019, from en.Wikipedia.org/w/index.php?title=Globular_protein&oldid=901360467
Wikipedia contributors. (2019, July 11). Intrinsically disordered proteins. In Wikipedia, The Free Encyclopedia. Retrieved 19:52, July 20, 2019, from en.Wikipedia.org/w/index.php?title=Intrinsically_disordered_proteins&oldid=905782287
Comprehensive Database for Protein Analysis - Biozon
SCOP: Structural Characterization of Proteins - Database showing folds, superfamiles, families, and domains




.png?revision=1)
.png?revision=1&size=bestfit&width=385)
.png?revision=1)
.png?revision=1)
_conformation_of_myosin_heavy_chain_(6xe9).png?revision=1)