6.4D: The FISH Technique
- Page ID
- 9171
- Describe how fluorescent in situ hybridization (FISH) is used in clinical and biomedical studies to detect and localize the presence or absence of specific DNA sequences and to identify pathogens
FISH (fluorescence in situ hybridization) is a cytogenetic technique developed by biomedical researchers in the early 1980s. It is used to detect and localize the presence or absence of specific DNA sequences on chromosomes. FISH uses fluorescent probes bind to those targets that show a high degree of sequence complementarity. FISH can be used to detect RNA or DNA sequences of interest. FISH is often used for finding specific features in DNA for use in genetic counseling, medicine, and species identification. FISH can also be used to detect and localize specific RNA targets, including mRNAs, in cells. In this context, it can help define the spatial-temporal patterns of gene expression within cells and tissues.
Central to FISH are the use of probes. The probe must be large enough to hybridize specifically with its target but not so large as to impede the hybridization process. They are anti-sense to the target mRNA or DNA of interest, thus they hybridize to targets. The probe can be tagged directly with fluorophores, or with targets for flourescently labelled antibodies or other substrates. Different types of tags can be used, therefore different targets can be detected in the same sample simultaneously (multi-colour FISH). Tagging can be done in various ways, such as nick translation, or PCR using tagged nucleotides. Probes can vary in length from 20 to 30 nucleotides to much longer sequences.
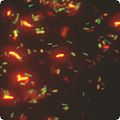
FISH is often used in clinical studies. If a patient is infected with a suspected pathogen, bacteria from the patient’s tissues or fluids, are typically grown on agar to determine the identity of the pathogen. Many bacteria, however, even well-known species, do not grow well under laboratory conditions. FISH can be used to directly detect the presence of the suspect on small samples of the patient’s tissue. FISH can also be used to compare the genomes of two biological species, to deduce evolutionary relationships. A similar hybridization technique is called a zoo blot. Bacterial FISH probes are often primers for the 16s rRNA region. FISH is widely used in the field of microbial ecology, to identify microorganisms. Biofilms, for example, are composed of complex (often) multi-species bacterial organizations. Preparing DNA probes for one species and performing FISH with this probe allows one to visualize the distribution of this specific species within the biofilm. Preparing probes (in two different colors) for two species allows to visualize/study co-localization of these two species in the biofilm, and can be useful in determining the fine architecture of the biofilm.
Key Points
- FISH can be used in a clinical setting to identify pathogens or DNA / RNA targets of interest.
- FISH is used to detect and localize the presence or absence of specific DNA or RNA sequences in tissue or cells.
- FISH can also be used to compare the genomes of two biological species, such as in ecological studies, where a bacteria may not be culturable, it can be identified using FISH.


