8.9: Generation of Single Strands
- Page ID
- 357
One of the major pathways for generating 3’ single-stranded termini uses the RecBCD enzyme, also known as exonuclease V (Figure 8.13). The three subunits of this enzyme are encoded by the genes recB, recC, and recD. Each model for recombination requires a single-strand with with a free end for strand invasion, and this enzyme does so, but with several unexpected features.
RecBCD has multiple functions, and it can switch activities. It is a helicase (in the presence of SSB), an ATPase and a nuclease. The nuclease can be a 3’ to 5’ exonuclease, and endonuclease or a 5’ to 3’ exonuclease, at different steps of the process.
The helicase activity of the RecBCD enzyme initiates unwinding only on DNA containing a free duplex end. It binds to the duplex end, using the energy of ATP hydrolysis to travel along the duplex, unwinding the DNA. The enzyme complex tracks along the top strand faster than it does on the bottom strand, so single‑stranded loops emerge, getting progressively larger as it moves down the duplex. These loops can be visualized in electron micrographs. RecBCD is also a 3' to 5' exonuclease during this phase, removing the end of one of the unwound strands (Figure 8.13).
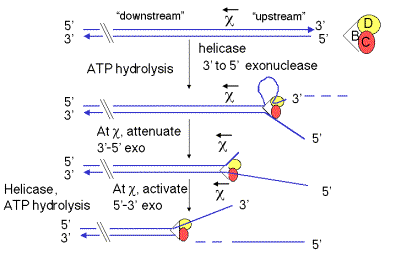
The activities of the RecBCD enzyme change at particular sequences in the DNA called chi sites(for the Greek letter c). The sequence of a chi site is 5' GCTGGTGG; this occurs about once every 4 kb on the E. coligenome. Genetic experiments show that RecBCD promotes recombination most frequently at chi sites. These sites were first discovered as mutations in bacteriophage l that led to increased recombination at those sites. These mutations altered the l sequence at the site of the mutation to become a chi site (GCTGGTGG).
When the RecBCD enzyme encounters a chi site, it will leave an extruded single strand close to this site (4 to 6 nucleotides 3' to it). A chi site serves as a signal to RecBCD to shift the polarity of its exonuclease function. Before reaching the chi site, RecBCD acts primarily as a 3’ to 5’ exonuclease, e.g. working on the top strand in Figure 8.13. At the chi site, the 3’ to 5’ exonuclease function is suppressed, and afterthe chi site, RecBCD converts to a 5’ to 3’ exonuclease, now working on the other strand (e.g. the bottom strand in Figure 8.13). Presumably, the strand that will be the substrate for the 5’ to 3’ exonuclease is nicked in concert with this conversion in polarity of the exonuclease. This process leaves the chi site at the 3’ end of a single stranded DNA. This is the substrate to which RecA can bind to initiate strand exchange (see below).
Some tests of the models for recombination have examined whether chi sites serve preferentially as either donors or recipients of the DNA during recombination. However, both results have been obtained, which makes it difficult to tie this activity precisely into either model for recombination. The genetic evidence is clear, however, that it is needed for one major pathway of recombination.
Exercise 8.4
What are the predictions of the Holliday model and the double-strand-break model for whether chi sites would be used as donors or recipients of genetic information during recombination?
An alternative pathway for generating single-strand ends for recombination uses the enzyme RecE, also known as exonuclease VIII. This pathway is revealed in recBCD- mutants. RecE is a 5’ to 3’ exonuclease that digests double‑stranded linear DNA, thereby generating single‑stranded 3' tails. RecE is encoded on a cryptic plasmid in E. coli. It is similar to the redexonuclease encoded by bacteriophage l.
A third pathway used the RecQhelicase, which is also a DNA-dependent ATPase. This pathway is revealed in recBCD- recE-mutants. The result of its helicase activity, in the presence of SSB, is the formation of a DNA molecule with single-stranded 3' tails, which can be used for strand invasion.


