7.1: Mutations and Mutagens
- Page ID
- 342
Mutations commonly are substitutions, in which a single nucleotide is changed into a different nucleotide. Other mutations result in the loss (deletion) or addition (insertion) of one or more nucleotides. These insertions or deletions can range from one to tens of thousands of nucleotides. Often an insertion or deletion is inferred from comparison of two homologous sequences, and it may be impossible to ascertain from the data given whether the presence of a segment in one sequence but not another resulted from an insertion of a deletion (in this case, it can be referred to as an "indel"). One mechanism for large insertions is the transposition of a sequence from one place in a genome to another (described in Chapter 9).
Types of muations
Nucleotide substitutions are one of two classes. In a transition, a purine nucleotide is replaced with a purine nucleotide, or a pyrimidine nucleotide is replaced with a pyrimidine nucleotide. In other words, the base in the new nucleotide is in the same chemical class as that of the original nucleotide. In a transversion, the chemical class of the base changes, i.e. a purine nucleotide is replaced with a pyrimidine nucleotide, or a pyrimidine nucleotide is replaced with a purine nucleotide.
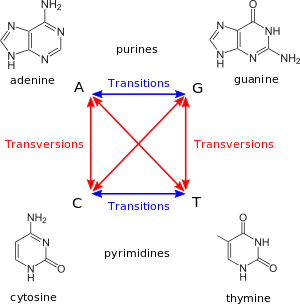
Comparison of the sequences of homologous genes between species reveals a pronounced preference for transitions over transversions (about 10-fold), indicating that transitions occur much more frequently than transversions.
Errors in Replication
Despite effective proofreading functions in many DNA polymerases, occasionally the wrong nucleotide is incorporated. It is estimated that E. coliDNA polymerase III holoenzyme (with a fully functional proofreading activity) uses the wrong nucleotide during elongation about 1 in 108 times. It is more likely for an incorrect pyrimidine nucleotide to be incorporated opposite a purine nucleotide in the template strand, and for a purine nucleotide to be incorporated opposite a pyrimidine nucleotide. Thus these misincorporations resulting in a transition substitution are more common. However, incorporation of a pyrimidine nucleotide opposite another pyrimidine nucleotide, or a purine nucleotide opposite another purine nucleotide, can occur, albeit at progressively lower frequencies. These rarer misincorporations lead to transversions.
Exercise \(\PageIndex{1}\)
If a dCTP is incorporated into a growing DNA strand opposite an A in the template strand, what mutation will result? Is it a transition or a transversion?
Exercise \(\PageIndex{2}\)
If a dCTP is incorporated into a growing DNA strand opposite a T in the template strand, what mutation will result? Is it a transition or a transversion?
A change in the isomeric form of a purine or pyrimidine base in a nucleotide can result in a mutation. The base-pairing rules are based on the hydrogen-bonding capacity of nucleotides with their bases in the ketotautomer. A nucleotide whose base is in the enoltautomer can pair with the "wrong" base in another nucleotide. For example, a T in the rare enolisomer will pair with a keto G (Figure \(\PageIndex{2}\)), and an enolG will pair with a ketoT.

The enoltautomers of the normal deoxynucleotides guanidylate and thymidylate are rare, meaning that a single molecule is in the ketoform most of the time, or within a population of molecules, most of them are in the ketoform. However, certain nucleoside and base analogs adopt these alternative isomers more readily. For instance 5-bromo-deoxyuridine (or 5-BrdU) is an analog of deoxythymidine (dT) that is in the enoltautomer more frequently than dT is (although most of the time it is in the ketotautomer).
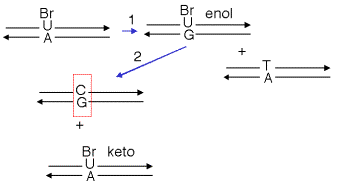
Thus the frequency of misincorporation can be increased by growth in the presence of base and nucleoside analogs. For example, growth in the presence of 5-BrdU results in an increase in the incorporation of G opposite a T in the DNA, as illustrated in Figure \(\PageIndex{3}\). After cells take up the nucleoside 5-BrdU, it is converted to 5-BrdUTP by nucleotide salvage enzymes that add phosphates to its 5’ end. During replication, 5-BrdUTP (in the ketotautomer) will incorporate opposite an A in DNA. The 5-BrdU can shift into the enolform while in DNA, so that when it serves as a template during the next round of replication (arrow 1 in the diagram below), it will direct incorporation of a G in the complementary strand. This G will in turn direct incorporation of a C in the top strand in the next round of replication (arrow 2). This leaves a C:G base pair where there was a T:A base pair in the parental DNA. Once the pyrimidine shifts back to the favored ketotautomer, it can direct incorporation of an A, to give the second product in the diagram below (with a BrU-A base pair).
Exercise \(\PageIndex{3}\)
Where are the hydrogen bonds in a base pair between enol –guanidine and keto-thymidine in DNA?
Likewise, misincorporation of A and C can occur when they are in the rare iminotautomers rather than the favored aminotautomers. In particular, iminoC will pair with aminoA, and iminoA will pair with aminoC (Figure \(\PageIndex{4}\)).

Misincorporation during replication is the major pathway for introducing transversions into DNA. Normally, DNA is a series of purine:pyrimidine base pairs, but in order to have a transversion, a pyrimidine has to be paired with another pyrimidine, or a purine with a purine. The DNA has to undergo local structural changes to accommodate these unusual base pairs. One way this can happen for a purine-purine base pair is for one of the purine nucleotides to shift from the preferred anticonformation to the synconformation. Atoms on the "back side" of the purine nucleotide in the syn-isomer can form hydrogen bonds with atoms in the rare tautomer of the purine nucleotide, still in the preferred anticonformation. For example, an A nucleotide in the syn-, amino- isomer can pair with an A nucleotide in the anti-, imino- form (Figure \(\PageIndex{5}\)). Thus the transversion required a shift in the tautomeric form of the base in one nucleotide as well as a change in the base-sugar conformation (antito syn) of the other nucleotide.

Exercise \(\PageIndex{4}\)
Why does the shift of a purine nucleotide from anti to synhelp allow a purine:purine base pair? Is this needed for a pyrimidine:pyrimidine base pair?
Errors in replication are not limited to substitutions. Slippage errors during replication will add or delete nucleotides. A DNA polymerase can insert additional nucleotides, more commonly when tandem short repeats are the template (e.g. repeating CA dinucleotides). Sometimes the template strand can loop out and form a secondary structure that the DNA polymerase does not read. In this case, a deletion in the nascent strand will result.


