3.1: Recombinant DNA, Polymerase Chain Reaction and Applications to Eukaryotic Gene Structure and Function
- Page ID
- 10521
Methods to purify some abundant proteins were developed early in the 20th century, and some of the experiments on the fine structure of the gene (colinearity of gene and protein for trpA and tryptophan synthase) used microbial genetics and proteins sequencing. However, methods to isolate genes were not developed until the 1960’s, and the were applicable to only a few genes.
All this changed in the late 1970’s with the development of recombinant DNA technology, or molecular cloning. This technique enabled researchers to isolate any gene from any organism from which one could isolate intact DNA (or RNA). The full potential to provide access to all genes of organisms is now being realized as full genomes are sequenced. One of the by-products of the intense investigation of individual DNA molecules after the advent of recombinant DNA was a procedure to isolate any DNA for which one knows the sequence. This technique, called the polymerase chain reaction (PCR), is far easier than traditional molecular cloning methods, and it has become a staple of many laboratories in the life sciences. After covering the basic techniques in recombinant DNA technology and PCR, their application to studies of eukaryotic gene structure and function will be discussed.
Like many advances in molecular genetics, recombinant DNA technology has its roots in bacterial genetics.
Transducing Phage
The first genes isolated were bacterial genes that could be picked up by bacteriophage. By isolating these hybrid bacteriophage, the DNA for the bacterial gene could be recovered in a highly enriched form. This is the basic principal behind recombinant DNA technology.
Some bacteriophage will integrate into a bacterial chromosome and reside in a dormant state (Figure 3.1). The integrated phage DNA is called a prophage, and the bacterium is now a lysogen. Phage that do this are lysogenic. Induction of the lysogen will result in excision of the prophage and multiplication to produce many progeny, i.e. it enters a lytic phasein which the bacteria are broken open and destroyed. The nomenclature is descriptive. The bacteria carrying the prophage show no obvious signs of the phage (except immunity to superinfection with the same phage, covered later in Part Four), but when induced (e.g. by stress or UV radiation) they will generate a lytic state, hence they are called lysogens. Induced lysogens make phage from the prophage that was integrated. Phage that always multiply when they infect a cell are called lytic.
Excision of a prophage from a lysogen is notalways precise. Usually only the phage DNA is cut out of the bacterial chromosome, but occassionally some adjacent host DNA is included with the excised phage DNA and encapsidated in the progeny. These transducing phage are usually biologically inactive because the piece of the bacterial chromosome replaces part of the phage chromosome; these can be propagated in the presence of helper phage that provide the missing genes when co-infected into the same bacteria. When DNA from the transducing phage is inserted into the newly infected cell, the bacterial genes can recombine into the host chromosome, thereby bringing in new alleles or even new genes and genetically altering the infected cell. This process is called transduction.
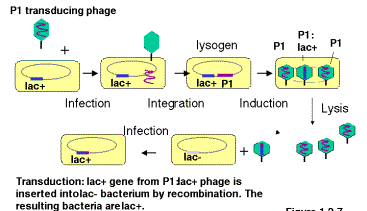
Note that the transducing phage are carrying one or a small number of bacterial genes. This is a way of isolating the genes. The bacterial gene in the transducing phage has been separated from the other 4000 bacterial genes (in E. coli). By isolating large numbers of the transducing phage, the phage DNA, including the bacterial genes, can be obtained in large quantitiesfor biochemical investigation. One can isolate mg or mg quantities of a single DNA molecule, which allows for precise structural determination and detailed investigation.
A generalized transducing phagecan integrate at many different locations on the bacterial chromosome. Imprecise excision from any of those locations generates a particular transducing phage, carrying a short sections of the bacterial genome adjacent to the integration site. Thus a generalized transducing phage such as P1 can pick up many different parts of the E. coli genome.
A specialized transducing phageintegrates into only one or very few sites in the host genome. Hence it can carryonly a few specific bacterial genes, e.g., l lac(Figure 3.2).

This process of isolating a particular bacterial gene on a transducing phage is mimicked in recombinant DNA technology, in which a gene or genome fragment from any organism is isolated on a recombinant phage or plasmid.


