5.3: Inferring the Mode of Inheritance
- Page ID
- 4037
Given a pedigree of an uncharacterized disease or trait, one of the first tasks is to determine which modes of inheritance are possible and then which mode of inheritance is most likely. This information is essential in calculating the probability that the trait will be inherited in any future offspring. We will mostly consider five major types of inheritance: autosomal dominant (AD), autosomal recessive (AR), X-linked dominant (XD), X-linked recessive (XR), and Y-linked (Y).
Autosomal Dominant (AD)
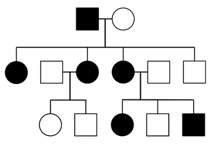
When a disease is caused by a dominant allele of a gene, every person with that allele will show symptoms of the disease (assuming complete penetrance), and only one disease allele needs to be inherited for an individual to be affected. Thus, every affected individual must have an affected parent. A pedigree with affected individuals in every generation is typical of AD diseases. However, beware that other modes of inheritance can also show the disease in every generation, as described below. It is also possible for an affected individual with an AD disease to have a family without any affected children, if the affected parent is a heterozygote. This is particularly true in small families, where the probability of every child inheriting the normal, rather than disease allele is not extremely small. Note that AD diseases are usually rare in populations, therefore affected individuals with AD diseases tend to be heterozygotes (otherwise, both parents would have had to been affected with the same rare disease). Achondroplastic dwarfism, and polydactyly are both examples of human conditions that may follow an AD mode of inheritance.

X-linked dominant (XD)
In X-linked dominant inheritance, the gene responsible for the disease is located on the X-chromosome, and the allele that causes the disease is dominant to the normal allele in females. Because females have twice as many X-chromosomes as males, females tend to be more frequently affected than males in the population. However, not all pedigrees provide sufficient information to distinguish XD and AD. One definitive indication that a trait is inherited as AD, and not XD, is that an affected father passes the disease to a son; this type of transmission is not possible with XD, since males inherit their X chromosome from their mothers.

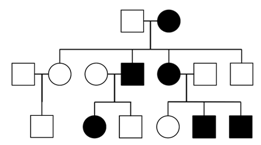
Autosomal recessive (AR)
Diseases that are inherited in an autosomal recessive pattern require that both parents of an affected individual carry at least one copy of the disease allele. With AR traits, many individuals in a pedigree can be carriers, probably without knowing it. Compared to pedigrees of dominant traits, AR pedigrees tend to show fewer affected individuals and are more likely than AD or XD to “skip a generation”. Thus, the major feature that distinguishes AR from AD or XD is that unaffected individuals can have affected offspring.

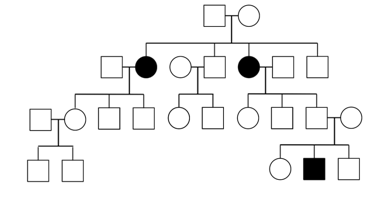
X-linked recessive (XR)
Because males have only one X-chromosome, any male that inherits an X-linked recessive disease allele will be affected by it (assuming complete penetrance). Therefore, in XR modes of inheritance, males tend to be affected more frequently than females in a population. This is in contrast to AR and AD, where both sexes tend to be affected equally, and XD, in which females are affected more frequently. Note, however, in the small sample sizes typical of human families, it is usually not possible to accurately determine whether one sex is affected more frequently than others. On the other hand, one feature of a pedigree that can be used to definitively establish that an inheritance pattern is not XR is the presence of an affected daughter from unaffected parents; because she would have had to inherit one X-chromosome from her father, he would also have been affected in XR.

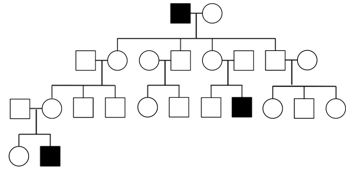
Y-linked and Mitochondrial Inheritance.
Two additional modes are Y-linked and Mitochondrial inheritance.
Only males are affected in human Y-linked inheritance (and other species with the X/Y sex determining system). There is only father to son transmission. This is the easiest mode of inheritance to identify, but it is one of the rarest because there are so few genes located on the Y-chromosome. An example of Y-linked inheritance is the hairy-ear-rim phenotype seen in some Indian families. As expected this trait is passed on from father to all sons and no daughters. Y-chromosome DNA polymorphisms can be used to follow the male lineage in large families or through ancient ancestral lineages. For example, the Y-chromosome of Mongolian ruler Genghis Khan (1162-1227 CE), and his male relatives, accounts for ~8% of the Y-chromosome lineage of men in Asia (0.5% world wide).
Mutations in Mitochondrial DNA are inherited through the maternal line (from your mother). There are some human diseases associated with mutations in mitochondria genes. These mutations can affect both males and females, but males cannot pass them on as the mitochondria are inherited via the egg, not the sperm. Mitochondrial DNA polymorphisms are also used to investigate evolutionary lineages, both ancient and recent. Because of the relative similarity of sequence mtDNA is also used in species identification in ecology studies.


