19.8: Enzyme Regulation
- Page ID
- 3436
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \) \( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)\(\newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\) \( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\) \( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\) \( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\) \( \newcommand{\Span}{\mathrm{span}}\) \(\newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\) \( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\) \( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\) \( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\) \( \newcommand{\Span}{\mathrm{span}}\)\(\newcommand{\AA}{\unicode[.8,0]{x212B}}\)
Briefly compare the genetic control of enzyme activity in bacteria with control of enzyme activity through feedback inhibition. Briefly compare an inducible operon with a repressible operon. Briefly compare competitive inhibition with noncompetitive inhibition.
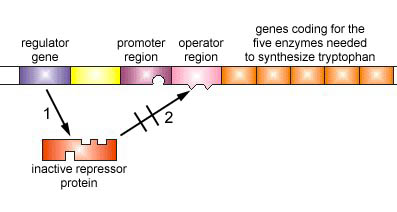
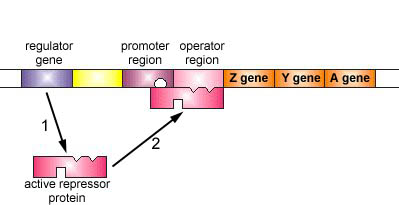
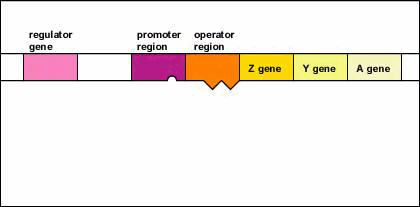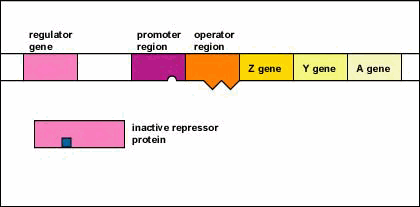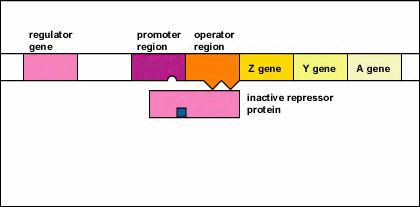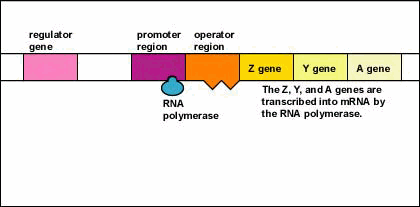
Figure \(\PageIndex{5}\): An Inducible Operon in the Absence of an Inducer (The Lactose Operon). Step 1: The regulator gene codes for an active repressor protein.
Step 2: The repressor protein then binds to the operator region of the operon.
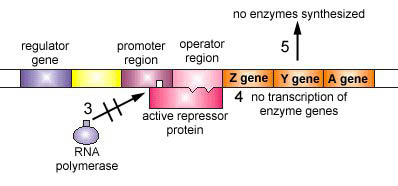
The regulator gene codes for an active repressor protein. The repressor protein then binds to the operator region of the operon. With the active repressor protein bound to the operator region, RNA polymerase (the enzyme responsible for the transcription of genes) is unable to bind to the promoter region of the operon. If RNA polymerase does not bind to the promoter region, the three enzyme genes (Z, Y, and A) are not transcribed into mRNA. Without the transcription of the three enzyme genes, the three enzymes needed for the utilization of the sugar lactose by the bacterium are not synthesized.
- The regulator gene codes for an active repressor protein.
- Lactose, the inducer molecule binds to the active repressor protein.
- The binding of the inducer alters the shape of the allosteric repressor causing it to become inactivated.
- The inactivated repressor protein is then unable to bind to the operator region of the operon.
- Since the inactive repressor protein is unable to bind to the operator region, RNA polymerase (the enzyme responsible for the transcription of genes) is now able to bind to the promoter region of the operon.
- RNA polymerase is now able to transcribe the three enzyme genes (Z, Y, and A) into mRNA.
- With the transcription of these genes, the three enzymes needed for the bacterium to utilize the sugar lactose are now synthesized. (The Z gene codes for beta-galactosidase, an enzyme that breaks down lactose into glucose and galactose. The Y gene codes for permease, an enzyme which transports lactose into the bacterium. The A gene codes for transacetylase, an enzyme which is thought to aid in the release of galactosides.)
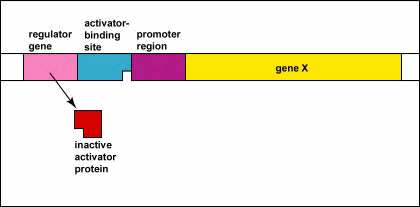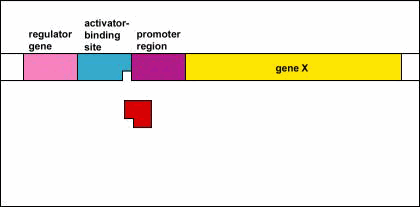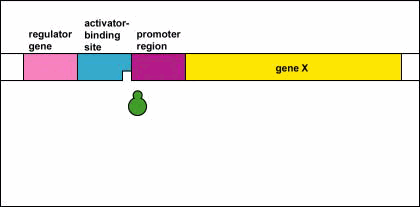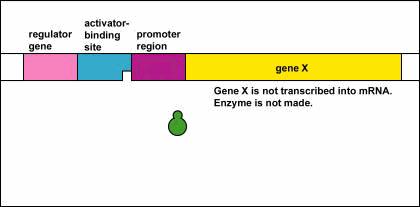
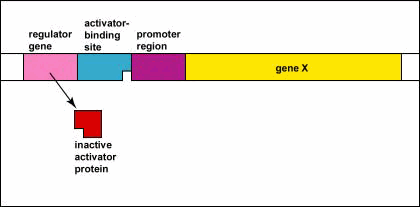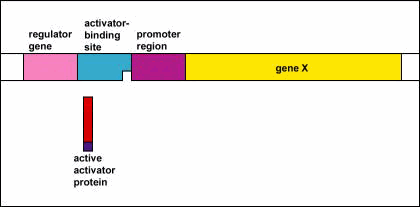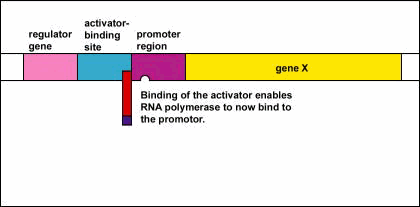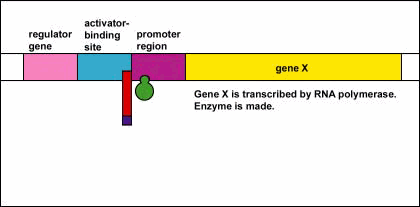
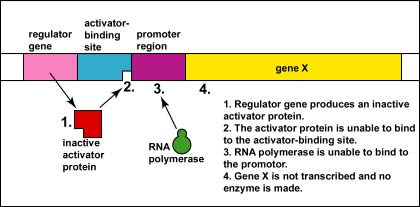
Genetic Control: Activators
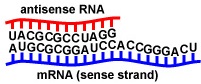
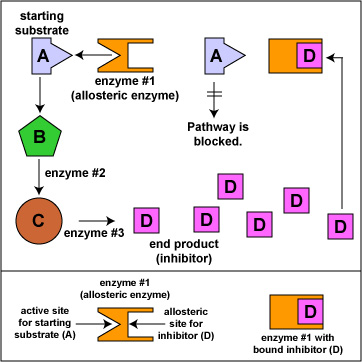
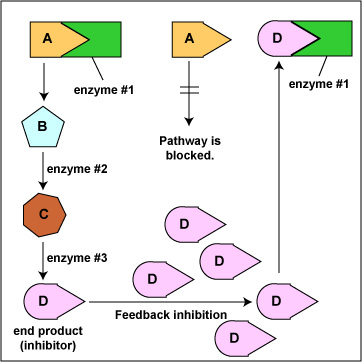
Feedback Inhibition