10.6C: The Life Cycle of HIV
- Page ID
- 3240
- Describe how the retrovirus HIV-1 accomplishes each of the following steps during its life cycle. (Include the following key words in your description: gp120, CD4, chemokine receptors, gp41, capsid, RNA genome, reverse transcriptase, double-stranded DNA intermediate, provirus, polyproteins, proteases, and budding.)
- viral attachment or adsorption to the host cell
- viral entry into the host cell
- viral movement to the site of replication within the host cell and production of a provirus.
- viral replication within the host cell
- viral assembly or maturation within the host cell and release from the host cell
- Name 3 types of cells HIV primarily infects and briefly explain why.
The Structure of the Human Immunodeficiency Virus (HIV)
HIV (see HIV A, HIV B and HIV C) has an envelope derived from host cell membranes during replication. Associated with the envelope are two HIV-encoded glycoproteins, gp120 and gp41. Underneath the envelope is a protein matrix composed of p17. Inside the virus is a capsid or core made of the protein p24. The nucleocapsid also contains p6, p7, reverse transcriptase (p66/p51), integrase (p32), protease (p10), and 2 molecules of single-stranded RNA, the viral genome (see Figure \(\PageIndex{3}\)).
![hivgenes [Converted].jpg](https://bio.libretexts.org/@api/deki/files/12124/hivgenes_%255BConverted%255D.jpg?revision=1)
To view further electron micrographs of HIV, see the AIDS Pathology Tutorial at the University of Utah.
The Life Cycle for the Human Immunodeficiency Virus (HIV)
1. Attachment or Adsorption to the Host Cell
Initially, HIV uses a cellular protein called cyclophilin that is a component of its envelope to bind a low affinity host cell receptor called heparin. This first interaction (not shown in the illustrations or animations) enables the virus to initially make contact with the host cell. In order to infect a human cell, however, an envelope glycoprotein on the surface of HIV called gp120 must adsorbs to both a CD4 molecule and then a chemokine receptor found on the surface of only certain types of certain human cells.
Human cells possessing CD4 molecules include:
- T4-helper lymphocytes (also called T4-cells and CD4+ cells)
- monocytes
- macrophages
- dendritic cells
Chemokines are cytokines that promote an inflammatory response by pulling white blood cells out of the blood vessels and into the tissue to fight infection. Different white blood cells have receptors on their surface for different chemokines. The chemokine receptors are now thought to determine the type of CD4+ cell HIV is able to infect. First, a portion or domain of the HIV surface glycoprotein gp120 binds to its primary receptor, a CD4 molecule on the host cell. This induces a change in shape that enables the chemokine receptor binding domains of the gp120 to interact with a host cell chemokine receptor. The chemokine receptor functions as the viral co-receptor. This interaction brings about another conformational change that exposes a previously buried portion of the transmembrane glycoprotein gp41 called the fusion peptide that enables the viral envelope to fuse with the host cell membrane (see Figure \(\PageIndex{1}\)A, Figure \(\PageIndex{1}\)B), and Figure \(\PageIndex{1}\)C).
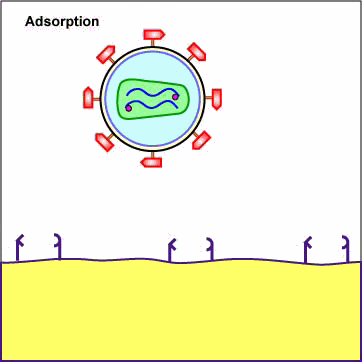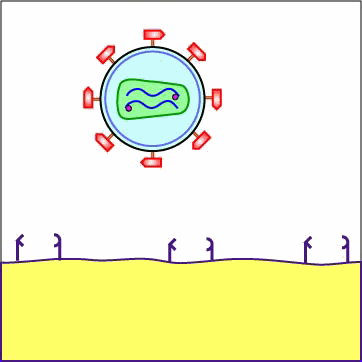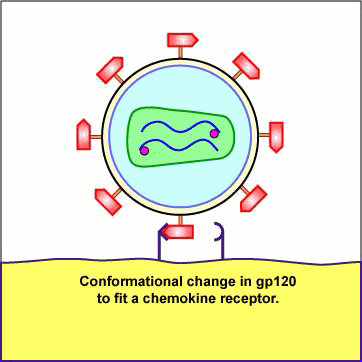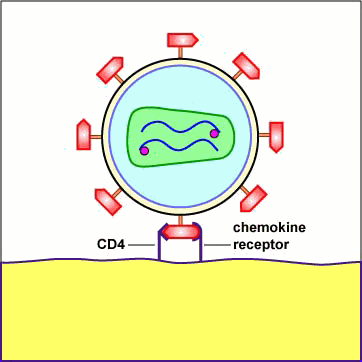
Animation: Adsorption of HIV to a T4-Helper Lymphocyte. The HIV envelope gp120 must attach to both a CD4 molecule and a chemokine receptor on the surface of such cells as macrophages and T4-helper lymphocytes in order to enter the cell. The gp120 first binds to a CD4 molecule on the plasma membrane of the host cell. The interaction between the gp120 and the CD4 molecule on the host cell induces a change in shape that brings the chemokine receptor binding domains of the gp120 into proximity with the host cell chemokine receptor
- Transmission electron micrograph showing envelope and glycoprotein spikes (gp120) of HIV; courtesy of CDC.
- Scanning electron micrograph showing HIV infecting a T4-lymphocyte; courtesy of CDC.
YouTube animation illustrating adsorption and penetration of HIV.
Most strains of HIV are referred to as M-tropic or T-tropic. The gp120 of M-tropic HIV (see Figure \(\PageIndex{2}\)) is able to adsorb to the CD4 molecules and the CCR5 chemokine receptors found on CD4+ macrophages, immature dendritic cells, and memory T4-lymphocytes. (M-tropic HIV are also called R5 viruses since they adsorb to the chemokine receptor CCR5.) M-tropic HIV require only low levels of CD4 molecules expressed on the surface of the host cell for infection. M-tropic HIV are thought to spread the infection. These strains appear to be slower-replicating and less virulent than the later T-tropic strains and do not cause the formation of syncytias. HIV initially replicates to high levels within macrophages without destroying them. (The T-tropic HIV, found later in HIV infection, are faster-replicating, more virulent, and lead to syncytia formation.)
As time goes by, mutation in the gene coding for gp120 enables some of the HIV to become dual tropic and able to infect both macrophages via the CCR5 chemokine receptor found on these cells, and T4-lymphocytes via the CCR5 and CXCR4 chemokine receptors found on these cells. (Duel-tropic HIV are also called R5X4 viruses since they adsorb to both the chemokine receptors CCR5 and CXCR4.)
Later during the course of HIV infection, most of the viruses have mutated their gp120 to become T- tropic (see Figure \(\PageIndex{2}\)) and infect primarily mature dendritic cells and T4-lymphocytes by way of CD4 and the CXCR4 co-receptors found on these cells. (T-tropic HIV are also called X4 viruses since they adsorb to the chemokine receptor CXCR4.) T-tropic HIV require high levels of CD4 molecules expressed on the surface of the host cell for infection. As mentioned, these T-tropic strains of HIV are faster-replicating and more virulent, and cause formation of syncytias and begin the cycles of T4-lymphocyte destruction.
HIV infecting microglia cells in the brain appear to bind to a CD4 molecule and a chemokine receptor called CCR3 found on these macrophage-like cells.
2. Viral Entry into the Host Cell
As mentioned above under adsorption, the binding of a portion or domain of the HIV surface glycoprotein gp120 to a CD4 molecule on the host cell induces a change in shape that brings the chemokine receptor binding domains of the gp120 into proximity with the host cell chemokine receptor. This, in turn, brings about a conformational change that exposes a previously buried portion of the transmembrane glycoprotein gp41 enabling the viral envelope to fuse with the host cell membrane (see Figure \(\PageIndex{5}\) and Figure \(\PageIndex{6}\)). After fusion of the viral envelope with the host cell cytoplasmic membrane, the genome-containing protein core of the virus enters the host cell's cytoplasm. (Occasionally the virus enters by endocytosis, after which the viral envelope fuses with the endocytic vesicle releasing the genome-containing core into the cytoplasm.)
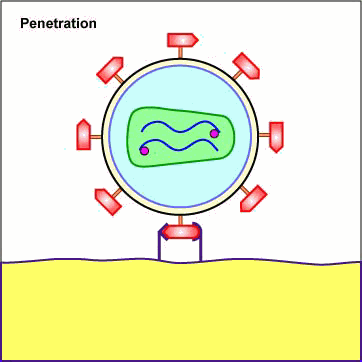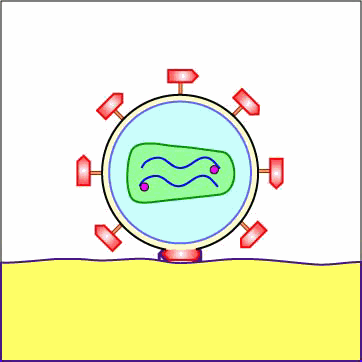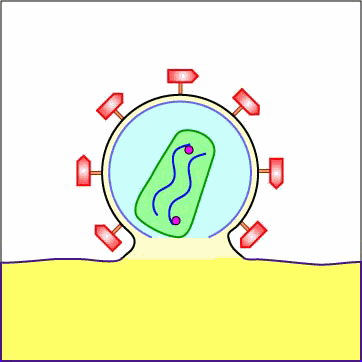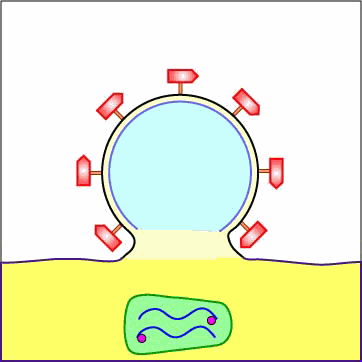
Animation: Penetration of HIV into Host Cell. The binding of a portion or domain of the HIV surface glycoprotein gp120 to a CD4 molecule on the host cell induces a change in shape that brings the chemokine receptor binding domains of the gp120 into proximity with the host cell chemokine receptor. This, in turn, brings about a conformational change that exposes a previously buried portion of the transmembrane glycoprotein gp41 enabling the viral envelope to fuse with the host cell membrane. After fusion of the viral envelope with the host cell cytoplasmic membrane, the genome-containing protein core of the virus enters the host cell's cytoplasm.
3. Viral Movement to the Site of Replication within the Host Cell and Production of a Provirus
During uncoating, the single-stranded RNA genomes within the core or capsid of the virus are released into the cytoplasm. HIV now uses the enzyme reverse transcriptase, associated with the viral RNA genome, to make a DNA copy of the RNA genome. (Normal transcription in nature is when the DNA genome is transcribed into mRNA which is then translated into protein. In HIV reverse transcription, RNA is reverse-transcribed into DNA.)
Reverse transcriptase has three enzyme activities:
- It has RNA-dependent DNA polymerase activity that copies the viral (+) RNA into a (-) viral complementary DNA (cDNA);
- It has ribonuclease activity that degrades the viral RNA during the synthesis of cDNA; and
- It has DNA-dependent DNA polymerase activity that copies the (-) cDNA strand into a (+) DNA to form a double-stranded DNA intermediate.
As the cDNA is being synthesized off of the RNA template the ribonuclease activity degrades the viral RNA genome (see Figure \(\PageIndex{7}\)A, Figure \(\PageIndex{7}\)B, and Figure \(\PageIndex{7}\)C). The reverse transcriptase then makes a complementary DNA strand to form a double-stranded viral DNA intermediate (see Figure \(\PageIndex{7}\)D).

Animation: HIV Copying RNA into DNA with Reverse Transcriptase. The single-stranded RNA genomes are released from the capsid. HIV uses the enzyme reverse transcriptase to transcribe its RNA genome into single-stranded DNA. As the DNA is being made, the RNA genome is degraded by an RNase. The reverse transcriptase then synthesizes a complementary DNA strand to produce a double-stranded DNA intermediate that enters the infected host cell's nucleus.
A viral enzyme called integrase then binds to the double-stranded viral DNA intermediate, transports it through the pores of the host cell's nuclear membrane, and inserts into one of the host cell's chromosomes to form a provirus (see Figure \(\PageIndex{8}\)A and Figure \(\PageIndex{8}\)B).
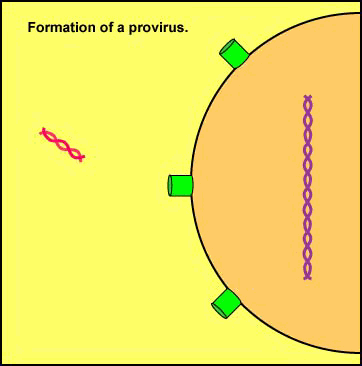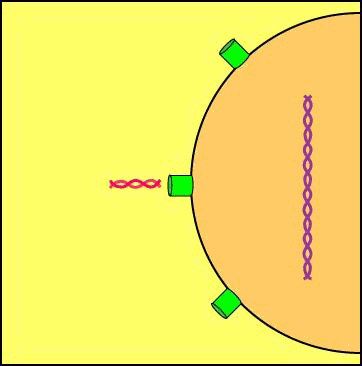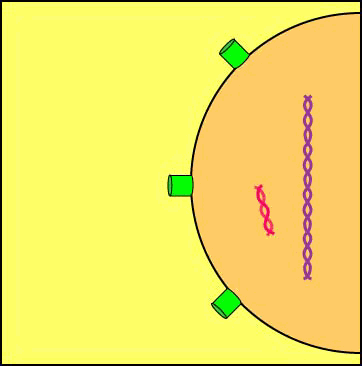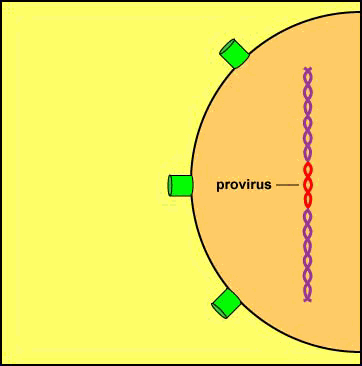
Animation: Formation of a Provirus. An HIV enzyme called integrase is used to insert the HIV double-stranded DNA intermediate into the DNA of a host cell's chromosome. HIV is now a provirus.
After integration, the HIV proviral DNA can exist in either a latent or productive state, which is determined by genetic factors of the viral strain, the type of cell infected, and the production of specific host cell proteins.
The majority of the proviral DNA is integrated into the chromosomes of activated T4-lymphocytes. These generally comprise between 93% and 95% of infected cells and are productively infected, not latently infected. However, a small percentage of HIV-infected memory T4-lymphocytes persists in a resting state because of a latent provirus. These, along with infected monocytes, macrophages, and dendritic cells, provide stable reservoirs of HIV capable of escaping host defenses and antiretroviral chemotherapy.
4. Replication of HIV within the Host Cell
The vast majority of T4-lymphocytes, which are productively infected, immediately begin producing new viruses. In the case of the small percentage of infected, resting memory T4-lymphocytes, before replication can occur, the HIV provirus must become activated. This is accomplished by such means as antigenic stimulation of the infected T4-lymphocytes or their activation by factors such as cytokines, endotoxins, and superantigens.
Following activation of the provirus, molecules of (+) mRNA are transcribed off of the (-) proviral DNA strand by the enzyme RNA polymerase II. Once synthesized,HIV mRNA goes through the nuclear pores into the rough endoplasmic reticulum to the host cell's ribosomes where it is translated into HIV structural proteins, enzymes, glycoproteins, and regulatory proteins(see Figure \(\PageIndex{3}\)).
A 9 kilobase mRNA is formed that is used for three viral functions:
a. Synthesis of Gag polyproteins (p55). These polyproteins will eventually be cleaved by HIV proteases to become HIV matrix proteins (MA; p17), capsid proteins (CA; p24), and nucleocapsid proteins (NC, p7). See Figure \(\PageIndex{9}\)A and Figure \(\PageIndex{9}\)B.
b. Synthesis of Gag-Pol polyproteins (p160). These polyproteins will eventually be cleaved by HIV proteases to become HIV matrix proteins (MA; p17), capsid proteins (CA; p24), proteinase molecules (protease or PR; p10), reverse transcriptase molecules (RT; p66/p51), and integrase molecules (IN; p32). See Figure \(\PageIndex{9}\)C and Figure \(\PageIndex{9}\)D.
c. During maturation, these RNA molecules also become the genomes of new HIV virions.
The 9kb mRNA can also be spliced to form a 4kb mRNA and a 2kb mRNA.
The 4kb mRNA is used to:
a. Synthesize the Env polyproteins (gp160). These polyproteins will eventually be cleaved by proteases to become HIV envelope glycoproteins gp120 and gp41. See Figure \(\PageIndex{9}\)E and Figure \(\PageIndex{9}\)F.
b. Synthesize 3 regulatory proteins called vif, vpr, and vpu.
The 2kb mRNA is used to synthesize 3 regulatory proteins known as tat, rev, and naf.
5. Viral Assembly or Maturation within the Host Cell and Release from the Host Cell
Assembly of HIV virions begins at the plasma membrane of the host cell. Maturation occurs either during the budding of the virion from the host cell or after its release from the cell.
- Transmission electron micrograph of HIV budding from a T4-lymphocyte; courtesy of Dennis Kunkel's Microscopy.
Prior to budding, the Env polyprotein (gp160) goes through the endoplasmic reticulum and is transported to the Golgi complex where it is cleaved by a protease (proteinase) and processed into the two HIV envelope glycoproteins gp41 and gp120. These are transported to the plasma membrane of the host cell where gp41 anchors the gp120 to the membrane of the infected cell. See Figure \(\PageIndex{10}\)A, Figure \(\PageIndex{10}\)B, Figure \(\PageIndex{10}\)C, and Figure \(\PageIndex{10}\)D.
|
GIF animation showing maturation of gp41 and gp120.
|
The Gag (p55) and Gag-Pol (p160) polyproteins also associate with the inner surface of the plasma membrane along with the HIV genomic RNA as the forming virion begins to bud from the host cell.
During maturation, HIV proteases (proteinases) will cleave the remaining polyproteins into individual functional HIV proteins and enzymes such as matrix proteins (MA; p17), capsid proteins (CA; p24), reverse transcriptase molecules (RT; p66/p51), and integrase molecules (IN; p32).. See Figure \(\PageIndex{10}\)E, Figure \(\PageIndex{10}\)F, Figure \(\PageIndex{10}\)G, and Figure \(\PageIndex{10}\)H.
a. The Gag polyproteins (p55) will be cleaved by HIV proteases to become HIV matrix proteins (MA; p17), capsid proteins (CA; p24), and nucleocapsid proteins (NC, p7 and p6).
b. The Gag-Pol polyproteins (p160) will be cleaved by HIV proteases to become HIV matrix proteins (MA; p17), capsid proteins (CA; p24), proteinase molecules (protease or PR; p10), reverse transcriptase molecules (RT; p66/p51), and integrase molecules (IN; p32).
The various structural components then assemble to produce a mature HIV virion.
|
GIF animation showing maturation of of HIV.
|
6. Reinfection
Free viruses now infect new susceptible body cells. HIV can also be transmitted by cell-to-cell contact. This can occur when an infected cell with gp120 on its cytoplasmic membrane attaches to CD4 molecules and chemokine receptors on the surface of an uninfected cell. The cells then fuse (see Figure \(\PageIndex{11}\) and Figure \(\PageIndex{12}\)).
|
Excellent Animation Summarizing the Life Cycle of HIV Courtesy of HHMI's Biointeractive. |
|
YouTube Animation Illustrating Reproduction of HIV.
Courtesy of 3D Medical Animations Library, Dr. Rufus Rajadurai |
- State the role(s) of gp120 and gp41 in the life cycle of HIV.
- Why does HIV primarily infect T4-lymphocytes, macrophages, and dendritic cells?
- How do antiretroviral drugs that bind to HIV-encoded protease help to reduce the number of HIV in the body.
- If one could destroy all of the infected white blood cells in a person infected with HIV and then reconstitute the cells by giving a bone marrow transplant from a person homozygous for a deletion mutation in their gene coding for the chemokine receptor CCR5 (he or she can not make CCR5 molecules), describe how this might prevent HIV infection in the person receiving the transplant.
|
Medscape article on infections associated with organisms mentioned in this Learning Object. Registration to access this website is free.
|
Summary
- During adsorption, an envelope glycoprotein on the surface of HIV called gp120 must adsorbs to both a CD4 molecule and then a chemokine receptor found on the surface of only certain types of certain human cells such as T4-lymphocytes, monocytes, macrophages, and dendritic cells.
- Following adsorption, glycoprotein gp41 enabling the viral envelope to fuse with the host cell membrane, allowing the nucleocapsid of the virus enters the host cell's cytoplasm.
- During uncoating, the single-stranded RNA genomes within the capsid of the virus are released into the cytoplasm and HIV now uses the enzyme reverse transcriptase to make a single-stranded DNA copy of its single-stranded RNA genome. The reverse transcriptase then makes a complementary DNA strand to form a double-stranded viral DNA intermediate.
- A viral enzyme called integrase then binds to the double-stranded viral DNA intermediate, transports it through the pores of the host cell’s nuclear membrane, and inserts into one of the host cell's chromosomes to form a provirus.
- Following activation of the provirus, molecules of mostly polycistronic mRNA are transcribed off of the proviral DNA strand, go through the nuclear pores into the rough endoplasmic reticulum where it is translated by host cell's ribosomes HIV structural proteins, enzymes, glycoproteins, and regulatory proteins.
- Polyproteins translated from polycistronic mRNAs must be cleaved into function proteins by HIV protease enzymes.
- The two HIV envelope glycoproteins gp41 and gp120 are transported to the plasma membrane of the host cell where gp41 anchors the gp120 to the membrane of the infected cell. HIV obtains its envelope from the plasma membrane by budding.
- Most maturation occurs either during the budding of the virion from the host cell or after its release from the cell.
Questions
Study the material in this section and then write out the answers to these questions. Do not just click on the answers and write them out. This will not test your understanding of this tutorial.
- Describe how the retrovirus HIV-1 accomplishes each of the following steps during its life cycle. (Include the following key words in your description: gp120, CD4, chemokine receptors, gp41, capsid, RNA genome, reverse transcriptase, double-stranded DNA intermediate, provirus, polyproteins, proteases, and budding.)
- viral attachment or adsorption to the host cell (ans)
- viral entry into the host cell (ans)
- viral movement to the site of replication within the host cell and production of a provirus. (ans)
- viral replication within the host cell (ans)
- viral assembly or maturation within the host cell and release from the host cell (ans)
- Name 3 types of cells HIV primarily infects and briefly explain why. (ans)
- HIV possesses a genome of RNA. How then is HIV able to insert into the DNA of host cells and form a provirus? (ans)
- Multiple Choice (ans)


