8.8: rII Locus of T4
- Page ID
- 4859
Mapping Within A Gene: the RII Locus
T2 and its close relative T4 are viruses that infect the bacterium E. coli. The infection ends with destruction (lysis) of the bacterial cell so these viruses are examples of bacteriophages ("bacteria eaters"). They have been enormously useful in genetic studies because:
- Viruses of two (or more) different genotypes can simultaneously infect a single bacterium.
- The DNA molecules of one of the infecting viruses can recombine with that of another forming recombinant molecules.
- The huge number of viruses released from a huge number of bacterial hosts enables even rare recombination events to be detected.
Another page — Bacteriophage Genetics — describes how T2 is used to map the order and relative spacing of genes on the single circular molecule of DNA that is the virus's genome. Here let us see how T4 can be used to detect mutations within a single gene and speed up the process of mapping these point mutations by the use of deletion mutants.
Detecting Mutation within a Single Gene
In Bacteriophage Genetics, we examined mutation of a gene designated r, for "rapid lysis". It turned out that actually there are three different gene loci — rI, rII, and rIII — mutations in any one of which produced a rapid-lysis phenotype. But, in addition, there were many mutations found in each of these. Could wild-type virus be formed by recombination between mutations within the same gene? Seymour Benzer decided to find out.
As we saw in Bacteriophage Genetics, the recombination frequency between different genes is low (on the order of 10-2). One would expect that recombination frequencies between mutations in a single gene would be far lower (10-4 or less). Fortunately Benzer could exploit a phenomenon to enable him to detect such rare events: rII mutants as well as wild-type T4 can infect and complete their life cycle in a strain of E. coli designated B. However, while rII mutants can infect a strain of E. coli designated K, they cannot complete their life cycle in strain K. Wild-type T4 can.
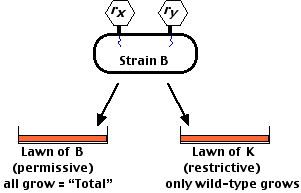
The procedure was to infect strain B in liquid culture with two mutants to be tested (designated here as rx and ry). After incubation, these were plated on a lawn of:
- strain B — which supports the growth of all viruses thus giving the total number of viruses liberated.
- strain K — on which only wild-type viruses can grow.
The recombination frequency between any pair of mutations is calculated as
\[\text{Recombination Frequency} = \dfrac{2 \times \text{number of wild-type plaques (strain K plaques)}}{\text{total number of plaques (on strain B)}}\]
You have to double the number found on strain K because you only see one-half the recombinants — the other half consists of double mutants. Using this technique, Benzer eventually found some 2000 different mutations in the rII gene. The recombination frequency between some pairs of these was as low as 0.02.
- The T4 genome has 160,000 base pairs of DNA extending over ~1,600 centimorgans (cM).
- So 1 cM ≅ 100 base pairs
- So 0.02 cM represents a pair of adjacent nucleotides.
- From these data, Benzer concluded that the
- smallest unit of mutation and
- the smallest unit of recombination
In other words, these mutations represent a change in a single base pair — we call these point mutations. Recombination between two molecules of DNA can occur at any pair of nucleotides.
Mapping Point Mutations Within A Gene
The relative order and spacing of any two point mutations in a single gene like rII can be done using the procedure describe in Bacteriophage Genetics. But with some 2000 different mutations to test, the process would be tremendously time-consuming. (Even using the procedure to be described now, Benzer spent some 10 years on the project.) Benzer was able to speed up the mapping process by taking advantage of the discovery that some of his mutants did not have point mutations but deletions instead. In contrast to the properties of T4 viruses with point mutations, T4 viruses with deletions in rII showed no recombination with other rII mutants or any other genes for that matter. Moreover, these deletions never back-mutated.
Deletion Mapping
Deletions can be mapped by the same procedure used for point mutations. Simply cross pairs of deletion mutants and see if they produce progeny that can grow on E. coli strain K. Here is a hypothetical example. Each of 6 strains of deletion mutants are crossed with each of the others.
| Strains | 1 | 2 | 3 | 4 | 5 | 6 | |
| 1 | 0 | 0 | + | 0 | 0 | 0 | 1 and 3 do not overlap |
| 2 | 0 | + | + | 0 | 0 | must shift 4 away from 2 | |
| 3 | 0 | + | + | 0 | 6 must extend under 3 | ||
| 4 | 0 | + | + | right-hand end of 4 must be removed from over 6 | |||
| 5 | 0 | + | left-hand end of 6 must not overlap 5 but must continue to overlap 2. ∴ shorten right-hand end of 5 | ||||
| 6 | 0 |
From the results, one can draw a map showing the order and relative size of the deletions.
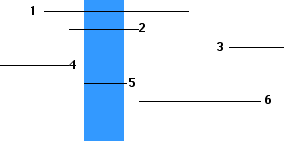
With such a deletion map, one can now quickly map the location of point mutations by coinfecting each of the different deletion strains (here 1–6) with the mutant strain ("x").
There is no longer any need to count plaques; simple see whether there is growth or not.
| Coinfect with strain | 1 | 2 | 3 | 4 | 5 | 6 |
| and mutant "x" Results → | 0 | 0 | + | + | 0 | + |
From these results, we learn that the point mutation "x" is located on the T4 DNA within the region shown above in blue.
Complementation
As we saw above, rapid lysis (r) mutants were found that mapped to three different regions of the T4 genome: rI, rII, and rIII. This meant that those in different regions were not alleles of the same gene and more than one gene product participated in the lysis function. Even within one "locus", rII, there turned out to be two different stretches of DNA both of which were needed intact for the lysis function. This was revealed by the complementation test that Benzer used. In this test,
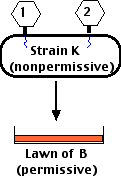
- E. coli strain K (which rII mutants can infect but not complete their life cycle) — growing in liquid culture — was
- coinfected with two different rII mutants (shown in the figure as "1" and "2").
Note that this procedure differs from the earlier one (recombination) in that the nonpermissive E. coli K is used for the initial infection (not strain B as before). Neither strain rII"1" nor strain rII"2" is able to grown in E. coli K. But if the lost function in rII"1" is NOT the same as the lost function in rII"2", then
- each should be able to produce the gene product missing in the other — complementation — and
- living phages will be produced. (Again, there is no need to count plaques; simply see if they are formed or not.)
| Mutant strains | 1 | 2 | 3 | 4 | 5 |
| 1 | 0 | 0 | + | 0 | + |
| 2 | 0 | + | 0 | + | |
| 3 | 0 | + | 0 | ||
| 4 | 0 | + | |||
| 5 | 0 |
From these results, you can deduce that these 5 rII mutants fall into two different complementation groups, which Benzer designated A (containing strains 1, 2, and 4) and B (containing strains 3 and 5). Later work showed that the function of rII depended on the polypeptide products encoded by two adjacent regions (A and B) of rII (perhaps acting as a heterodimer). In terms of function, then, both A and B qualify as independent genes. In coinfections by two mutant strains,
- If either A or B is mutated on the same DNA molecule ("cis"), there is no function while
- if A is mutated in one DNA molecule and B in the other ("trans"), function is restored.
Complementation, then, is the ability of two different mutations to restore wild-type function when they are in the "trans" (on different DNA molecules), but not when they are in "cis" (on the same DNA molecule). Benzer coined the term cistron for these genetic units of function. But today, we simply modify earlier concepts of the "gene" to fit this operational definition.


