E2. The Biggest Ribozyme - The Ribosome
- Page ID
- 5544
Protein synthesis from a mRNA template occurs on a ribosome, a nanomachine composed of proteins and ribosomal RNAs (rRNA). The ribosome is composed of two very large structural units. The smaller unit (termed 30S and 40S in bacteria and eukaryotes, respectively) coordinates the correct base pairing of the triplet codon on the mRNA with another small adapter RNA, transfer or tRNA, that brings a covalently connected amino acid to the site. Peptide bond formation occurs when another tRNA-amino acid molecule binds to an adjacent codon on mRNA. The tRNA has a cloverleaf tertiary structure with some intrastranded H-bonded secondary structure. The last three nucleotides at the 3' end of the tRNA are CpCpA. The amino acid is esterified to the terminal 3'OH of the terminal A by a protein enzyme, aminoacyl-tRNA synthetase.
Covalent amide bond formation between the second amino acid to the first, forming a dipeptide, occurs at the peptidyl transferase center, located on the larger ribosomal subunit (50S and 60S in bacteria and eukaryotes, respectively). The ribosome ratchets down the mRNA so the dipeptide-tRNA is now at the the P or Peptide site, awaiting a new tRNA-amino acid at the A or Amino site. The figure below shows a schematic of the ribosome with bound mRNA on the 30S subunit and tRNAs covalently attached to amino acid (or the growing peptide) at the A and P site, respectively.
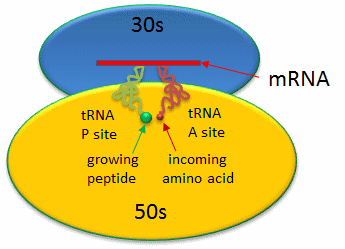
Figure: Prokaryotic Ribosme - P and A sites
A likely mechanism (derived from crystal structures with bound substrates and transition state analogs) for the formation of the amide bond between a growing peptide on the P-site tRNA and the amino acid on the A-site tRNA is shown below. Catalysis does not involve any of the ribosomal proteins (not shown) since none is close enough to the peptidyl transferase center to provide amino acids that could participate in general acid/base catalysis, for example. Hence the rRNA must acts as the enzyme (i.e. it is a ribozyme). Initially it was thought that a proximal adenosine with a perturbed pKa could, at physiological pH, be protonated/deprotonated and hence act as a general acid/base in the reaction. However, none was found. The most likely mechanism to stabilize the oxyanion transition state at the electrophilic carbon attack site is precisely located water, which is positioned at the oxyanion hole by H-bonds to uracil 2584 on the rRNA. The cleavage mechanism involves the concerted proton shuffle shown below. In this mechanism, the substrate (Peptide-tRNA) assists its own cleavage in that the 2'OH is in position to initiate the protein shuttle mechanism. (A similar mechanism might occur to facilitate hydrolysis of the fully elongated protein from the P-site tRNA.) Of course all of this requires perfect positioning of the substrates and isn't that what enzymes do best? The main mechanisms for catalysis of peptide bond formation by the ribosome (as a ribozyme) are intramolecular catalysis and transition state stabilization by the appropriately positioned water molecule.
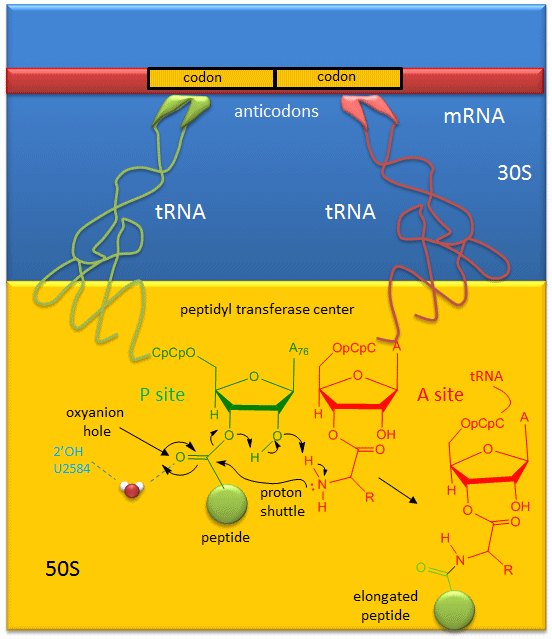
Figure: Mechanism Peptide Bond Formation by the Ribosome
The crystal structure of the eukaryotic ribosome has recently been published (Ben-Shem et al). It is significantly larger (40%) with mass of around 3x106 Daltons. The 40S subunit has one rRNA chain (18) and 33 associated proteins, while the larger 60S subunit has 3 rRNA chains (25S, 5.8S and 5S) and 46 associated proteins. The larger size of the eukaryotic ribosome facilitates more interactions with cellular proteins and greater regulation of cellular events. The Jmol structure of a bacterial 70S ribosome showing mRNA and tRNA interactions is shown below.
![]() Jmol: Updated 70 S Ribosome from Thermus Thermophilus showing mRNA and tRNA interactions Jmol14 (Java) | JSMol (HTML5) Not done; Fix
Jmol: Updated 70 S Ribosome from Thermus Thermophilus showing mRNA and tRNA interactions Jmol14 (Java) | JSMol (HTML5) Not done; Fix


