1.21: Eukaryotes
- Page ID
- 8235
Eukaryotes
Living things fall into three large groups: Archaea, Bacteria, and Eukarya. The first two groups include non-nucleated cells, and the third contains all eukaryotes. The comparative biology- particularly the comparative genomics- of extant organisms and the limited fossil record provide some insight into the history of Eukarya.
The earliest fossils found appear to be Bacteria, most likely cyanobacteria. They are about 3.5 billion years old and are recognizable because of their colonial nature (formation of mats) and the preservation of their pigments (or rather, degraded forms of these). The earliest record of what are thought to be the chemical remnants of eukaryotes- the presence of a family of fatty acids termed sterols, currently found only in eukaryotes- were found in oil deposits thought to be approximately 2.7 billion years old.
Characteristics of Eukaryotes
Data from DNA sequencing led biologists to the conclusion that living eukaryotes are all descendants of a single common ancestor. Mapping the characteristics found in all major groups of eukaryotes reveals that the following characteristics almost certainly were present in their last common ancestor, because these characteristics are present in at least some of the members of each major lineage.
- Cells with nuclei surrounded by a nuclear envelope with nuclear pores. This is the single characteristic that is both necessary and sufficient to define an organism as a eukaryote. All extant eukaryotes have cells with nuclei.
- Mitochondria. Some extant eukaryotes have very reduced remnants of mitochondria in their cells, whereas other members of their lineages have “typical” mitochondria.
- A cytoskeleton containing the structural and motility components called actin microfilaments and microtubules. All extant eukaryotes have these cytoskeletal elements.
- Flagella and cilia, organelles associated with cell motility. Some extant eukaryotes lack flagella and/or cilia, but they are descended from ancestors that possessed them.
- Chromosomes, each consisting of a linear DNA molecule coiled around basic (alkaline) proteins called histones. The few eukaryotes with chromosomes lacking histones clearly evolved from ancestors that had them.
- Mitosis, a process of nuclear division wherein replicated chromosomes are assorted into daughter cells using elements of the cytoskeleton.
- Sex, a process of genetic recombination unique to eukaryotes in which diploid nuclei at one stage of the life cycle undergo meiosis to yield haploid nuclei and subsequent karyogamy, a stage where two haploid nuclei fuse together to create a diploid zygote nucleus.
Endosymbiosis and the Evolution of Eukaryotes
In order to understand eukaryotic organisms fully, it is necessary to understand that all extant eukaryotes are descendants of a chimeric organism that was a composite of a host cell and the cell(s) of an alpha-proteobacterium that “took up residence” inside it. This major theme in the origin of eukaryotes is known as endosymbiosis, one cell engulfing another such that the engulfed cell survives and both cells benefit. Over many generations, a symbiotic relationship can result in two organisms that depend on each other so completely that neither could survive on its own. Endosymbiotic events likely contributed to- if not defined- the origin of the last common ancestor of today’s eukaryotes and to later diversification in certain lineages of eukaryotes.
Bacterial and Archaeal Contributions
Many important metabolic processes arose in bacteria and archaea, and some of these, such as nitrogen fixation, are never found in eukaryotes. The process of aerobic respiration is found in all major lineages of eukaryotes, and it is localized in the mitochondria. Aerobic respiration is also found in many lineages of bacteria and archaea, but it is not present in all of them, and many forms of evidence suggest that the ancestors of many existing anaerobic microbes never had the ability to perform aerobic respiration.
While today’s atmosphere is about one-fifth molecular oxygen (O2), geological evidence shows that it originally lacked O2. Without oxygen, aerobic respiration cannot occur, and living things would have relied on fermentation or anaerobic respiration instead to derive energy from their fuel. At some point about 3.5 billion years ago, the group of bacteria that gave rise to cyanobacteria used water as the electron source and released O2 as a waste product. Given the highly reactive nature of O2, this molecule was toxic to the early biosphere (reacting with Fe++, H2, and other reduced compounds that acted as electron sources for many living things) and more directly to living things (oxidizing and therefore inactivating many enzymes).
Eventually, the amount of photosynthetic oxygen built up in some environments to levels that posed a risk to living organisms. Various metabolic processes evolved that detoxified oxygen though its controlled reduction, one of which, aerobic respiration, also generated high levels of ATP. It became widely present among microbes, including a group we the call alpha-proteobacteria. Organisms that did not acquire aerobic respiration or other O2-protective mechanisms had to remain in oxygen-free environments. Originally, oxygen-rich environments were likely localized around places where cyanobacteria were active, but geological evidence shows that by about 2 billion years ago, oxygen was building up to higher concentrations in the atmosphere and the upper oceans. Oxygen levels similar to today’s levels only arose within the last 700 million years.
Recall that the first fossils that we believe to be eukaryotes date to about 2 billion years ago, so they appeared as oxygen levels were increasing. Also, recall that all extant eukaryotes descended from an ancestor with mitochondria. This organelle was first observed by microscopists in the late 1800s, where they appeared to be somewhat worm-shaped structures that seemed to be moving around in the cell. Some early observers suggested that they might be bacteria living inside host cells, but this hypothesis remained unknown or rejected in most scientific communities.
Endosymbiotic Theory
As cell biology developed in the twentieth century, it became clear that mitochondria were the organelles responsible for producing ATP using aerobic respiration. In the 1960s, American biologist Lynn Margulis developed the endosymbiotic theory, which states that eukaryotes may have been a product of one cell engulfing another, with one surviving within another, and evolving over time until the separate cells were no longer recognizable as such. In 1967, Margulis introduced new work on the theory and substantiated her findings through microbiological evidence. Although Margulis’ work initially was met with resistance, this once-revolutionary hypothesis is now widely accepted, with work progressing on uncovering the steps involved in this evolutionary process and the key players involved. Much still remains to be discovered about the origins of the cells that now make up the cells in all living eukaryotes.
It has become clear that many of our nuclear genes and the molecular machinery responsible for DNA replication, repair, recombination and gene expression appear closely related to those in Archaea. On the other hand, the organelles and genes responsible for many energy-harvesting processes had their origins in bacteria. These genes are most similar to those found in the Alpha-proteobacteria, a group that includes species symbiotic with plants, disease organisms that can infect humans via ticks, and many free-living species that use light for energy. Although our closest ancestor (on our bacterial side) is still a matter of debate, DNA sequence analysis suggests that today's mitochondria are most closely related to the very ancient and successful SAR11 clade of this bacterial group, a clade of largely marine bacteria that often participate in interspecific cell colonization events. Much remains to be clarified about how this relationship occurred; this continues to be an exciting field of discovery in biology. For instance, it is not known whether the endosymbiotic event that led to mitochondria occurred before or after the host cell had a nucleus. The details of the process of endosymbiosis are also unclear- the "Hydrogen Hypothesis" suggests this partnership originated as an external symbiosis between a hydrogen-seeking methanogen and a hydrogen-excreting, metabolically flexible alpha-proteobacterium. This hypothesis is supported by the fact that many eukaryotes retain a mitochondria-like organelle termed a hydrogenosome, which performs a particularly efficient type of fermentation of pyruvate which releases H2 as a waste product.
Introduction to eukaryotic cells
Although it is quite possible that the common ancestor of all eukaryotes- the ancestor that originally cohabited with an aerobically respiring alpha-proteobacterium, did not have a nucleus, today, by definition, eukaryotic cells are cells that contain a membrane-bound nucleus, a structural feature that is not present in bacterial or archaeal cells. In addition to the nucleus eukaryotic cells are characterized by numerous membrane-bound organelles such as the endoplasmic reticulum, Golgi apparatus, chloroplasts (sometimes), mitochondria (or related organelles, such as hydrogenosomes), and others.
In previous sections we began to consider the Design Challenge of making cells larger than a small bacterium - more precisely, growing cells to sizes where, in the eyes of natural selection, relying on diffusion of substances for transport through a highly viscous cytosol comes with inherent functional trade-offs that offset most selective benefits of getting larger. In the lectures and readings on bacterial cell structure we discovered some morphological features of large bacteria that allow them to effectively overcome diffusion-limited size barriers (e.g. filling the cytoplasm with a large storage vacuole maintains a small volume for metabolic activity that remains compatible with diffusion-driven transport). As we transition our focus to eukaryotic cells we want you to approach the study by constantly returning to the Design Challenge. We will cover a large number of sub-cellular structures that are unique to eukaryotes and you will be expected to know the names of these structures or organelles, to associate them with one or more functions, and to identify the structure on a canonical cartoon representation of a eukaryotic cell. This memorization exercise is necessary but not sufficient. We will also ask you to think about some of the functional and evolutionary costs and benefits (trade-offs) of evolving eukaryotic cells and various eukaryotic organelles and how a eukaryotic cell might coordinate the functions of different organelles.
Your instructors will, of course, propose some functional hypotheses for you to consider that address these broader points. Our hypotheses may sometimes come in the form of statements like "Thing A exists because of rationale B." If we are completely honest, however, in many cases we don't actually know all of the selective pressures that led to the creation or maintenance of certain cellular structures and the likelihood that one explanation will fit all cases in biology is slim. The causal linkage/relationship implied by the use of terms like because should be treated as good hypotheses rather than objective concrete undisputed factual knowledge. We want you to understand these hypotheses and be able to discuss the ideas presented in class, but we also want you to indulge your own curiosity and to begin thinking about these ideas critically yourself. Try using the Design Challenge rubric to explore some of your ideas. In the following we will try to seed questions to encourage this activity.
In the section immediately below, we will discuss the various organelles found in various eukaryotic cells. Some organelles are found in all eukaryotic cells, and some are not. These organelles are visible through the light and/or electron microscopy long before people had any idea what their functions were. In many cases their functions are still being debated today.
As always, I encourage you to focus- for exams- on topics that are described in lecture.

These figures show the major organelles and other cell components of a typical eukaryotic plant cell. The plant cell has a cell wall, chloroplasts, plastids, and a central vacuole—structures not found in animal cells. Plant cells do not have lysosomes- organelles for the digestion of phagocytosed food particles. Why not?
The Plasma Membrane
Like bacteria and archaea, eukaryotic cells have a plasma membrane, a phospholipid bilayer with embedded proteins that separates the internal contents of the cell from its surrounding environment. The plasma membrane controls the passage of organic molecules, ions, water, and oxygen into and out of the cell. Wastes (such as carbon dioxide and ammonia) also leave the cell by passing through the plasma membrane (if hydrophobic) or with the help of protein transporters.
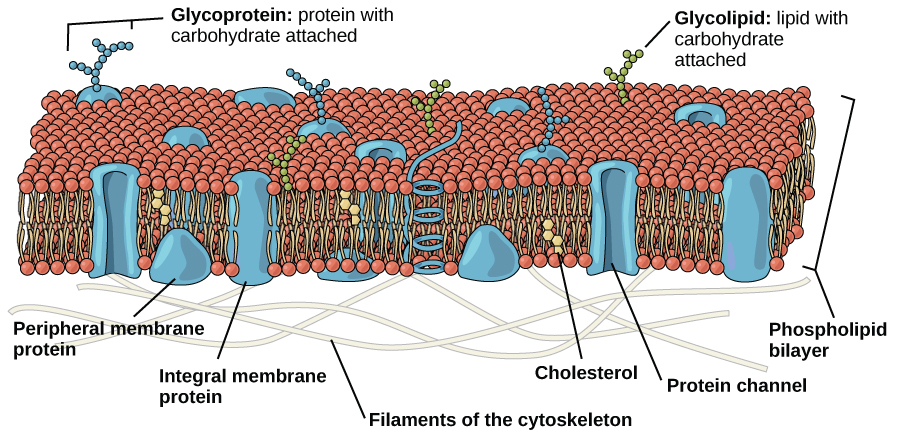
The eukaryotic plasma membrane is a phospholipid bilayer with proteins and cholesterol embedded in it.
As discussed in the context of bacterial cell membranes, the plasma membranes of eukaryotic cells may also adopt unique conformations. For instance, the plasma membrane of cells that in multicellular organisms specialize in absorption are often folded into fingerlike projections called microvilli (singular = microvillus); (see figure below). The "folding" of the membrane into microvilli effectively increases the surface area for absorption while minimally impacting the cytosolic volume. Such cells can be found lining the small intestine, the organ that absorbs nutrients from digested food.
An aside: People with celiac disease have an immune response to gluten, a protein found in wheat, barley, and rye. The immune response damages microvilli. As a consequence, afflicted individuals have an impaired ability to absorb nutrients. This can lead to malnutrition, cramping, and diarrhea.

Microvilli, shown here as they appear on cells lining the small intestine, increase the surface area available for absorption. These microvilli are only found on the area of the plasma membrane that faces the cavity from which substances will be absorbed. (credit "micrograph": modification of work by Louisa Howard)
The Cytoplasm
The cytoplasm refers to the entire region of a cell between the plasma membrane and the nuclear envelope. It is composed of organelles suspended in the gel-like cytosol, the cytoskeleton, and various chemicals (see figure below). Even though the cytoplasm consists of 70 to 80 percent water, it nevertheless has a semi-solid consistency. It's crowded in the cytoplasm. Proteins, simple sugars, polysaccharides, amino acids, nucleic acids, fatty acids, ions and many other water soluble molecules are all competing for space and water.
The Nucleus
Typically, the nucleus is the most prominent organelle in a cell (see figure below) when viewed through a microscope. The nucleus (plural = nuclei) houses the cell’s DNA. Let’s look at it in more detail.

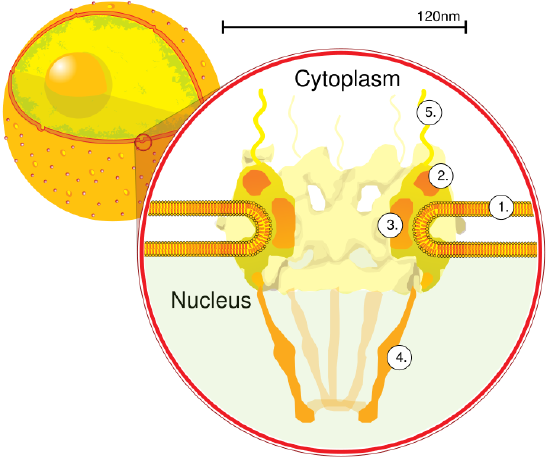
Left- a cross-section of a nucleus. Right- a close-up cross section of a nuclear pore. The nucleus stores chromatin (DNA plus proteins) in a gel-like substance called the nucleoplasm. The nucleolus is a condensed region of chromatin where ribosome synthesis (and the assembly of other ribonucleocomplexes) occurs. The boundary of the nucleus is called the nuclear envelope. It consists of two phospholipid bilayers: an outer membrane and an inner membrane. The nuclear membrane is continuous with the endoplasmic reticulum. Nuclear pores (right) allow substances to enter and exit the nucleus. Smaller molecules simply diffuse through pores, in contrast, the pores are highly selective in terms of allowing passage of macromolecules, allowing only specific proteins and nucleic acids to pass in or out.
The Nuclear Envelope
The nuclear envelope, a structure that constitutes the outermost boundary of the nucleus, is a double-membrane - both the inner and outer membranes of the nuclear envelope are phospholipid bilayers. The nuclear envelope is also punctuated with protein-based pores that control the passage of ions, small molecules, small proteins (as well as larger proteins carrying a "nuclear targeting" sequence) and messenger RNAs, and ribosomal components between the nucleoplasm and cytoplasm. Nuclear pores are not just openings in the nuclear membrane- they are very selective and highly regulated organelles. The nucleoplasm is the semi-solid fluid inside the nucleus, where we find the chromatin and the nucleolus, a condensed region of chromatin where ribosome synthesis occurs.
Chromatin and Chromosomes
To understand chromatin, it is helpful to first consider chromosomes. Chromosomes are structures within the nucleus that are made up of DNA, the hereditary material. You may remember that in bacteria and archaea, DNA is typically organized into one or more circular chromosome(s). In eukaryotes, chromosomes are linear structures. Every eukaryotic species has a specific number of chromosomes in the nuclei of its cells. In humans, for example, the chromosome number is 23, while in fruit flies, it is 4.
Chromosomes are only clearly visible and distinguishable from one another by visible optical microscopy when the cell is preparing to divide and the DNA is tightly packed by proteins into easily distinguishable shapes. When the cell is in the growth and maintenance phases of its life cycle, numerous proteins are still associated with the nucleic acids, but the DNA strands more closely resembles an unwound, jumbled bunch of threads, too thin to be observed by light microscopy. The term chromatin is used to describe chromosomes (the protein-DNA complexes) when they are both condensed and decondensed.
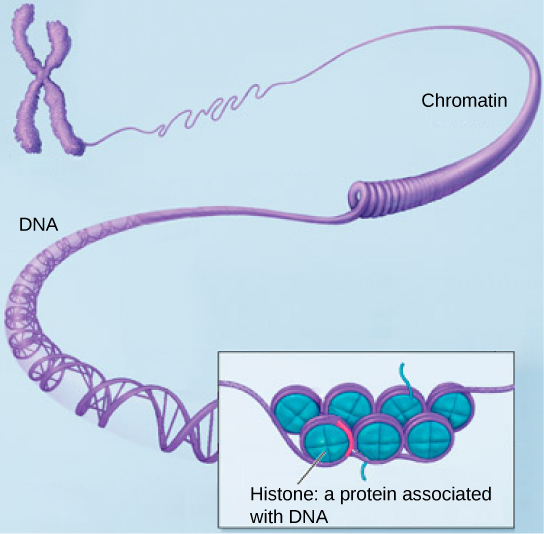
(a) This image shows various levels of the organization of chromatin (DNA and protein). (b) This image shows paired chromosomes. (credit b: modification of work by NIH; scale-bar data from Matt Russell)
The Nucleolus
Some chromosomes have sections of DNA that encode ribosomal RNA. A darkly staining area within the nucleus called the nucleolus (plural = nucleoli) aggregates the ribosomal RNA with associated proteins to assemble the ribosomal subunits that are then transported out through the pores of the nuclear envelope to the cytoplasm. Other nucleoprotein complexes- including complexes involved in splicing of mRNAs, are also assembled in the nucleolus. These remain in the nucleus, where splicing occurs.
Ribosomes
Ribosomes are the cellular structures responsible for protein synthesis. They are present in all cells, prokaryotic and eukaryotic. When viewed through an electron microscope, ribosomes appear either as clusters (polyribosomes) or single, tiny dots that float freely in the cytoplasm. They can also be found attached to the cytoplasmic side of the plasma membrane or the cytoplasmic side of the endoplasmic reticulum and the outer membrane of the nuclear envelope (see cartoon of cell above).
Electron microscopy has shown us that ribosomes, which are large complexes of protein and RNA, consist of two subunits, simply called large and small (figure below). Ribosomes receive their "instructions" for protein synthesis from the nucleus where the DNA is transcribed into messenger RNA (mRNA). The mRNA travels to the ribosomes, which translate the code provided by the sequence of the nitrogenous bases in the mRNA into a specific order of amino acids in a protein. We will cover all of this in greater detail in the section covering the process of translation.
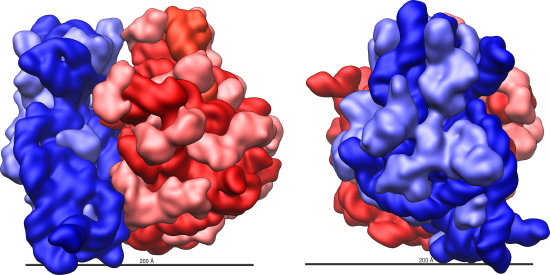
Ribosomes are made up of a large and a small subunit. During protein synthesis, ribosomes use the message encoded on mRNA to assemble amino acids into proteins.
Mitochondria
Mitochondria (singular = mitochondrion) are often called the “powerhouses” or “energy factories” of a cell because they are the primary site of metabolic respiration in eukaryotes. Although some bacteria can perform anawrobic respiration, muoitochondria are limited to aerobic respiration- the can only use O2 as their external terminal electron acceptor. Mitochondria produce ATP via oxidative phosphorylation, hence the use of terms "powerhouse" and/or "energy factory" to describe this organelle. The mitochondrion is also the site of the TCA cycle, which produces the NADH employed in respiration. A little bit of substrate-level phosphorylation also occurs as part of the TCA cycle, producing GTP or ATP. Nearly all mitochondria also possess a small genome that encodes genes whose function are typically restricted to the mitochondrion.
In some cases, the number of mitochondria per cell is "tunable" depending, typically, on energy demand. For instance, muscle cells that are frequently used - that have a higher demand for ATP - may often be found to have a significantly higher number of mitochondria than cells that do not have a high energy load.
The structure of the mitochondria can vary significantly depending on the organism. The typical text-book image, however, depicts mitochondria as oval-shaped organelles with a double inner and outer membrane (see figure below); learn to recognize this generic representation. Both the inner and outer membrane are phospholipid bilayer embedded with proteins that mediate transport across them and catalyze various other biochemical reactions. The inner membrane layer has folds called cristae that increase the surface area into which respiratory chain proteins can be embedded. The interior of the mitochondrion- the space enclosed by both the inner and outer membrane, is called the mitochondrial matrix and contains - among other things - enzymes of the TCA cycle (though some are embedded on the inner membrane). During respiration protons are pumped by respiratory chain complexes from the matrix into a region known as the intermembrane space (between the inner and outer membranes). Once a proton gradient has formed, ATP synthase crosses the inner membrane, allowing protons to move from the intermembrane space to the matrix. This exergonic proton flow is coupled to the formation of ATP from ADP + Pi, a reaction that occurs in the matrix
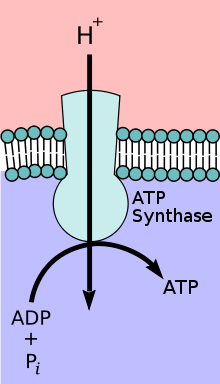
This electron micrograph shows a mitochondrion as viewed with a transmission electron microscope. This organelle has an outer membrane and an inner membrane. The inner membrane contains folds, called cristae, which increase its surface area. The space between the two membranes is called the intermembrane space, and the space inside the inner membrane is called the mitochondrial matrix. ATP synthase crosses the inner membrane (facilitating proton flow across the inner membrane, from the intermembrane space to the matrix). This reaction is coupled to the endergonic conversion of ADP + Pi to ATP in the matrix. (credit: modification of work by Matthew Britton; scale-bar data from Matt Russell)
Possible discussion
Discuss: Processes like glycolysis, lipid biosynthesis and nucleotide biosynthesis all have compounds that feed into the TCA cycle - some of which occurs in the mitochondria. What are some of the functional challenges associated with coordinating processes that have a common set of molecules if the enzymes are sequestered into different cellular compartments?
Peroxisomes
Peroxisomes are small, round organelles enclosed by single membranes. These organelles carry out redox reactions that oxidize and break down fatty acids and amino acids. They also help to detoxify many toxins that may enter the body. Many of these redox reactions release hydrogen peroxide, H2O2, which would be damaging to cells; however, when these reactions are confined to peroxisomes, where enzymes safely break down the H2O2 into oxygen and water. For example, alcohol is detoxified by peroxisomes in liver cells. Glyoxysomes, which are specialized peroxisomes in plants, are responsible for converting stored fats into sugars.
Vesicles and Vacuoles
Vacuoles are membrane-bound sacs that function in storage, breakdown, and containment of small molecules, including toxins. They have a wide variety of functions. They are found in fungi and plants, but only rarely in animals or protists. The large central vacuole of plants helps maintain turgor pressure (among many other functions). While the membranes of vesicles can fuse with either the plasma membrane or other membrane systems within the cell, the membranes of vacuoles do not. Why is that an important distinction?
Vesicles, like vacuoles and many other organelles, are surrounded by a phospholipid bilayer. Vesicles are much smaller than vacuoles and are are generally involved in transport- they are filled with cargo (such as proteins). In some cases the lipids in their membranes- and proteins embedded in the membrane- may be the cargo. Vesicles are too large to spontaneously diffuse in the cell, and so their diffusion is facilitated by motor proteins, which walk along microtubules.
The Endomembrane System
The endomembrane system (endo = “within”) is a group of membranes and organelles in eukaryotic cells that works together to modify, package, and transport lipids and proteins. It includes the nuclear envelope, lysosomes, and vesicles, which we’ve already mentioned, as well as the endoplasmic reticulum and Golgi apparatus, which we will cover shortly. Although not technically within the cell, the plasma membrane is included in the endomembrane system because, as you will see, it interacts with the other endomembranous organelles. The endomembrane system does not include the membranes of either mitochondria or chloroplasts. This is not an arbitrary distinction, as you'll see when we cover protein compartmentalization.
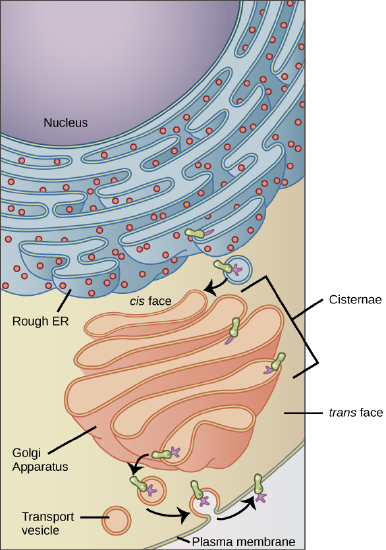
Membrane and secretory proteins are synthesized in the rough endoplasmic reticulum (RER). The RER also sometimes modifies proteins (for example, via glycosylation (= the addition of carbohydrates)). In this illustration, a (green) integral membrane protein in the ER is modified by attachment of a (purple) carbohydrate. Vesicles with the integral protein bud from the ER and fuse with the cis face of the Golgi apparatus. As the protein passes along the Golgi’s cisternae, it is further modified by the addition of more carbohydrates. After its synthesis is complete, it exits as an integral membrane protein of the vesicle that buds from the Golgi’s trans face. When the vesicle fuses with the cell membrane the protein becomes integral portion of that cell membrane. (credit: modification of work by Magnus Manske)
Possible discussion
If a peripheral membrane protein were synthesized in the lumen (inside) of the ER, would it end up on the inside or outside of the plasma membrane?
The Endoplasmic Reticulum
The endoplasmic reticulum (ER) (see figure above) is a series of interconnected membranous sacs and tubules that collectively modifies proteins and synthesizes lipids. However, these two functions are performed in separate areas of the ER: the rough ER and the smooth ER, respectively.
The hollow portion of the ER tubules is called the lumen or cisternal space. The membrane of the ER, which is a phospholipid bilayer embedded with proteins, is continuous with the nuclear envelope.
Rough ER
The rough endoplasmic reticulum (RER) is so named because the ribosomes attached to its cytoplasmic surface give it a studded appearance when viewed through an electron microscope (see figure below).
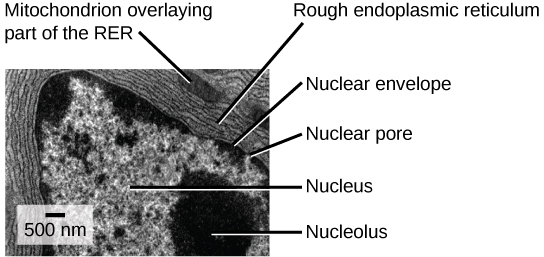
This transmission electron micrograph shows the rough endoplasmic reticulum and other organelles in a pancreatic cell. (credit: modification of work by Louisa Howard)
Ribosomes transfer their newly synthesized proteins into the lumen of the RER where they undergo structural modifications, such as folding or the acquisition of side chains. These modified proteins will be incorporated into cellular membranes—the membrane of the ER or those of other organelles—or secreted from the cell (such as protein hormones, enzymes). The RER also makes phospholipids for cellular membranes.
If the phospholipids or modified proteins are not destined to stay in the RER, they will reach their destinations via transport vesicles that bud from the RER’s membrane.
Since the RER is engaged in modifying proteins (such as enzymes, for example) that will be secreted from the cell, you would be correct in assuming that the RER is abundant in cells that secrete proteins. This is the case with cells of the liver, for example.
Smooth ER
The smooth endoplasmic reticulum (SER) is continuous with the RER but has few or no ribosomes on its cytoplasmic surface. Functions of the SER include synthesis of carbohydrates, lipids, and steroid hormones; detoxification of medications and poisons; and storage of calcium ions.
In muscle cells, a specialized SER called the sarcoplasmic reticulum is responsible for storage of the calcium ions that are needed to trigger the coordinated contractions of the muscle cells.
The Golgi Apparatus
We have already mentioned that vesicles can bud from the ER and transport their contents elsewhere, but where do the vesicles go? Before reaching their final destination, the lipids or proteins within the transport vesicles still need to be sorted, packaged, and tagged so that they wind up in the right place. Sorting, tagging, packaging, and distribution of lipids and proteins takes place in the Golgi apparatus (also called the Golgi body), a series of flattened membranes (see figure below).
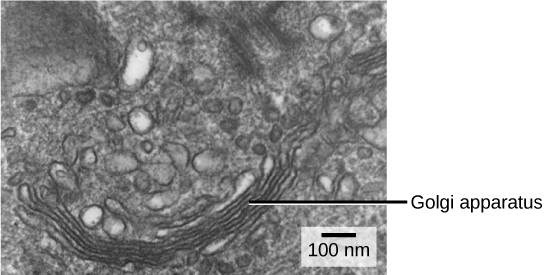
The Golgi apparatus in this white blood cell is visible as a stack of semicircular, flattened rings in the lower portion of the image. Several vesicles can be seen near the Golgi apparatus. (credit: modification of work by Louisa Howard)
The receiving side of the Golgi apparatus is called the cis face. The opposite side is called the trans face. The transport vesicles that formed from the ER travel to the cis face, fuse with it, and empty their contents into the lumen of the Golgi apparatus. As the proteins and lipids travel through the Golgi, they undergo further modifications that allow them to be sorted. The most frequent modification is the addition of short chains of sugar molecules. These newly modified proteins and lipids are then tagged with phosphate groups or other small molecules so that they can be routed to their proper destinations.
Finally, the modified and tagged proteins are packaged into secretory vesicles that bud from the trans face of the Golgi. While some of these vesicles deposit their contents into other parts of the cell where they will be used, other secretory vesicles fuse with the plasma membrane and release their contents outside the cell.
In another example of form following function, cells that engage in a great deal of secretory activity (such as cells of the salivary glands that secrete digestive enzymes or cells of the immune system that secrete antibodies) have an abundance of Golgi.
In plant cells, the Golgi apparatus has the additional role of synthesizing polysaccharides, some of which are incorporated into the cell wall and some of which are used in other parts of the cell.
Summary of Endomembranes
The endomembrane system includes the nuclear envelope, lysosomes, vesicles, the ER, and Golgi apparatus, as well as the plasma membrane. These cellular components work together to modify, package, tag, and transport proteins and lipids that form the membranes.
The RER modifies proteins and synthesizes phospholipids used in cell membranes. The SER synthesizes carbohydrates, lipids, and steroid hormones; engages in the detoxification of medications and poisons; and stores calcium ions. Sorting, tagging, packaging, and distribution of lipids and proteins take place in the Golgi apparatus. Lysosomes are created by the budding of the membranes of the RER and Golgi. Lysosomes digest macromolecules, recycle worn-out organelles, and destroy pathogens.
The Cell Wall
The cell wall of both prokaryotes and eukaryotes should be thought of as a web or mesh rather than a solid wall. It's major function is to provide shape and stability to the cell- preventing over-expansion of the plasma membrane in response to a hypotonic environment and contributing to the overall shape of the organism. Fungal and algal cells also have cell walls. The major organic molecule in the plant cell wall is cellulose (see structure below), a polysaccharide made up of glucose subunits. These chains of glucose molecules are spun together by cellulose synthase complexes at the plasma membrane to generate cellulose microfibrils. The resulting mesh is very porous- allowing even proteins to diffuse within the wall.
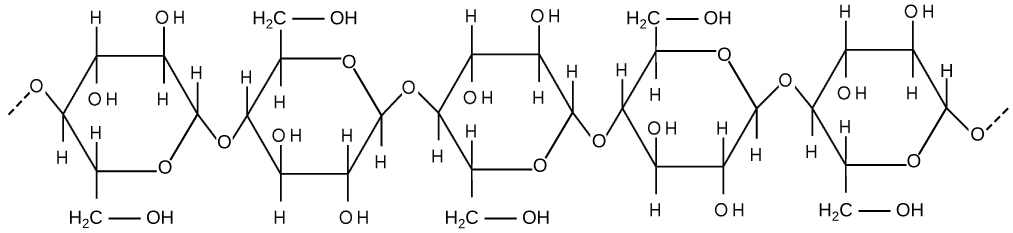
Above: Cellulose is a long chain of β-glucose molecules connected by a 1-4 linkage. The dashed lines at each end of the figure indicate more glucose units.
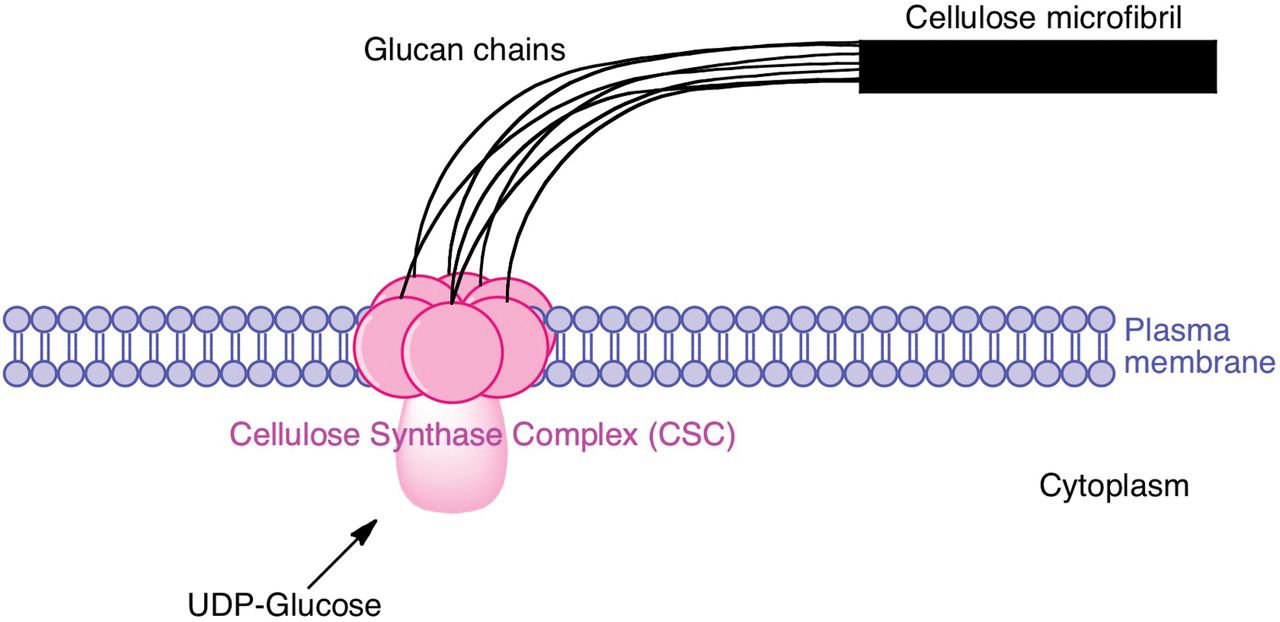
Below: A cellulose synthase complex weaves a cellulose microfibril, which is extruded to the exterior of the cell.
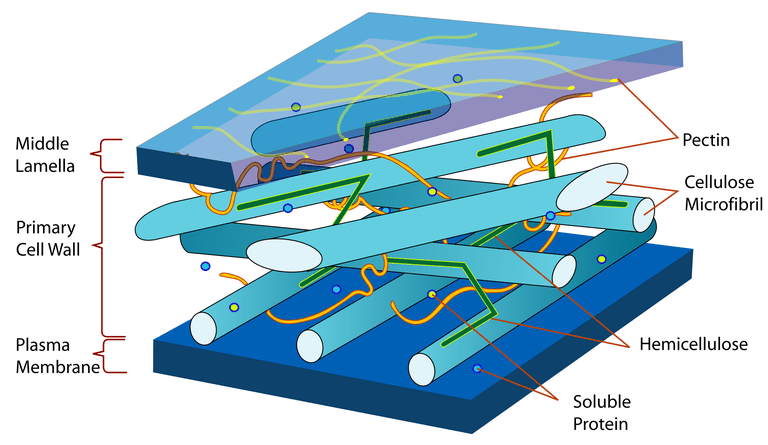
A plant cell wall. Note the relative size of the cellulose microfibrils vs. a typical protein.
Chloroplasts
Chloroplasts are plant cell organelles that carry out photosynthesis. Like the mitochondria, chloroplasts have their own DNA and ribosomes, but chloroplasts have an entirely different function.
Like mitochondria, chloroplasts have outer and inner membranes, but within the space enclosed by a chloroplast’s inner membrane is a set of interconnected and stacked fluid-filled membrane sacs called thylakoids (figure below). Each stack of thylakoids is called a granum (plural = grana). The fluid enclosed by the inner membrane that surrounds the grana is called the stroma.
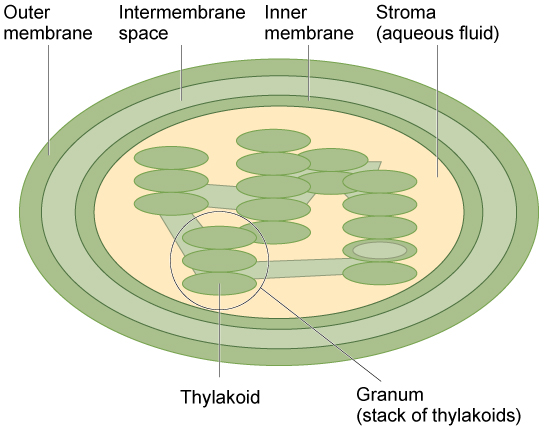
The chloroplast has an outer membrane, an inner membrane, and membrane structures called thylakoids that are stacked into grana. The space inside the thylakoid membranes is called the thylakoid lumen. The light harvesting reactions take place in the thylakoid membranes, and the synthesis of sugar takes place in the fluid inside the inner membrane, which is called the stroma. Chloroplasts also have their own genome, which is contained on a single circular chromosome.
The thylakoid membranes contain a green pigment called chlorophyll, which captures the light energy that drives the reactions of photosynthesis. The light reactions, including electron transport, NADPH production, water splitting, and ATP synthesis all take place on enzymes that are attached to the thylakoid membrane, binding their substrates and releasing their products in the stroma. This provides energy and reducing power for the Calvin cycle, which occurs in the stroma ( as do many other biosynthetic reactions). Like plant cells, photosynthetic protists also have chloroplasts.
The Centrosome
The centrosome is a microtubule-organizing center found near the nuclei of animal cells. It contains a pair of centrioles, two structures that lie perpendicular to each other (see figure below). Each centriole is a cylinder of nine triplets of microtubules.
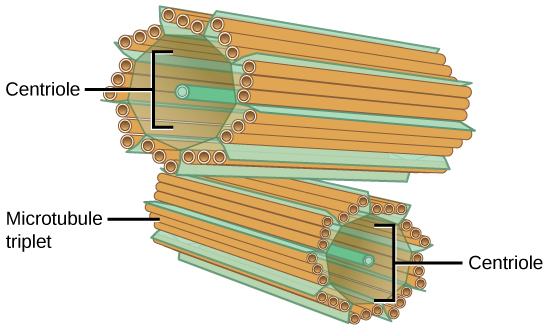
The centrosome consists of two centrioles that lie at right angles to each other. Each centriole is a cylinder made up of nine triplets of microtubules. Nontubulin proteins (indicated by the green lines) hold the microtubule triplets together.
The centrosome (the organelle where all microtubules originate in animals) replicates itself before a cell divides, and the centrioles appear to have some role in pulling the duplicated chromosomes to opposite ends of the dividing cell. However, the exact function of the centrioles in cell division isn’t clear, because cells that have had the centriole removed can still divide, and plant cells, which lack centrioles, are capable of cell division.
Lysosomes
In addition to their role as the digestive component and organelle-recycling facility of animal cells, lysosomes are considered to be parts of the endomembrane system. Lysosomes also use their hydrolytic enzymes to destroy pathogens (disease-causing organisms) that might enter the cell. A good example of this occurs in a group of white blood cells called macrophages, which are part of your body’s immune system. In a process known as phagocytosis or endocytosis, a section of the plasma membrane of the macrophage invaginates (folds in) and engulfs a pathogen. The invaginated section, with the pathogen inside, then pinches itself off from the plasma membrane and becomes a vesicle. The vesicle fuses with a lysosome. The lysosome’s hydrolytic enzymes then destroy the pathogen (figure below).
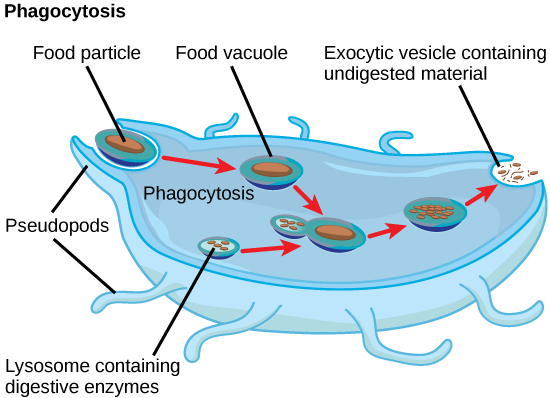
A macrophage has engulfed (phagocytized) a potentially pathogenic bacterium and then fuses with a lysosomes within the cell to destroy the pathogen. Other organelles are present in the cell but for simplicity are not shown.
The Cytoskeleton
Section Summary
The cytoskeleton is a network of different protein fibers that function to: maintain or change the shape of the cell, secure some organelles in specific positions, enable movement of cytoplasm and vesicles within the cell, and enable the cell to move in response to stimuli. There are three types of fibers within the cytoskeleton: microfilaments, intermediate filaments, and microtubules. Some of the cytoskeletal fibers work in conjunction with molecular motors which move along the fibers within the cell to carry out a diverse set of functions. There are two main classes of molecular motors; the actin- associated myosins, and the microtubule-associated dyneins and kinesins.
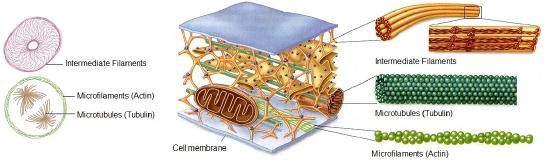
Microfilaments thicken the cortex around the inner edge of a cell; like rubber bands, they resist tension. They are also involves in changing cell shape- n cell division, and in phagocytosis or cell crawling. Microtubules are found in the interior of the cell where they maintain cell shape by resisting compressive forces. They're also involved in the trafficking of vesicles and chromosomes. Intermediate filaments represent a variety of proteins, and not involved in movement and trafficking, are found throughout the cell and help hold organelles in place. The can also act to hold adjacent cells together.
Design Challenge
Eukaryotic cells contain membrane-bound organelles that effectively separate materials, processes, and reactions from one another and from the cytoplasm. This in itself poses a problem for eukaryotes.
How can the cell purposely move and control the location of materials between these organelles? More specifically, how can a eukaryotic cell transport compounds from their place of origin (for example, the cytoplasm) to where they are needed (perhaps the nucleus, the mitochondria or the cell surface)?
Possible discussion
Propose some reasons why cells - particularly large cells and/or cells with organelles - cannot rely on simple diffusion to move metabolites, building blocks, proteins, etc. to the locations in the cell where they are needed.
One possible solution is for the cell to create a network that can connect all the different parts of the cell together. This network could be used not only as a scaffold to hold components in place but also as a reference for direction. For example, we can use a map to determine the direction we need to travel and roads to connect and travel from home to campus. Likewise, an interconnecting network inside the cell can be used to direct and move compounds from one location to a final destination. Some of the required characteristics of this network are listed below. Can you add to this list?
Intracellular Network
- The network would need to be extensive, and connect every area of the cell.
- The network would need to be flexible, able to change and adapt as the cell grows larger, divides into two cells, or physically moves from one environment to another.
- The network needs to be strong, able to hold up to mechanical pressure from inside the cell or from outside of the cell.
- The network needs to be composed of different fibers and each of these fibers needs to be for a specific connection in the cell. For example, certain fibers might be involved in holding organelles in place, and other fibers would be involved in connecting two different organelles.
- The fibers need to have directionality (or polarity), meaning, they need to have a defined starting point and a defined end to help direct movement from one location to another.
- The fibers need to work with proteins that can convert chemical energy into kinetic energy, to actively transport compounds along the fibers.
Microfilaments
Actin
Microfilaments are cytoskeleton fibers composed of actin subunits. Actin is one of the most abundant proteins in eukaryotic cells and comprises 20% of total cellular protein by weight in muscle cells. The actin amino acid sequence is highly conserved in eukaryotic cells, meaning that the protein amino acid sequence, and therefore its final 3D shape, has changed little over the course of evolution, maintaining more than 80% similarity between algae and humans.
Actin can be present as either a free monomer called G-actin (globular) or as part of a polymer microfilament called F-actin ('F' for filamentous). Actin must be bound to ATP in order to assemble into its filamentous form and maintain the structural integrity of the filament. The actin filament itself has structural polarity. This term "polarity" in reference to a cytoskeleton filament, does not mean what it did when we discussed polar functional groups earlier in this course. Polarity here refers to the fact that there are two distinct ends to the filament. These ends are called the "(-) end" and the "(+) end". At the "(+) end", actin subunits are added onto the elongating filament and at the "(-) end", actin subunits are disassembling, or falling off of the filament. This process of assembly and disassembly is controlled by the ATP:ADP ratio in the cytoplasm.
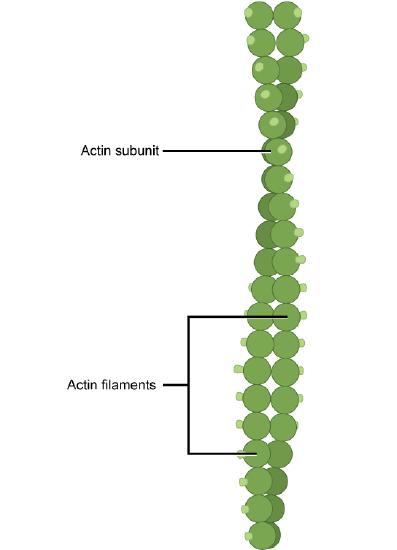
Microfilaments are the narrowest of the three cytoskeleton fibers, with a diameter of about 7 nm. Microfilaments are composed of actin subunits which form into two intertwined strands.
Actin participates in many cellular processes, including muscle contraction, cell motility, cytokinesis during cell division, vesicle and organelle movement, and the maintenance of cell shape. Actin filaments serve as a track for the movement of a family of motor proteins called myosins discussed in more detail in a section below.
Link to Learning:
To see an example of a white blood cell in action, click here and watch a short time-lapse video of the cell capturing two bacteria. It engulfs one and then moves on to the other.
Animations on actin filaments and how they work
- Actin filament assembly
- Sliding movement of actin filaments
Intermediate Filaments
Intermediate filaments are associated with animal cell biology. They are made of several strands of fibrous proteins that are wound together. These elements of the cytoskeleton get their name from the fact that their diameter, 8 to 10 nm, is between those of the smaller microfilaments and the larger microtubules. The intermediate filaments are the most diverse group of cytoskeletal elements. Several types of fibrous proteins are found in the intermediate filaments. You are probably most familiar with keratin, the fibrous protein that strengthens your hair, nails, and the epidermis of the skin.
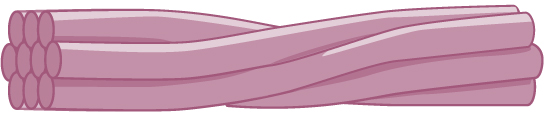
Intermediate filaments consist of several intertwined strands of fibrous proteins.
Intermediate filaments have no role in cell movement. Their function is purely structural. They bear tension, thus maintaining the shape of the cell, and anchor the nucleus and other organelles in place. The figure above shows how intermediate filaments create a cable-like supportive scaffolding inside the cell.
Microtubules
Microtubules are the largest component of the cytoskeleton and are found throughout the cytoplasm. These polymers are made up of globular protein subunits called α-tubulin and β-tubulin. Microtubules are found not only in eukaryotic cells but in some bacteria as well.
Both the α-tubulin and β-tubulin subunits bind to GTP. When bound to GTP, the formation of the microtubule can begin, this is called the nucleation event. As more GTP tubulin dimers assemble onto the filament, GTP is slowly hydrolyzed by β-tubulin to form GDP. Tubulin bound to GDP is less structurally robust and can lead to disassembly of the microtubule.
Much like the actin filaments discussed above, microtubules also have a distinct polarity that is critical for their biological function. Tubulin polymerizes end to end, with the β-subunits of one tubulin dimer contacting the α-subunits of the next dimer. These differences lead to different subunits being exposed on the two ends of the filament. The ends are designated the "(−)" and "(+)" ends. Unlike actin filaments, microtubules can elongate at both the (+) and (-) ends, but elongation is significantly more rapid at the (+) end.
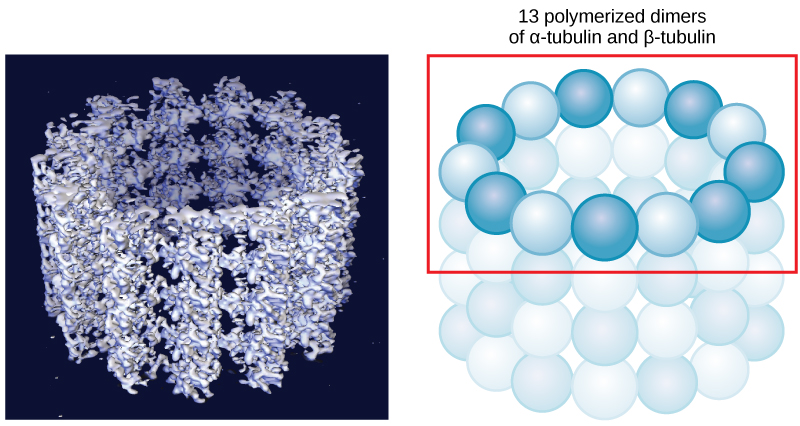
Microtubules are hollow. Their walls consist of 13 polymerized dimers of α-tubulin and β-tubulin (right image). The left image shows the molecular structure of the tube.
Microtubules help the cell resist compression, provide a track along which vesicles move through the cell, pull replicated chromosomes to opposite ends of a dividing cell, and are the structural elements of flagella, cilia, and centrioles.
HIGHLY RECOMMENDED:
Animations of the cytoskeleton
Where did these fibers come from?
The cytoskeleton probably has its origins in bacterial and/or archaeal ancestry. There are ancient relatives to both Actin and Tubulin in bacterial systems. In bacteria, the MreB protein and the ParM protein are believed to be early ancestors to Actin. MreB functions in maintaining cell shape and ParM functions in plasmid (DNA) partitioning. The FtsZ protein in bacteria functions in cytokinesis, it is a GTPase, spontaneously forms filaments and is hypothesized to be an ancient form of Tubulin. These findings support the hypothesis that the eukaryotic cytoskeleton has its origins in the bacterial world.
Flagella and Cilia
Flagella (singular = flagellum) are long, hair-like structures that extend from the plasma membrane and are used to move an entire cell (for example, sperm, Euglena). When present, the cell has just one flagellum or a few flagella. Cilia are short, hair-like structures that are used to move entire cells (such as paramecia) or substances along the outer surface of the cell (for example, the cilia of cells lining the Fallopian tubes that move the ovum toward the uterus, or cilia lining the cells of the respiratory tract that trap particulate matter and move it toward your nostrils.) When cilia are present, there can be many of them, extending along the entire surface of the plasma membrane.
Despite their differences in length and number, flagella and cilia share a common structural arrangement of microtubules called a “9 + 2 array.” This is an appropriate name because a single flagellum or cilium is made of a ring of nine microtubule doublets, surrounding a single microtubule doublet in the center Figure below).
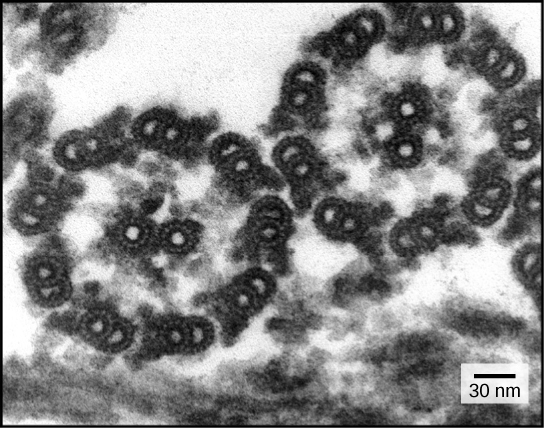
This transmission electron micrograph of two flagella shows the 9 + 2 array of microtubules: nine microtubule doublets surround a single microtubule doublet. (credit: modification of work by Dartmouth Electron Microscope Facility, Dartmouth College; scale-bar data from Matt Russell)
For a video on flagellar and ciliar movement in eukaryotes see the YouTube video: click here (you can skip the commercial).
Motor Proteins
One function of the cytoskeleton is to move cellular components from one part of the cell to another. These cellular components are called 'cargo' and are often stored within a vesicle for transport. You can think of the cytoskeleton as "railroad tracks" providing support and directionality inside of the cell.
Of course, if there are "railroad tracks" there needs to be an engine that can both move on the tracks and pull or push cargo along. In this case the engines are molecular motors that can move along the tracks in a specific direction. There are two families of molecular motors associated with the cytoskeleton; dyneins and kinesins. These motor proteins (train engines) and the cytoskeleton create a comprehensive network within the cell for moving vesicles (box cars) from one organelle to another or from one organelle to the cell surface.
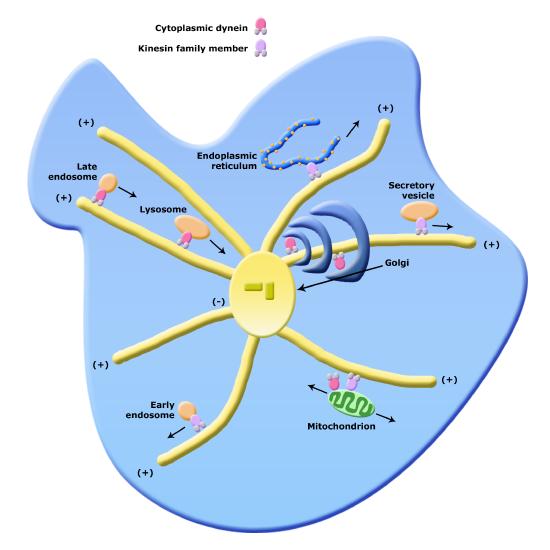
Organelle transport via microtubules and kinesins and dynes. Note that the figure is conceptual and only intended to show directionality of movement of various organelles, not necessarily represent all of their forms faithfully.
Cytoplasmic Dyneins
Dynein is a protein complex that functions as a molecular motor. In cells, it converts the chemical energy from ATP hydrolysis into the mechanical energy of movement to 'walk' along the microtubule while carrying a vesicle. Dyneins bind to microtubules and move or "walk" from the plus (+) end of the cytoskeletal microtubule filament to the minus (-) end of the filament, which is usually oriented towards the cell center. Thus, they are often referred to as "minus-end directed motors" and this vesicular transport is refereed to as retrograde transport. Cytoplasmic dynein moves processively along the microtubule, hydrolyzing ATP with each 'step' it takes along the microtubule. During this process one or the other of its "stalks" is always attached to the microtubule, this allows for the dynein motor (and its cargo) to "walk" a considerable distance along a microtubule without detaching.
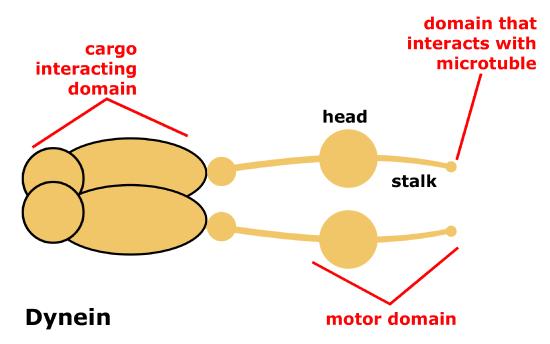
Schematic of cytoplasmic dynein motor protein. Dyneins are protein complexes composed of many smaller polypeptide subunits. The overall structure of the Dynein motors are relatively simple, consisting of 2 identical complexes each having a motor domain that interacts with the microtubule, a stalk or stem region region that connects the motor head to the cargo interacting domain.
Cytoplamic dyneins are involved in organelle movement such as the positioning of the Golgi complex and other organelles in the cell, are used in the transport of cargo such as the movement of vesicles made by the endoplasmic reticulum, endosomes, and lysosomes, and are responsible for the movement of chromosomes during cell division. Axonemal dyneins are motor proteins used in the sliding of microtubules in the axonemes of cilia and flagella in eukaryotic cells.
Kinesins
Kinesins, like cytoplasmic dyneins are motor protein-complexes that "walk" along the microtubules and are involved in vesicle transport. Unlike cytoplasmic dyneins, the polarity of kinesin movement is from the (-) end of the microtubule to the (+) end with the hydrolysis of ATP. In most cells, this entails transporting cargo from the center of the cell towards the periphery (the opposite direction to dyneins). Like cytoplasmic dyneins, kinesins are involved in a variety of cellular processes including vesicle movement and chromosome movement during cell division.
The structure of kinesins are similar to cytoplasmic dyneins. Members of the kinesin superfamily vary in shape but the overall structure is that of a heterotetramer whose motor subunits (heavy chains) form a protein dimer (molecule pair) that binds two light chains.
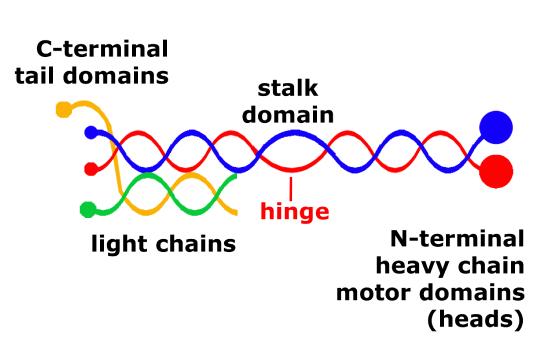
Schematic of kinesin motor proteins. The heavy chains comprise a globular head (the motor domain) at the amino terminal end connected via a short, flexible neck linker to the stalk – a long, central alpha-helical coiled-coil domain – that ends in a carboxy terminal tail domain which associates with the light-chains. The stalks of two light chains intertwine to form a coiled-coil that directs dimerization of the two heavy chains. In most cases transported cargo binds to the kinesin light chains, but in some cases cargo binds to the C-terminal domains of the heavy chains.
How do vesicles "know" they have reached their destination?
Microtubule based trafficking gives vesicles repeated pushes or pulls in a general direction (i.e., away from the nucleus). In cells that are much longer than they are wide (neurons, root hairs) vesicular trafficking can be quite dramatic. However, kinesins do not "know" that they are headed for, for example, the plasma membrane vs. the lysosomal membrane. They're just helping their cargo diffuse. In fact, it is the vesicle itself that recognizes its destination (though without the motor protein it would be unable to move). SNARE proteins embedded in the vesicle and its target recognize one another and facilitate the fusion of the vesicle with the membrane of its target compartment. For example, these proteins facilitate the release of neurotransmitters, allowing neurons to transmit intercellular signals. These same SNAREs are the target of many neurotoxins, include botulinum toxin (Botox). The inhibition of neurotransmitter release by Botox induces flaccid paralysis of muscles.
Energy story: Write an energy story for vesicular transport with in the cell. Include a description of the overall process and the initial and final states of the energy being used in the process. Overall, what work is being done?
HIGHLY RECOMMENDED:
A day in the life of a motor protein
Animations of Kinesin and Dynein at work
- Animation of a cyplasmic dynein motor on a microtubule
- How dynein moves along a microtubule
- mechanism of Kinesin moving on a microtubule
- Kinesin and dynein motors
How do the motors interact with cargo and the microtubules
Cytoplasmic dyneins and kinesins interact with both cargo and microtubules in similar fashion. The light chains interact with receptors on the various cargo vesicles and the globular motor domains, specifically interact with the microtubules.
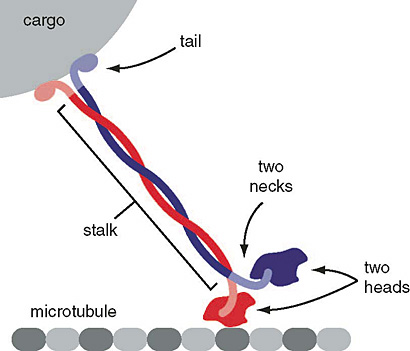
Schematic of kinesin motor protein carrying a cargo vesicle along a microtubule filament.
Possible discussion
What are the benefits for having multiple types of motor proteins? Multiple types of filaments? Filaments with polarity?

