4.8: Genetic screens - Inherited Phenotypes Reveal Functions of Genes
- Page ID
- 40141
Learning Objectives
- Compare forward and reverse genetic approaches.
- Identify examples and explain the value of using model organisms to understand the functions of genes.
- Recall that in genetic screens mutations occur random but that specific phenotypes can be identified by screening large numbers of organisms.
The principle of genetic screens is simple (even though they can be complicated and labor-intensive to perform).
Genetic screens rely on essentially two things:
- That the researcher can induce mutations
- Making mutations at random to find unusual phenotypes without knowing which gene is affected = forward genetics
- Making a mutation in (or inhibiting the function of) a specific gene to observe changes in phenotype = reverse genetics
- That a mutation in a gene could result in an observable change in phenotype
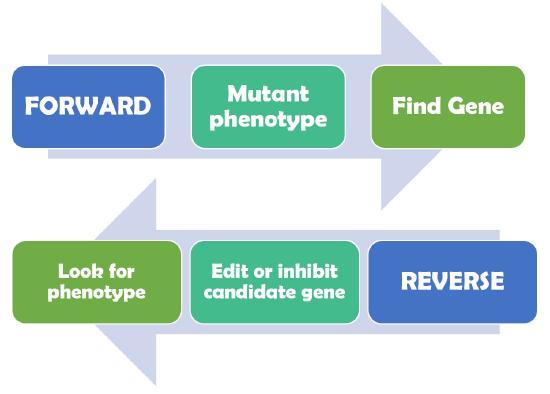
Figure \(\PageIndex{1}\): Forward genetics begins by identifying a mutant phenotype following by identifying the gene that was mutated. Reverse genetics approaches became possible by advances in molecular biology that allowed scientists to specifically manipulate individual genes of interest and then determine if the genetic manipulation resulted in a change in phenotype. (Copyright Leacock own work)
Random mutations can be used as a screening tool
Identifying genes involved in development
Forward genetic screens for mutations that altered embryonic development became foundational to the fields of genetics and developmental biology. The basic principle is to expose males to a mutagen and then screen their offspring (often in the second generation after exposure) for phenotypes of interest. The vast majority of mutations will not result in a phenotypic change, so the most of the animals will appear wild-type. However, the rare mutants show that a gene important for that phenotype has been disrupted in some way.
Drosophila patterning and segmentation
Eric Wieschaus, Christiane Nüsslein-Volhard, and Edward B. Lewis shared the 1995 Nobel Prize in Physiology or Medicine for discoveries made using forward genetic screens in the fruit fly Drosophila melanogaster. This groundbreaking work revealed sets of genes that set up the body patterning and segmentation of the fruit fly embryo and adult structures. Edward Lewis examined mutants in which the fates of segments were transformed into the fates of other segments, leading to the identification of a conserved set of genes involved in segment identity. Notably these genes are transcription factors, which supports the important role that regulation of gene transcription has on cell fate.
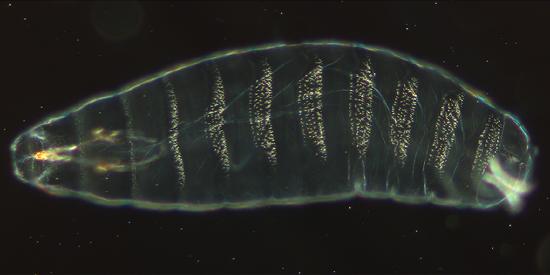
Figure \(\PageIndex{4}\): Cuticle preparation of a Drosophila melanogaster embryo, about 22 hours old. Ventral view with denticles. Anterior (the head) is on the left side. 200x magnification. (CC BY-SA Nina via https://commons.wikimedia.org/wiki/F...laKutikula.jpg)
Nüsslein-Volhard and Wieschaus Forward Genetic Screen Overview
- Exposed male fruit flies to EMS (an alkylating agent)
- Established 5,800 mutant lines (using genetic markers on the second and third chromosome)
- Identified dead embryos in 4,500 mutant lines (an embryonic lethal phenotype)
- Examined the cuticle pattern in 2,600 embryonic lethal lines
- Grouped 15 abnormal cuticle patterns into three classes and mapped to chromosomal locations
- Segment Polarity (6 genes identified)
- Pair-rule (6 genes identified)
- Gap (3 genes identified)
Summarized from Nüsslein-Volhard and Wieschaus, 1980
In the video below, Drs. Wieschaus and Nüsslein-Volhard discuss how they worked together to pay attention to small details that made these discoveries possible in the interview below. Although the genes that were revealed in this screen turned out to be extremely important genes throughout the animal kingdom, remember that the mutations occurred at random in response to a mutagen. In this way, forward genetics screens rely on the organism to inform us which genes are important for a process.
Video \(\PageIndex{1}\): Interview with Eric Wieschaus and Christiane Nüsslein-Volhard: Collaborating to Find Developmental Genes. (Copyright iBiology via https://www.youtube.com/watch?v=GmQ9eI1vdGM)
Cell Death Genes in C. elegans
Caenorhabditis elegans, a microscopic roundworm, serves as a model organism for numerous aspects of cell and molecular biology. These worms are primarily self-fertilizing hermaphrodites, although males also exist.
Hermaphrodites and sexual reproduction
Remember that hermaphrodite means having both male and female organs for sexual reproduction. In C. elegans, hermaphrodites produce sperm via meiosis during the last larval stage and store the sperm in a specialized organ. Then in the adult stage, the germ cells undergoing meiosis become oocytes. The oocytes are then fertilized by the stored sperm.
Meiosis, including the rules of chromosome segregation and inheritance, still occurs and the worms reproduce using sexual reproduction (not asexual reproduction).
C. elegans males occur when only one X chromosome is present. These males have different structural features and produce only sperm, but can fertilize hermaphrodites, still using gametes and sexual reproduction.
C. elegans are transparent, which allowed scientists first studying this organism to follow the pattern of cell division, differentiation, or programmed cell death of every cell under the microscope. This foundational work revealed that these worms develop in an invariant pattern, in which every wild-type animal undergoes the same number and pattern of cell divisions. This knowledge allowed for forward genetic screens in which this pattern was abnormal.
In one such screen, male worms were exposed to ethyl methanesulfonate (EMS), an alkylating agent, to induce mutations. These males were mated with hermaphrodites and researchers examined the second generation (F2) offspring for abnormal persistence of cells that normally undergo programmed cell death in wild-type animals (Hedgecock et al. 1983). The first two mutants were named ced-1 and ced-2 (for programmed cell death) and the genes that were eventually identified as mutated were named ced-1 and ced-2. Additional ced mutants were identified in subsequent screens, ultimately identifying a pathway of proteins that act to trigger cell death, kill the cell, and execute the engulfment of the dead cell.
In 2002, Sydney Brenner, John E. Sulston, and H. Robert Horvitz shared the Nobel Prize "for their discoveries concerning genetic regulation of organ development and programmed cell death" (https://www.nobelprize.org/prizes/medicine/2002/press-release/). Cloning of cell death genes in the nematode had revealed a conserved set of proteins involved in programmed cell death in eukaryotes, giving insight into normal functions of programmed cell death as well as abnormal avoidance of cell death in cancer.
References
Conradt B and Xue D. (2005) Programmed cell death. Wormbook.org (http://www.wormbook.org/chapters/www_programcelldeath/programcelldeath.html#bib20)
Ellis HM, Horvitz HR. Genetic control of programmed cell death in the nematode C. elegans. Cell. 1986 Mar 28;44(6):817-29. doi: 10.1016/0092-8674(86)90004-8. PMID: 3955651.
Hedgecock EM, Sulston JE, Thomson JE (1983) Mutations affecting programmed cell deaths in the nematode Caenorhabditis elegans. Science. Vol. 220, Issue 4603, pp. 1277-1279 DOI: 10.1126/science.6857247
Lewis EB (1978). A gene complex controlling segmentation in Drosophila. Nature 276, 565-570.
Nüsslein-Volhard, C., Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801 (1980). https://doi.org/10.1038/287795a0


