15.1.3.1: Escherichia coli
- Page ID
- 42663
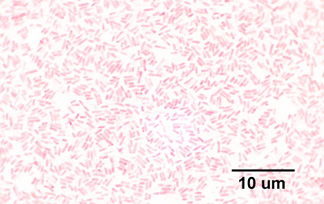
Gram Stain of Escherichia coli. Note Gram-negative (pink) bacilli.
Organism
- E. coli is a member of the Enterobacteriaceae family which are all Gram-negative rods
- Catalase positive
- Oxidase negative
- Facultative anaerobe
- E. coli characteristically ferments lactose
Habitat
- Normal inhabitant of the intestines of humans and other mammals
Source
- Fecal contamination of food or water
- Inadequate hygiene
- Can be endogenous (from own flora) or exogenous (from outside the body)
Epidemiology
- Certain strains can cause urinary tract infection (UTI) or gastrointestinal infection in people without specific risk factors
- Extremely common cause of nosocomial infections
- E. coli and other Enterobacteriaceae cause about 25-30% of the approximately 1.7 million nosocomial infections in the US each year
- Outbreaks of enterohemorrhagic E. coli (EHEC) infection are associated with contaminated food
Clinical Disease and Virulence Factors
- Urinary tract infection
- E. coli is the most common cause of UTI
- Much more common in women (about 14 times)
- E. coli can be introduced into the urinary tract from the periurethral region during intercourse
- Symptoms include:
- A strong, persistent urge to urinate
- A burning sensation when urinating
- Passing frequent, small amounts of urine
- Cloudy urine
- Blood in the urine
- Strong-smelling urine
- Pelvic pain
- Attach to the urinary tract with fimbriae and induce inflammation
- Food poisoning (enterotoxigenic E. coli (ETEC)) (a.k.a. traveler's diarrhea)
- due to consumption of food contaminated with enterotoxin-producing strains of E. coli
- more common in areas with less developed sanitation
- caused enterotoxins produced by the infecting bacteria
- causes diarrhea without fever
- Bismuth subsalicylate (Pepto-Bismol) binds to the enterotoxin and can be useful for treatment of symptoms or prophylaxis
- Antibiotics may be warranted in more severe cases
- Enterohemorrhagic E. coli (EHEC) (such as strain O157:H7)
- Primarily associated with contaminated food. As an inhabitant of the GI tract of cows, it is commonly associated with beef. Ground beef is of particular concern because it is a mixture of various cuts, and if there is contamination it is distributed throughout the product rather than just on the surface as it would be in a steak or roast. Other foods have also been associated with outbreaks, such as leafy greens, sprouts, nuts, and nut butters.
- Causes hemorrhagic (bloody) diarrhea, sometimes with cramping or vomiting
- Can lead to potentially fatal hemolytic uremic syndrome (HUS)
- blood vessels in kidneys are damaged
- can cause kidney failure
- The transfer of several proteins to the host cell through a pilus induces intestinal epithelial cells to produce pedestals to which the E. coli can tightly bind (Figure \(\PageIndex{1}\))
- Produces shiga toxin, which damages intestinal epithelium and causes hemorrhagic diarrhea. It may also play a significant role in the development of HUS.
- Gene for the shiga toxin is phage-encoded
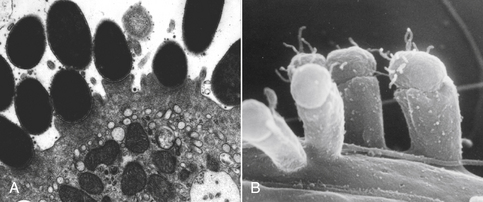
Figure \(\PageIndex{1}\): Scanning and transmission electron micrographs showing actin pedestals induced by enteropathogenic and enterohemorrhagic Escherichia coli (EPEC and EHEC). A, EPEC generates attaching and effacing (AE) lesions on the intestinal epithelium after infection of gnotobiotic piglets. Note the pedestal-like structures on host cells beneath attached bacteria. (From Campellone KG, Leong JM. Tails of two Tirs: actin pedestal formation by enteropathogenic E. coli and enterohemorrhagic E. coli. Curr Opin Microbiol 2003;6(1):82-90.) B, Actin pedestals that resemble AE lesions formed in vivo are also generated on cultured epithelial (HeLa) cells. (Courtesy of Knutton S. In: Knutton S, Rosenshine L, Pallen, MJ, et al. A novel EspA-associated surface organelle of enteropathogenic Escherichia coli involved in protein translocation into epithelial cells. EMBO J. 1998;17:2166-2176.)
- Common nosocomial infections
- (although these are listed for E. coli, many other members of the Enterobacteriaceae such as Klebsiella and Citrobacter can cause similar infections)
- neonatal meningitis
- UTI
- wound infections/abcesses
- pneumonia
- bacteremia/septicemia
- Carbapenem-resistant Enterobacteriaceae (CRE)
- CRE are resistant to nearly all antibiotics and are classified as an "urgent threat" by the CDC (fact sheet)
- Produce beta-lactamases that can break down carbapenems and most other beta-lactam antibiotics
- Genes for these beta-lactamases (such as NDM-1) are primarily found on mobile genetic elements (plasmids or phages) and can spread rapidly among the similarly pathogenic species of Enterobacteriaceae, especially since they are close together in the intestines and common in wastewater.
- Prudent use of antibiotics, proper aseptic technique and hygiene, and monitoring for CRE infections are all critical to controlling the spread of CRE
Additional Information:

