17.4B: Pharmacogenomics, Toxicogenomics, and Metagenomics
- Page ID
- 13385
- Explain how microbial metagenomics can give researchers a more complete picture of a particular environment
Pharmacogenomics, also called toxicogenomics, involves evaluating the effectiveness and safety of drugs on the basis of information from an individual’s genomic sequence. Genomic responses to drugs can be studied using experimental animals (such as laboratory rats or mice) or live cells in the laboratory before embarking on studies with humans. Studying changes in gene expression could provide information about the transcription profile in the presence of the drug, which can be used as an early indicator of the potential for toxic effects. For example, genes involved in cellular growth and controlled cell death, when disturbed, could lead to the growth of cancerous cells. Genome-wide studies can also help to find new genes involved in drug toxicity. Personal genome sequence information can be used to prescribe medications that will be most effective and least toxic on the basis of the individual patient’s genotype. The gene signatures may not be completely accurate, but can be tested further before pathologic symptoms arise.
Microbial Genomics: Metagenomics
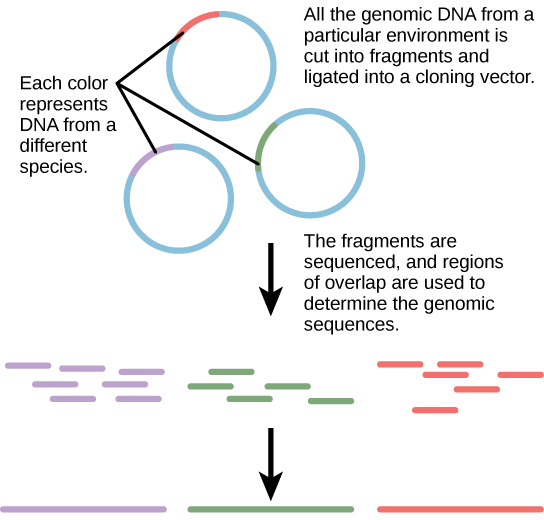
Traditionally, microbiology has been taught with the view that microorganisms are best studied under pure culture conditions, which involves isolating a single type of cell and culturing it in the laboratory. Because microorganisms can go through several generations in a matter of hours, their gene expression profiles adapt to the new laboratory environment very quickly. In addition, the vast majority of bacterial species resist being cultured in isolation. Most microorganisms do not live as isolated entities, but in microbial communities known as biofilms. For all of these reasons, pure culture is not always the best way to study microorganisms. Metagenomics is the study of the collective genomes of multiple species that grow and interact in an environmental niche. Metagenomics can be used to identify new species more rapidly and to analyze the effect of pollutants on the environment.
Key Points
- Pharmacogenomics involves evaluating the effectiveness and safety of drugs on the basis of information from an individual’s genomic sequence.
- Genomic responses to drugs can be studied using experimental animals or live cells in the laboratory, which help indicate the potentially toxic effects of a drug.
- Personal genome sequence information can be used to prescribe medications that will be most effective and least toxic on an individual level.
- Metagenomics, the study of the collective genomes of multiple species that grow and interact in an environmental niche, is often a better way to study microrganisms rather than pure culture.
Key Terms
- pharmacogenomics: the study of genes that code for enzymes that metabolize drugs, and the design of tailor-made drugs adapted to an individual’s genetic make-up
- metagenomics: the study of the collective genomes of multiple species that grow and interact in an environmental niche


