15.4: Prokaryotic Transcription - Initiation of Transcription in Prokaryotes
- Page ID
- 13302
RNA polymerase initiates transcription at specific DNA sequences called promoters.
- Summarize the initial steps of transcription in prokaryotes
Key Points
- Transcription of mRNA begins at the initiation site.
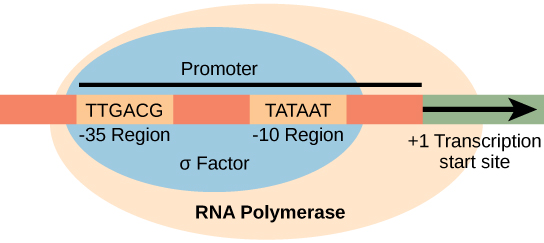
- Two promoter consensus sequences are at the -10 and -35 regions upstream of the initiation site.
- The σ subunit of RNA polymerase recognizes and binds the -35 region.
- Five subunits (α, α, β, β’, and σ) make up the complete RNA polymerase holoenzyme.
Key Terms
- holoenzyme: a fully functioning enzyme, composed of all its subunits
- promoter: the section of DNA that controls the initiation of RNA transcription
Prokaryotic RNA Polymerase
Prokaryotes use the same RNA polymerase to transcribe all of their genes. In E. coli, the polymerase is composed of five polypeptide subunits, two of which are identical. Four of these subunits, denoted α, α, β, and β’, comprise the polymerase core enzyme. These subunits assemble each time a gene is transcribed; they disassemble once transcription is complete. Each subunit has a unique role: the two α-subunits are necessary to assemble the polymerase on the DNA; the β-subunit binds to the ribonucleoside triphosphate that will become part of the nascent “recently-born” mRNA molecule; and the β’ binds the DNA template strand. The fifth subunit, σ, is involved only in transcription initiation. It confers transcriptional specificity such that the polymerase begins to synthesize mRNA from an appropriate initiation site. Without σ, the core enzyme would transcribe from random sites and would produce mRNA molecules that specified protein gibberish. The polymerase comprised of all five subunits is called the holoenzyme.
Prokaryotic Promoters and Initiation of Transcription
The nucleotide pair in the DNA double helix that corresponds to the site from which the first 5′ mRNA nucleotide is transcribed is called the +1 site, or the initiation site. Nucleotides preceding the initiation site are given negative numbers and are designated upstream. Conversely, nucleotides following the initiation site are denoted with “+” numbering and are called downstream nucleotides.
A promoter is a DNA sequence onto which the transcription machinery binds and initiates transcription. In most cases, promoters exist upstream of the genes they regulate. The specific sequence of a promoter is very important because it determines whether the corresponding gene is transcribed all the time, some of the time, or infrequently. Although promoters vary among prokaryotic genomes, a few elements are conserved. At the -10 and -35 regions upstream of the initiation site, there are two promoter consensus sequences, or regions that are similar across all promoters and across various bacterial species. The -10 consensus sequence, called the -10 region, is TATAAT. The -35 sequence, TTGACA, is recognized and bound by σ. Once this interaction is made, the subunits of the core enzyme bind to the site. The A–T-rich -10 region facilitates unwinding of the DNA template; several phosphodiester bonds are made. The transcription initiation phase ends with the production of abortive transcripts, which are polymers of approximately 10 nucleotides that are made and released.