14: Translation (Protein synthesis)
- Page ID
- 384
A reminder: mRNA encodes the polypeptide with each amino acid designated by a string of three nucleotides. tRNAs serve as the adaptors to translate from the language of nucleic acids to that of proteins. Ribosomes are the factories for protein synthesis.
A. tRNAs
1. The transfer RNAs, or tRNAs serve as adaptors to align the appropriate amino acids on the mRNA templates.
Figure 3.5.1.

2. Primary structure of tRNAs
a. tRNAs are short, being only 73 to 93 nts long.
b. All tRNAs have the trinucleotide CCA at the 3' end.
- The amino acid is attached to the terminal A of the CCA.
- In most prokaryotic tRNA genes, the CCA is encoded at the 3' end of the gene. No known eukaryotic tRNA gene encodes the CCA, but rather it is added posttranscriptionally by the enzyme tRNA nucleotidyl transferase.
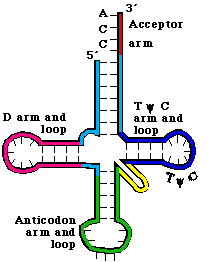
a. tRNAs have 4 arms with 3 loops (see Figure 3. 5.2. for yeast phenylalanine tRNA)
b. Theamino acid acceptor arm is formed by complementary base‑pairing between the intial 7 nts of tRNA and a short segment near the 3' end. Again, the amino acid will be added to the terminal A.
c. The D arm ends in the D loop. It contains several dihydrouridines, which are abbreviated "D".
d. The anticodon arm ends in anticodon loop. The anticodon is located in the center of the loop. It will align 3' to 5' with the mRNA (reading 5' to 3').
e. The variable loop varies in size in different tRNAs. The difference in size between the 73 nt versus 93 nt tRNAs is found in the variable loop.
f. The TyC arm is named for this highly conserved motif found in the loop.
4. The tertiary structure of tRNA is a "fat L". (See Fig 3.5.3.)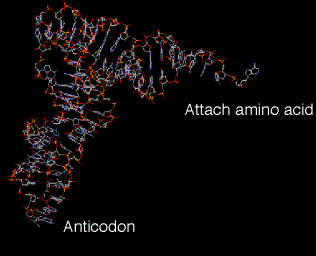
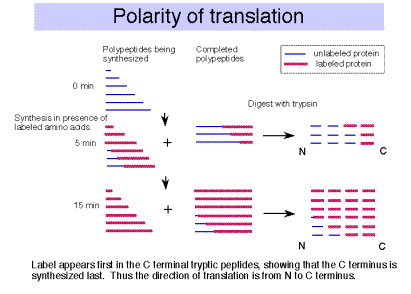
F. The polarity of translation is from the amino (N) terminus to the caboxy (C) terminus.
This was demonstrated in a classic experiment by Dintzis.
- Actively translating proteins were labeled with radioactive amino acids for a brief time (short relative to the time required to complete synthesis).
- Completed polypeptides were collected, digested with trypsin, and the amount of radioactivity in tryptic fragments was determined.
- Tryptic fragments from the C‑terminal end of the polypeptide had radioactivity at the earliest times of labeling.
- As the period of labeling was increased (longer pulse), tryptic fragments closer to the N terminus were labeled.
- This shows that the direction of polypeptide growth is from the N teminus to the C terminus, i.e. translation begins at the N terminal amino acid. This corresponds to mRNA chain growth in a 5' to 3' direction.
- Note that this experimental protocol is also used to map origins of replication, as we covered in Part Two of the course.
b. IF3= Initiation Factor 3
- An antiassociation factor; prevents association between the large and small ribosomal subunits.
- It also must be associated with the small subunit for it to form an initiation complex, i.e. for the small subunit to correctly bind mRNA and fmet-tRNAf.
- It dissociates prior to binding of the large subunit.
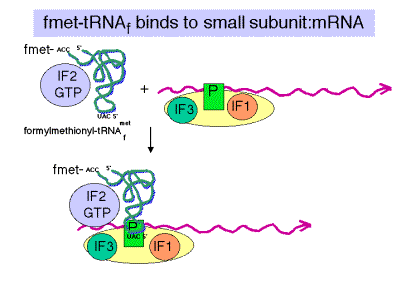
c. IF2
- Brings fmet‑tRNAf to the partial P site on the small subunit.
- At least in eukaryotes, it does this in a ternary complex with IF2, fmet‑tRNAf and GTP. In bacteria, the GTP may bind the initiation complex separately. [In some texts, such as MBOG, p. 412, the GTP-IF2 complex binds to the 30S subunit separately from fmet-tRNAf. How would you test the differences in these two models?]
- IF2 activates a GTPase activity in the small subunit. The resulting change in conformation may allow the large subunit to bind.
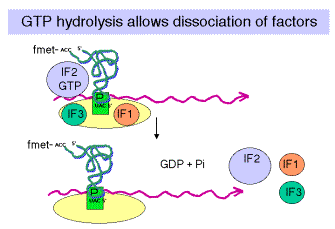
3. Binding of 50S (large) subunit to initiation complex gives a complete ribosome ready for the elongation phase of translation. Note that f-met-tRNAfmet is positioned at the P site. It has recognized the initiator AUG in the mRNA.
4. Identification of initiator AUG in eukaryotes
a. Bases around AUG influence efficiency of initiation.
- The most important effects are from a purine 3 nt before AUG and a G after it. The preferred context is RNNAUGG.
- The consensus sequence for a large number of mRNAs is GCCRCCAUGG, but these other nucleotides have little effect in mutagenesis experiments.
a. Modified scanner model
(1) The mRNA is "prepared" for binding to the ribosome by the action of eukaryotic initiation factor 4, abbreviated eIF4 (Figure 3.5.16). eIF4 is a multisubunit factor; it includes a cap‑binding protein, eIF4F, that recognizes the 5' cap structure. It also includes proteins eIF4A and eIF4B. These are RNA helicases, which unwind secondary structures in the 5' untranslated region of the mRNA at the expense of ATP hydrolysis.
The mRNA then binds to the small ribosomal subunit. The met-tRNAi has already been brought to the small ribosomal subunit by eIF2, in a complex with GTP.
eIF3 keeps the small ribosomal subunit apart from the large subunit during the process of binding the mRNA.
(2) The small subunit, with associated factors, scans along the mRNA until it reaches (usually) the first AUG. Factors eIF1 and eIF1A help move the preinitiation complex to the AUG start.
Figure 3.5.16.
H. The elongation cycle during translation
1. Binding of aminoacyl‑tRNA to the A site
Recent review: Weijland, A. and A. Parmeggiani (1994) TIBS 19:188-193. Schroeder, R. (1994) Nature 370:597.
a. Elongation factor EF‑Tu
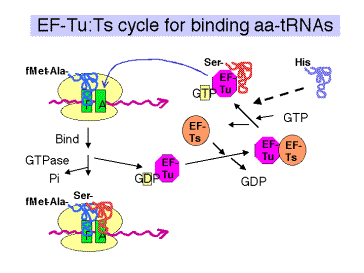
- The ternary complex of aminoacyl‑tRNA, EF‑Tu, and GTP brings the aminoacyl‑tRNA to the A site on the 70S ribosome (fig. 3.5.17).
- After the aminoacyl‑tRNA is deposited at the A site of the ribosome, the GTP is cleaved to GDP + Pi. The binary complex of EF‑Tu and GDP dissociates from the ribosome.
- This is one of many examples of guanine‑nucleotide‑binding proteins that are active when GTP is bound and inactive when GDP is bound.
The general model is that the GTP-bound state of EF-Tu adopts a conformation with a high affinity for aminoacyl-tRNA. The conformation (shape, charge density, etc.) of the resulting ternary complex (containing EF-Tu,GTP, and aminoacyl-tRNA) then allows it to bind to the A site of the ribsosome. Hydrolysis of GTP to form GDP and inorganic phosphate causes the EF-Tu to adopt a different conformation. The aminoacyl-tRNA now has a lower affinity for EF-Tu in the GDP bound state, and presumably a higher affinity for the A site on the ribosome, so it stays on the ribosome when EF-Tu in the GDP bound state dissociates (both from aminoacyl-tRNA and from the ribosome).
Figure 3.5.17.
(4) EF‑Tu is one of the most abundant proteins in E. coli, at 70,000 copies per cell. This is almost equal to the number of aminoacyl‑tRNAs per cell, so most of the aminoacyl‑tRNAs are likely to be in the ternary complex when the concentration of GTP is sufficiently high.
b. GTP
- Required for binding aminoacyl‑tRNA.
- Hydrolysis promotes dissociation of the complex EF‑Tu plus GDP from the ribosome.
c. EF‑Ts
- Aids in the recycling of EF‑Tu by GDP‑GTP exchange.
- EF‑Ts binds to EF‑Tu complexed with GDP, causing dissociation of GDP. GTP can now bind to the EF‑Tu‑Ts complex, causing EF‑Ts to dissociate and leaving EF‑Tu complexed with GTP. This latter binary complex is ready to bind another aminoacyl‑tRNA.
d. The antibiotic kirromycin prevents release of EF‑Tu‑GDP, thereby blocking elongation. This demonstrates that one step must be completed before the next can take place, and illustrates the importance of the EF‑Tu‑GTP/GDP cycle.
2. Peptidyl transferase on the large ribosomal subunit
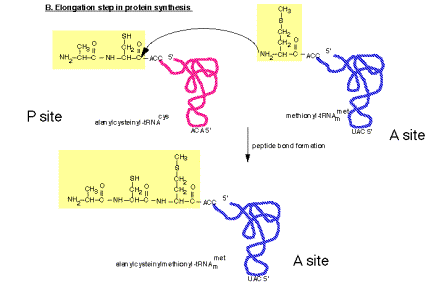
a. The peptidyl transferase reaction occurs via nucleophilic displacement. The amino group from aminoacyl‑tRNA (position n) attacks the "C‑terminal" carboxyl group of peptidyl‑tRNA (position n‑1in the mRNA). This results in cleavage of the high energy peptidyl‑tRNA ester linkage, thereby providing the free energy to drive the reaction. The resulting products of the reaction are deacylated tRNA at the P site and peptidyl‑tRNA at the A site.
Figure 3.5.18. Peptidyl transferase reaction
b. Role of rRNA in catalysis
It is likely that rRNA provides the catalytic center for the peptidyl transferase activity, with perhaps some ribosomal proteins aiding in holding the rRNA in the correct conformation for catalysis. This conclusion is supported by several lines of investigation, some of which are listed below.
- No protein, singly or in combination with other proteins, has been shown to catalyze peptide bond formation.
- Specific regions of 16S rRNA (in the small subunit) interact with the anticodon regions of tRNA in both the A and P sites. In contrast, 23S rRNA in the large subunit interacts with the CCA terminus of peptidyl‑tRNA, thus placing it in the right location for peptidyl transferase.
- The antibiotics erythromycin and chloramphenicol block peptidyl transferase. Some mutations that confer resistance to them map to the 23S rRNA sequence (others map to some 50S ribosomal proteins).
- A preparation consisting of 23S rRNA and some remnants of large subunit proteins retains peptidyl transferase activity. For more information, see Noller et al. (1992) Unusual resistance of peptidyl transferase to protein extraction procedures. Science 256: 1416-1419.
- Ribozyme RNAs can be selected that catalyze peptide bond formation. In this experiment, the investigators started with a pool of 1.3 ´ 1015 different RNAs of 72 nucleotides, flanked by constant regions. They let this large population of RNAs catalyze a peptide bond formation that adds a biotinyl-labeled amino acid (in a chemical mimic of a P site) to an amino acid connected to the RNA (in a chemical mimic of an A site). The RNAs that successfully catalyzed the reaction were extremely rare, but were now covalently attached to a biotin label. Thus they could be selected from the population by binding to streptavidin. PCR was used to amplify the successful RNAs, and the procedure repeated 19 times. At this point, the investigators characterized 9 RNAs that catalyzed the reaction. They found that these RNAs increased the reaction rate by a factor of 106 over the uncatalyzed reaction.
- The three-dimensional structure of the ribosome shows that the active site is comprised of RNA. The structure of a ribosome crystallized with an active site directed inhibitor has been determined, as well as the structure without the inhibitor. This allowed researchers to see precisely where the peptidyl transferase active site is within the structure. Only RNA is seen around this site. The nearest protein is 20 Angstroms away, too far to participate in catalysis.
3. Translocation
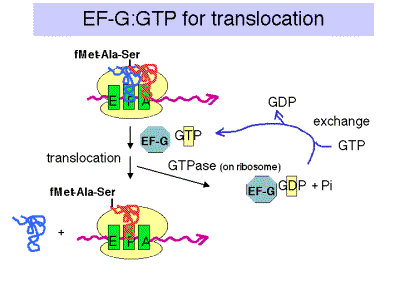
a. The translocation step moves the ribosome another 3 nucleotides downstream (one codon) and moves peptidyl‑tRNA to the P site (position n), deacylated tRNA exits through the E site, and the A site (position n+1) is vacant for another round of elongation.
b. Elongation Factor G = EF‑G
- This is another very abundant protein, with about 20,000 copies per cell, which is equivalent to the number of ribosomes.
- EF‑G‑GTP binds to the ribosome to aid translocation, and is released upon GTP hydrolysis (GTPase is from some ribosomal component).
- Recent structural studies (from A. Dahlberg and colleagues) show that EF-G in the GTP-bound state has a shape similar to that of the ternary complex of EF-Tu, GTP and aminoacyl-tRNA. Like the latter ternary complex, EF-G in the GTP-bound state also has a high affinity for the A site on the ribosome. This may help drive the movement of the peptidyl-tRNA from the A site to the P site, replacing it with EF-G (GTP) in the A site.
c. Hydrolysis of GTP is required for dissociation of EF‑G after translocation. The GTPase is part of the ribosome, not EF-G.
Figure 3.5.20.
d. Action of fusidic acid revealed the need for release of EF‑G‑GDP. In the presence of fusidic acid, EF‑G‑GTP binds the ribosome, GTP is hydrolyzed, and the ribosome moves three nucleotides. But the ribosome‑EF‑G‑GDP complex is stabilized by this compound, and translation is halted.
e. Ribosomes cannot bind EF‑Tu and EF‑G simultaneously. EF‑Tu must finish its action before EF‑G can act, and EF‑G must complete its cycle before EF‑Tu can act again to bring in another aminoacyl‑tRNA.
f. Effect of diptheria toxin
- The eukaryotic analog to EF‑G is eEF2, which is also a translocase dependent on GTP hydrolysis. It is also is blocked by fusidic acid.
- Diptheria toxin will catalyze the addition of ADP‑ribose (from substrate NAD+) to eEF2, thereby inactivating it. The target for ADP‑ribosylation is modified histidine found in eEF2 from many species.
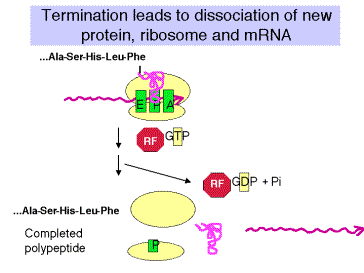
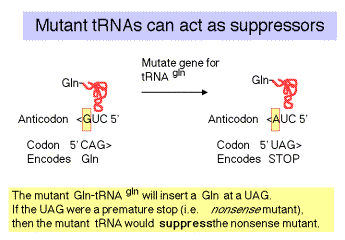
c. The "down side" to nonsense suppression is that the suppressor tRNA can act at any amber codon, Therefore it competes with the releasing factors in recognizing the normal termination codons. When the suppressor tRNA is used instead of releasing factors, translation proceeds further down the mRNA than it is supposed to, leading to production of aberrant proteins. Suppressor strains of E. colican be pretty sick (i.e. they don't grow as well as wild type strains).
d. Two other amber suppressors are encoded by the supDgene, which encodes a tRNA that will insert Ser at a UAG, and supF, which will insert Tyr.
3. Missense suppressors: These are mutant tRNAs that lead to the insertion of an amino acid that is compatible with the wild type amino acid (altered by the original mutation).
4. Frameshift suppressors: These are mutant tRNAs whose anticodon has been expanded (or contracted?) to match the length‑altering mutation in the mRNA.
E.g. Consider an original mutation 5'GGG ‑> 5'GGGG (insert a G).
A frameshift suppressor would also have an additional C in the anticodon.
wt tRNA anticodon 3'CCC ‑‑> suppressor tRNA 3'CCCC.


