2.4: Bayesian Statistics
- Page ID
- 21582
Recent years have seen tremendous growth of Bayesian approaches in reconstructing phylogenetic trees and estimating their branch lengths. Although there are currently only a few Bayesian comparative methods, their number will certainly grow as comparative biologists try to solve more complex problems. In a Bayesian framework, the quantity of interest is the posterior probability, calculated using Bayes' theorem:
\[ Pr(H|D) = \dfrac{Pr(D|H) \cdot Pr(H)}{Pr(D)} \label{2.19}\]
The benefit of Bayesian approaches is that they allow us to estimate the probability that the hypothesis is true given the observed data, \(Pr(H|D)\). This is really the sort of probability that most people have in mind when they are thinking about the goals of their study. However, Bayes theorem also reveals a cost of this approach. Along with the likelihood, \(Pr(D|H)\), one must also incorporate prior knowledge about the probability that any given hypothesis is true - Pr(H). This represents the prior belief that a hypothesis is true, even before consideration of the data at hand. This prior probability must be explicitly quantified in all Bayesian statistical analyses. In practice, scientists often seek to use “uninformative” priors that have little influence on the posterior distribution - although even the term "uninformative" can be confusing, because the prior is an integral part of a Bayesian analysis. The term \(Pr(D)\) is also an important part of Bayes theorem, and can be calculated as the probability of obtaining the data integrated over the prior distributions of the parameters:
\[Pr(D)=∫_HPr(H|D)Pr(H)dH \label{2.20}\]
However, \(Pr(D)\) is constant when comparing the fit of different models for a given data set and thus has no influence on Bayesian model selection under most circumstances (and all the examples in this book).
In our example of lizard flipping, we can do an analysis in a Bayesian framework. For model 1, there are no free parameters. Because of this, \(Pr(H)=1\) and \(Pr(D|H)=P(D)\), so that \(Pr(H|D)=1\). This may seem strange but what the result means is that our data has no influence on the structure of the model. We do not learn anything about a model with no free parameters by collecting data!
If we consider model 2 above, the parameter pH must be estimated. We can set a uniform prior between 0 and 1 for pH, so that f(pH)=1 for all pH in the interval [0,1]. We can also write this as “our prior for \(p_h\) is U(0,1)”. Then:
\[ Pr(H|D) = \frac{Pr(D|H) \cdot Pr(H)}{Pr(D)} = \frac{P(H|p_H,N) f(p_H)}{\displaystyle \int_{0}^{1} P(H|p_H,N) f(p_h) dp_H} \label{2.21} \]
Next we note that \(Pr(D|H)\) is the likelihood of our data given the model, which is already stated above as Equation \ref{2.2}. Plugging this into our equation, we have:
\[ Pr(H|D) = \frac{\binom{N}{H} p_H^H (1-p_H)^{N-H}}{\displaystyle \int_{0}^{1} \binom{N}{H} p_H^H (1-p_H)^{N-H} dp_H} \label{2.22} \]
This ugly equation actually simplifies to a beta distribution, which can be expressed more simply as:
\[ Pr(H|D) = \frac{(N+1)!}{H!(N-H)!} p_H^H (1-p_H)^{N-H} \label{2.23} \]
We can compare this posterior distribution of our parameter estimate, pH, given the data, to our uniform prior (Figure 2.3). If you inspect this plot, you see that the posterior distribution is very different from the prior – that is, the data have changed our view of the values that parameters should take. Again, this result is qualitatively consistent with both the frequentist and ML approaches described above. In this case, we can see from the posterior distribution that we can be quite confident that our parameter pH is not 0.5.
As you can see from this example, Bayes theorem lets us combine our prior belief about parameter values with the information from the data in order to obtain a posterior. These posterior distributions are very easy to interpret, as they express the probability of the model parameters given our data. However, that clarity comes at a cost of requiring an explicit prior. Later in the book we will learn how to use this feature of Bayesian statistics to our advantage when we actually do have some prior knowledge about parameter values.
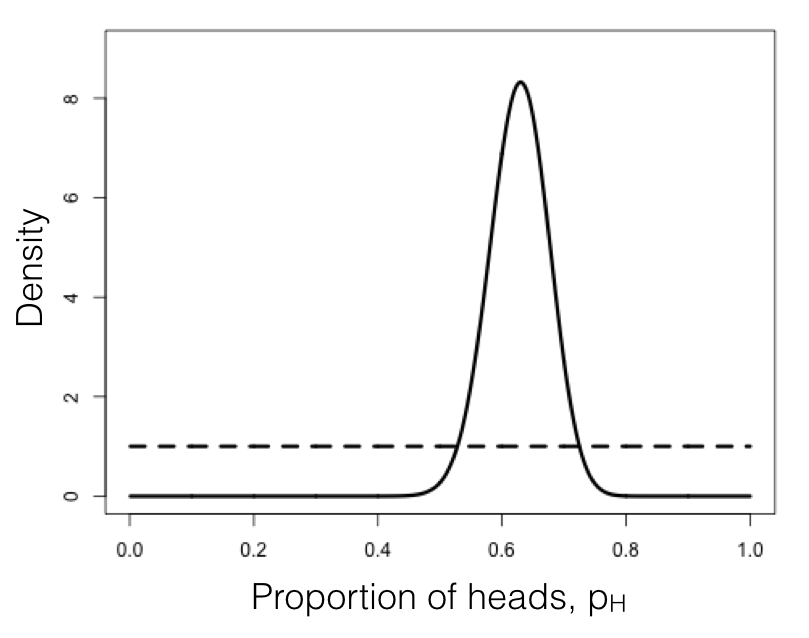
Figure 2.3. Bayesian prior (dotted line) and posterior (solid line) distributions for lizard flipping. Image by the author, can be reused under a CC-BY-4.0 license.
Section 2.4b: Bayesian MCMC
The other main tool in the toolbox of Bayesian comparative methods is the use of Markov-chain Monte Carlo (MCMC) tools to calculate posterior probabilities. MCMC techniques use an algorithm that uses a “chain” of calculations to sample the posterior distribution. MCMC requires calculation of likelihoods but not complicated mathematics (e.g. integration of probability distributions, as in Equation \ref{2.22}, and so represents a more flexible approach to Bayesian computation. Frequently, the integrals in Equation \ref{2.21} are intractable, so that the most efficient way to fit Bayesian models is by using MCMC. Also, setting up an MCMC is, in my experience, easier than people expect!
An MCMC analysis requires that one constructs and samples from a Markov chain. A Markov chain is a random process that changes from one state to another with certain probabilities that depend only on the current state of the system, and not what has come before. A simple example of a Markov chain is the movement of a playing piece in the game Chutes and Ladders; the position of the piece moves from one square to another following probabilities given by the dice and the layout of the game board. The movement of the piece from any square on the board does not depend on how the piece got to that square.
Some Markov chains have an equilibrium distribution, which is a stable probability distribution of the model’s states after the chain has run for a very long time. For Bayesian analysis, we use a technique called a Metropolis-Hasting algorithm to construct a special Markov chain that has an equilibrium distribution that is the same as the Bayesian posterior distribution of our statistical model. Then, using a random simulation on this chain (this is the Markov-chain Monte Carlo, MCMC), we can sample from the posterior distribution of our model.
In simpler terms: we use a set of well-defined rules. These rules let us walk around parameter space, at each step deciding whether to accept or reject the next proposed move. Because of some mathematical proofs that are beyond the scope of this chapter, these rules guarantee that we will eventually be accepting samples from the Bayesian posterior distribution - which is what we seek.
The following algorithm uses a Metropolis-Hastings algorithm to carry out a Bayesian MCMC analysis with one free parameter.
Metropolis-Hastings algorithm
- Get a starting parameter value.
- Sample a starting parameter value, p0, from the prior distribution.
- Starting with i = 1, propose a new parameter for generation i.
- Given the current parameter value, p, select a new proposed parameter value, p′, using the proposal density Q(p′|p).
- Calculate three ratios.
- a. The prior odds ratio. This is the ratio of the probability of drawing the parameter values p and p′ from the prior. \[R_{prior} = \frac{P(p')}{P(p)} \label{2.24}\]
- b. The proposal density ratio. This is the ratio of probability of proposals going from p to p′ and the reverse. Often, we purposefully construct a proposal density that is symmetrical. When we do that, Q(p′|p)=Q(p|p′) and a2 = 1, simplifying the calculations . \[R_{proposal} = \frac{Q(p'|p)}{Q(p|p')} \label{2.25}\]
- c. The likelihood ratio. This is the ratio of probabilities of the data given the two different parameter values. \[R_{likelihood} = \frac{L(p'|D)}{L(p|D)} = \frac{P(D|p')}{P(D|p)} \label{2.26}\]
- Multiply. Find the product of the prior odds, proposal density ratio, and the likelihood ratio.\[R_{accept} = R_{prior} ⋅ R_{proposal} ⋅ R_{likelihood} \label{2.27}\]
- Accept or reject. Draw a random number x from a uniform distribution between 0 and 1. If x < Raccept, accept the proposed value of p′ ( pi = p′); otherwise reject, and retain the current value p ( pi = p).
- Repeat. Repeat steps 2-5 a large number of times.
Carrying out these steps, one obtains a set of parameter values, pi, where i is from 1 to the total number of generations in the MCMC. Typically, the chain has a “burn-in” period at the beginning. This is the time before the chain has reached a stationary distribution, and can be observed when parameter values show trends through time and the likelihood for models has yet to plateau. If you eliminate this “burn-in” period, then, as discussed above, each step in the chain is a sample from the posterior distribution. We can summarize the posterior distributions of the model parameters in a variety of ways; for example, by calculating means, 95% confidence intervals, or histograms.
We can apply this algorithm to our coin-flipping example. We will consider the same prior distribution, U(0, 1), for the parameter p. We will also define a proposal density, Q(p′|p) U(p − ϵ, p + ϵ). That is, we will add or subtract a small number ( ϵ ≤ 0.01) to generate proposed values of p′ given p.
To start the algorithm, we draw a value of p from the prior. Let’s say for illustrative purposes that the value we draw is 0.60. This becomes our current parameter estimate. For step two, we propose a new value, p′, by drawing from our proposal distribution. We can use ϵ = 0.01 so the proposal distribution becomes U(0.59, 0.61). Let’s suppose that our new proposed value p′=0.595.
We then calculate our three ratios. Here things are simpler than you might have expected for two reasons. First, recall that our prior probability distribution is U(0, 1). The density of this distribution is a constant (1.0) for all values of p and p′. Because of this, the prior odds ratio for this example is always:
\[ R_{prior} = \frac{P(p')}{P(p)} = \frac{1}{1} = 1 \label{2.28}\]
Similarly, because our proposal distribution is symmetrical, Q(p′|p)=Q(p|p′) and Rproposal = 1. That means that we only need to calculate the likelihood ratio, Rlikelihood for p and p′. We can do this by plugging our values for p (or p′) into Equation \ref{2.2}:
\[ P(D|p) = {N \choose H} p^H (1-p)^{N-H} = {100 \choose 63} 0.6^63 (1-0.6)^{100-63} = 0.068 \label{2.29} \]
Likewise,
\[ P(D|p') = {N \choose H} p'^H (1-p')^{N-H} = {100 \choose 63} 0.595^63 (1-0.595)^{100-63} = 0.064 \label{2.30}\]
The likelihood ratio is then:
\[ R_{likelihood} = \frac{P(D|p')}{P(D|p)} = \frac{0.064}{0.068} = 0.94 \label{2.31}\]
We can now calculate Raccept = Rprior ⋅ Rproposal ⋅ Rlikelihood = 1 ⋅ 1 ⋅ 0.94 = 0.94. We next choose a random number between 0 and 1 – say that we draw x = 0.34. We then notice that our random number x is less than or equal to Raccept, so we accept the proposed value of p′. If the random number that we drew were greater than 0.94, we would reject the proposed value, and keep our original parameter value p = 0.60 going into the next generation.
If we repeat this procedure a large number of times, we will obtain a long chain of values of p. You can see the results of such a run in Figure 2.4. In panel A, I have plotted the likelihoods for each successive value of p. You can see that the likelihoods increase for the first ~1000 or so generations, then reach a plateau around lnL = −3. Panel B shows a plot of the values of p, which rapidly converge to a stable distribution around p = 0.63. We can also plot a histogram of these posterior estimates of p. In panel C, I have done that – but with a twist. Because the MCMC algorithm creates a series of parameter estimates, these numbers show autocorrelation – that is, each estimate is similar to estimates that come just before and just after. This autocorrelation can cause problems for data analysis. The simplest solution is to subsample these values, picking only, say, one value every 100 generations. That is what I have done in the histogram in panel C. This panel also includes the analytic posterior distribution that we calculated above – notice how well our Metropolis-Hastings algorithm did in reconstructing this distribution! For comparative methods in general, analytic posterior distributions are difficult or impossible to construct, so approximation using MCMC is very common.
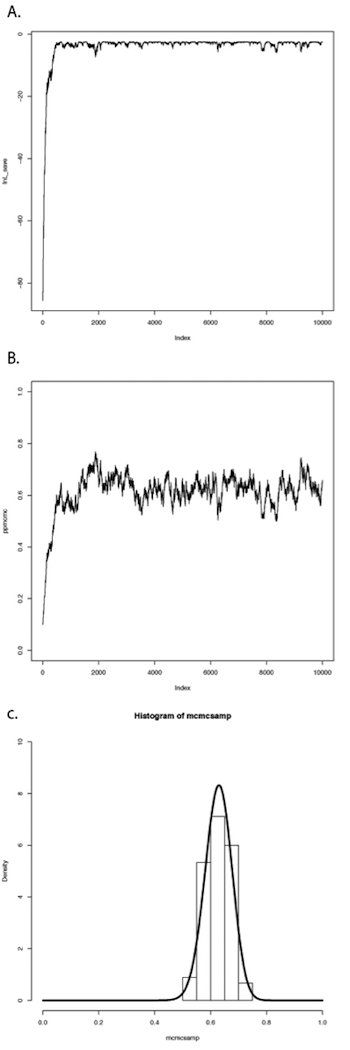
This simple example glosses over some of the details of MCMC algorithms, but we will get into those details later, and there are many other books that treat this topic in great depth (e.g. Christensen et al. 2010). The point is that we can solve some of the challenges involved in Bayesian statistics using numerical “tricks” like MCMC, that exploit the power of modern computers to fit models and estimate model parameters.
Section 2.4c: Bayes factors
Now that we know how to use data and a prior to calculate a posterior distribution, we can move to the topic of Bayesian model selection. We already learned one general method for model selection using AIC. We can also do model selection in a Bayesian framework. The simplest way is to calculate and then compare the posterior probabilities for a set of models under consideration. One can do this by calculating Bayes factors:
\[ B_{12} = \frac{Pr(D|H_1)}{Pr(D|H_2)} \label{2.32}\]
Bayes factors are ratios of the marginal likelihoods P(D|H) of two competing models. They represent the probability of the data averaged over the posterior distribution of parameter estimates. It is important to note that these marginal likelihoods are different from the likelihoods used above for AIC model comparison in an important way. With AIC and other related tests, we calculate the likelihoods for a given model and a particular set of parameter values – in the coin flipping example, the likelihood for model 2 when pH = 0.63. By contrast, Bayes factors’ marginal likelihoods give the probability of the data averaged over all possible parameter values for a model, weighted by their prior probability.
Because of the use of marginal likelihoods, Bayes factor allows us to do model selection in a way that accounts for uncertainty in our parameter estimates – again, though, at the cost of requiring explicit prior probabilities for all model parameters. Such comparisons can be quite different from likelihood ratio tests or comparisons of AICc scores. Bayes factors represent model comparisons that integrate over all possible parameter values rather than comparing the fit of models only at the parameter values that best fit the data. In other words, AICc scores compare the fit of two models given particular estimated values for all of the parameters in each of the models. By contrast, Bayes factors make a comparison between two models that accounts for uncertainty in their parameter estimates. This will make the biggest difference when some parameters of one or both models have relatively wide uncertainty. If all parameters can be estimated with precision, results from both approaches should be similar.
Calculation of Bayes factors can be quite complicated, requiring integration across probability distributions. In the case of our coin-flipping problem, we have already done that to obtain the beta distribution in Equation \ref{2.22. We can then calculate Bayes factors to compare the fit of two competing models. Let’s compare the two models for coin flipping considered above: model 1, where pH = 0.5, and model 2, where pH = 0.63. Then:
\[ \begin{array}{lcl} Pr(D|H_1) &=& \binom{100}{63} 0.5^{0.63} (1-0.5)^{100-63} \\\ &=& 0.00270 \\\ Pr(D|H_2) &=& \int_{p=0}^{1} \binom{100}{63} p^{63} (1-p)^{100-63} \\\ &=& \binom{100}{63} \beta (38,64) \\\ &=& 0.0099 \\\ B_{12} &=& \frac{0.0099}{0.00270} \\\ &=& 3.67 \\\ \end{array} \label{2.33}\]
In the above example, β(x, y) is the Beta function. Our calculations show that the Bayes factor is 3.67 in favor of model 2 compared to model 1. This is typically interpreted as substantial (but not decisive) evidence in favor of model 2. Again, we can be reasonably confident that our lizard is not a fair flipper.
In the lizard flipping example we can calculate Bayes factors exactly because we know the solution to the integral in Equation \ref{2.33. However, if we don’t know how to solve this equation (a typical situation in comparative methods), we can still approximate Bayes factors from our MCMC runs. Methods to do this, including arrogance sampling and stepping stone models (Xie et al. 2011; Perrakis et al. 2014), are complex and beyond the scope of this book. However, one common method for approximating Bayes Factors involves calculating the harmonic mean of the likelihoods over the MCMC chain for each model. The ratio of these two likelihoods is then used as an approximation of the Bayes factor (Newton and Raftery 1994). Unfortunately, this method is extremely unreliable, and probably should never be used (see Neal 2008 for more details).


