5.1: Splitting up the A/P axis: Beginning-Hox Genes, Another Level of Regionalization
- Page ID
- 21539
Broad Scale Regionalization: Hox Genes and the Segmentation Cascade
We already took a quick look at Hox genes, both in your readings and in the Genetic Toolkit section. Now we will put them into a broader Evo-Devo context by looking at a specific example of Hox patterning (following up the The curious case of Bicoid story) and by seeing evidence that regionalization by Hox genes is conserved across animals. Some cis-regulatory elements driving Hox gene expression patterns are also conserved. However, the earlier developmental events that set up Hox expression domains are not conserved across animals, they differ widely sometimes even within a phylum. For example, even though all arthropods are segmented in Drosophila segments are patterned simultaneously, while in some other insects and arthropods they arise one at a time. The conservation of the Hox pattern despite the variability of broader A/P pattern generation suggests that Hox patterning represents a developmental constraint in animal evolution. That is, once the Hox pattern evolved, it was so advantageous that it was mostly retained for millions of years of animal evolution.
We focus on Drosophila even though it is an odd case of simultaneous regionalization because it is very well-studied and easier to understand than many other model organisms. Our look at Drosophila A/P regionalization and patterning will illustrate three major concepts in Evo-Devo:
- There are multiple ways to get to constrained developmental stages.
- Much of development occurs by repeatedly dividing up tissues/regions and specifying them into more restricted fates.
- Multiple gene expression outputs can overlie each other to give a cell/tissue its specific identity.
In cleavage and gastrulation, we looked at the idea of symmetry breaking in creating axes and germ layers in animals. We also saw that opposing Bicoid and Nanos protein gradients are able to specify the first anteroposterior regionalization in developing fruit flies. But if we think about an adult fly, they have many more anteroposterior regions than simply a head and a tail. For example, only some segments bear wings or legs, other bear eyes or antennae and others have genitalia or excretory organs. How does the developing fly "regionalize" - that is go from having two opposing anteroposterior gradients to having fine-scale anteroposterior patterning? To answer this, we will look at the developmental genetics of the segmentation cascade - the series of steps that divides the embryonic fly into finer and finer subdivisions.
The Drosophila Segmentation Cascade
Here I will very briefly summarize the segmentation cascade, for more details please visit The Origins of Anterior-Posterior Polarity
The segmentation cascade is hierarchical in both developmental time and regulation of gene expression. Maternal effect genes are expressed earliest and they regulate the gap genes. Gap genes are expressed next, acting with the maternal effect genes to regulate the pair-rule genes. Pair-rule genes and gap genes regulate the expression of the segment-polarity genes, which are expressed last (Figure 1).
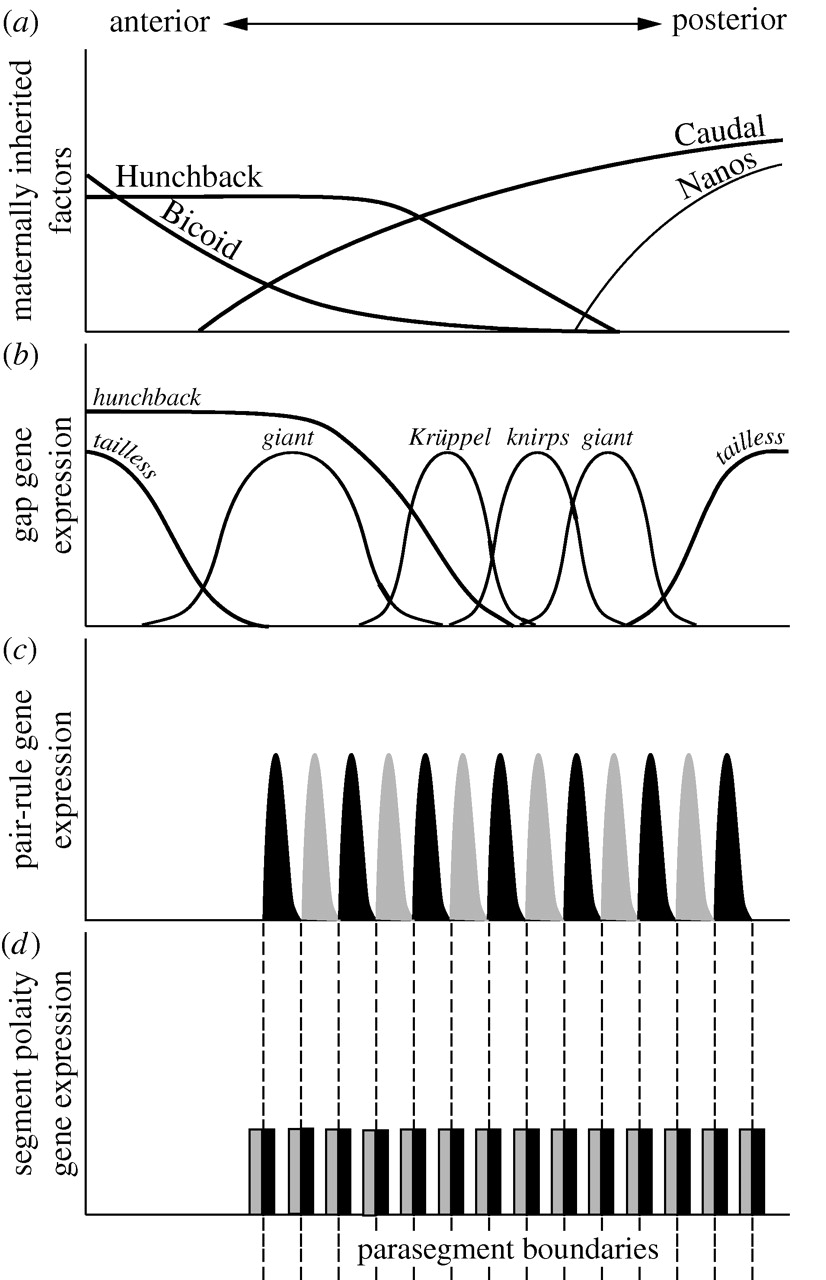 |
Figure 1: The Drosophila segmentation cascade. Drosophila divides up its segments all at once, but most arthropods (an animals in general) grow from a posterior growth zone where anterior-most regions develop first. Consider what parts of this expression and specification hierarchy might be conserved across different modes of development and which parts could not be. From "The evolution of developmental gene networks: lessons from comparative studies on holometabolous insects" Andrew Peel, 2008 in Philosophical Transactions of the Royal Society, B. Published under The Royal Society Academic Institution Single Site Licence DOI: 10.1098/rstb.2007.2244 |
Maternal effect genes: Bicoid, Nanos, Hunchback, and a handful of other Drosophila genes are known as "maternal effect" genes. This means that if you mutate them in a female fly, the mutant phenotype will be seen in her offspring. Since you already know that Bicoid and Nanos mRNAs are made by the mother's cells and pumped into the oocyte before fertilization it should be pretty clear why this is. The maternal effect genes set up broad anterior and posterior regions in the fruit fly as well as dorsal/ventral regions (which we will not cover here, though it's a pretty juicy story).
Gap genes: The maternal effect genes are transcription and translation factors that affect the expression of the gap genes. The gap genes are transcription factors that are expressed in broad strips along the A/P axis (Figure 2). They are called gap genes because if one is mutated it leads to a "gap" in the embryo - the larva will be missing a large portion of its body2.
Figure 2: Gap genes. Left panel shows mRNA expression of three Drosophila gap genes, from Wikimedia commons by user Celefin. This is a derivative of an original figure by Haecker A, Qi D, Lilja T, Moussian B, Andrioli LP, Luschnig S, Mannervik, M. DOI 10.1371/journal.pbio.0050145 published under a CC BY 2.5 license Right panel shows the effect of a homozygous Kruppel mutation in an embryo. Grey shapes are denticle bands (bristles) found in each abdominal segment. The Kruppel mutant has lost 6 of these bands, corresponding with Kruppel expression over the first 6 abdominal segments. Right panel by Amanda Lo and Ajna Rivera. An editable svg version of this figure is available at https://scholarlycommons.pacific.edu/open-images/24/
Pair-rule genes: Gap genes and maternal effect genes affect the expression of the pair-rule genes - which are each expressed either in odd or even segments2. These are particularly interesting because these genes bridge the gap between regionalization and modularity. They are expressed in a modular pattern, but get their input from a regionalization pattern. Genetically, this means that the expression of each of these genes is activated by multiple combinations of gap and maternal effect proteins. For example, the even-skipped gene is expressed in 7 stripes along the A/P axis. It has 4 major cis-regulatory enhancers that control this. One element controlling stripes 3 and 7 has binding sites for Hunchback and Knirps, both of which repress the expression driven by the 3+7 enhancer (Figure 2). This enhancer is activated by the ubiquitous (expressed everywhere) activator Zelda3. The element controlling stripe 2 is even more complex with binding sites for Bicoid, Hunchback, Caudal, Kruppel, and Knirps (Figure 3)4.

Segment polarity genes: Segment polarity genes are expressed in either the anterior or posterior of each segment and help to define segment boundaries as well as the A/P patterning of each segment (Figure 4). Like the pair-rule genes, these are expressed in stripes along the entire A/P axis, rather than in a regional pattern like Maternal Effect and Gap genes. The expression domains of the segment polarity genes are regulated largely by the pair-rule and gap genes.
Hox genes: Another Level of Regionalization
While the maternal effect genes and the gap genes regionalize the Drosophila embryo to some extent, the adult animal will have an even more regionalized body. The Hox gene pattern provides segment-level regionalization. It specifies where legs, wings, antennae, mouthparts, genitalia, and other unique structures will go (Figure 5). As you already know, most animals use the Hox genes to do A/P patterning and it's a bit of a mystery how and why so many diverse early developmental strategies coalesce onto the same A/P patterning, and then diverge into diverse body plans later in development.
How conserved is the Hox patterning? First, nearly all animals have Hox genes and many of them are syntenic - meaning that the order of the Hox genes is conserved along the chromosome. This is related to the regulation of the Hox genes, they have distal regulatory factors (one of which we will see later in Ectodermal Appendages) but also have proximal regulatory factors. In vertebrates, some genes in the cluster can share regulatory factors5. Because of this, gene order matters. An analogy is: if you share a car with your neighbor, you cannot move to a new neighborhood without losing access to the car. This sharing of elements is an evolutionary constraints that has resulted in the conservation of these regulatory factors and the preservation of synteny of the Hox genes. In invertebrate lineages, synteny may be preserved due to tight clustering of the genes with each other and their proximal cis regulatory elements5. Some species and lineages however, have lost this conserved synteny, notably the nematodes and the tunicates5. Despite these losses, synteny still seems to be the rule with at least some conservation of gene order found in most phyla examined, including the most basally branching Hox-containing phylum, the cnidarians6.
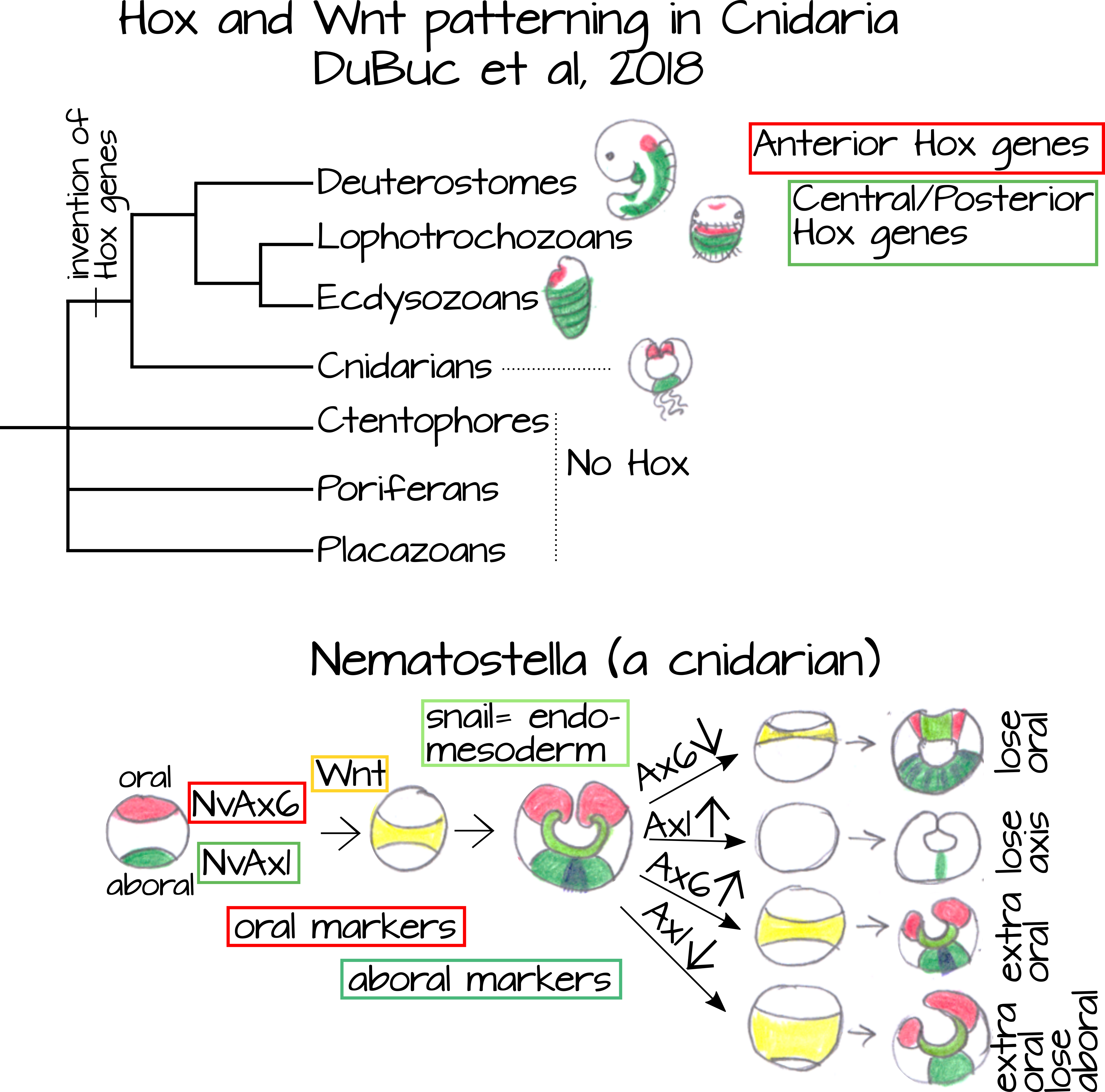
Cnidarians (such as jellies and sea anemones) are an interesting case in which to look at the Hox genes because while the Hox genes are known to be conserved A/P patterning elements, the cnidarians lack an A/P axis. How can they be missing something so important? Well it's because of how we define anterior and posterior. A standard definition is that anterior is towards the head and posterior is towards the tail. What about animals that don't have an obvious head? We count their mouth and any conglomeration of major sensory organs as the head end and keep moving along their long axis to the nearest tip of the body. Typically, along the long axis of the body, the tip nearest the mouth is the anterior end and the tip nearest the anus is the posterior end. Cnidarians, however, not only lack a head they also lack major sensory organs and a mouth. Instead they have a ring of tentacles (that can point up or down depending on the species) that surrounds a single opening that serves as a mouth and anus.
Many evolutionary scenarios have been proposed to homologize the cnidarian "radial" axis with the A/P and D/V bilaterian axes. Hox genes are one way to look at this. DuBuc et al6 found that cnidarian homologs of anterior Hox genes (red in Figures 6 and 7) are expressed at the oral end of an anemone (Cnidaria) embryo and cnidarian homologs of central/posterior Hox genes (blue/green in Figure 6) are expressed at the aboral end (the end opposite the mouth/anus). They also found that Wnt, a signal transduction gene involved in many developmental processes including anteroposterior specification, was expressed between the expression domains of Ax6,an anterior Hox gene, and Ax1, a posterior Hox gene