10.7: Eukaryotic Translation
- Page ID
- 17701
Eukaryotic translation, as with transcription, is satisfyingly similar (from a student studying point of view, or from an evolutionary conservation one) to the prokaryotic case. The initiation process is slightly more complicated, but the elongation and termination processes are the same, but with eukaryotic homologues of the appropriate elongation and release factors.
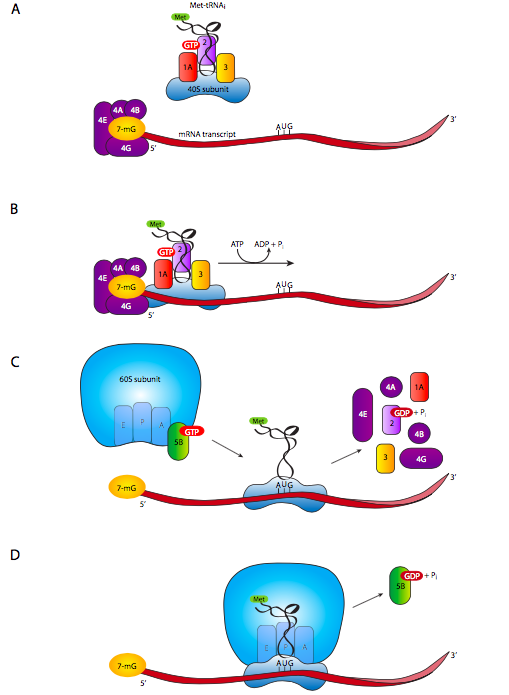
For eukaryotes, each mRNA encodes one and only one gene (as opposed to multi-gene transcripts such as operons), so there isn’t much question of which AUG is a start codon, and which are just regular methionines. Therefore, there is no requirement for a Shine-Dalgarno sequence in eukaryotes. The small ribosomal subunit, accompanied by eukaryotic initiation factors eIF-3, eIF-2, and Met-tRNAi, together known as the terneary complex, binds to eIF-1A. Meanwhile, eIF-4A, -4B, -4E, and -4G bind to the 5’ (7-methyguanosine) cap of the mRNA ( g. 7A). The small subunit complex and the eIF4/ mRNA cap-binding complex interact to form the 43S complex, which then begins scanning the mRNA from 5’ to 3’ looking for the first AUG.
Usually, but not always, the first AUG is the start codon for eukaryotic genes. However, the context of the AUG matters, and it is a much stronger (i.e. more frequently recognized and used) start codon if there is a purine residue (A or G) at -3 and a G at +4. See Kozak, M., Biochimie 76: 815-821, 1994.
Once the 43S scanning complex has found the start codon, the initiation factors drop off, and the large ribosomal subunit arrives. The large ribosomal subunit has bound eIF-6, which prevents it from reassociating with small subunits, and its removal is required first. Another factor, eIF-5 enters the scene during the coupling process between the large and small ribosomal subunits, and hydrolysis of an eIF-5-attached GTP is required to complete the docking of the subunits and the formation of a complete functional ribosome on the mRNA.
Elongation is functionally the same as in prokaryotes except that the functions of EF-Tu is taken care of by EF-1a, also with hydrolysis of GTP. EF-2 is the eukaryotic analog of EF-G, and utilizes GTP hydrolysis for translocation of the ribosome. Termination uses eukaryotic homologues of the release factors, though eRF-1 takes the place of both prokaryotic RF-1 and RF-2.
Although polyribosomes (aka polysomes) can form on both prokaryotic and eukaryotic mRNAs, eukaryotic polysomes have an additional twist. Technically, a polysome is simply an mRNA with multiple ribosomes translating it simultaneously, but in eukaryotes, the polysome also has a unique morphology because it utilizes PABPI, or poly-A binding protein. This protein not only binds to the 3’ poly-A tail of an mRNA, it also interacts with the eIF-4 initiation factors, which thus loops the mRNA into a circular shape. That way, once the ribosome reaches the end of the gene and releases from the mRNA, it is physically near the beginning of the mRNA to start translating again.

