10.6: Prokaryotic Translation
- Page ID
- 16154
As soon as the RNA has emerged from the RNAP and there is sufficient space to accommodate a ribosome, translation can begin in prokaryotes. In fact, for highly expressed genes, it would not be unusual to see multiple RNA polymerases transcribing the DNA and multiple ribosomes on each of the transcripts translating the mRNA to protein! The process begins with the small ribosomal subunit (and only the small subunit - if it is attached to the large subunit, it is unable to bind the mRNA), which binds to the mRNA loosely and starts to scan it for a recognition sequence called the Shine-Dalgarno sequence, after its discoverers. Once this is recognized by the small ribosomal subunit rRNA, the small subunit is positioned around the start codon (AUG). This process is facilitated by initiation factors as follows.
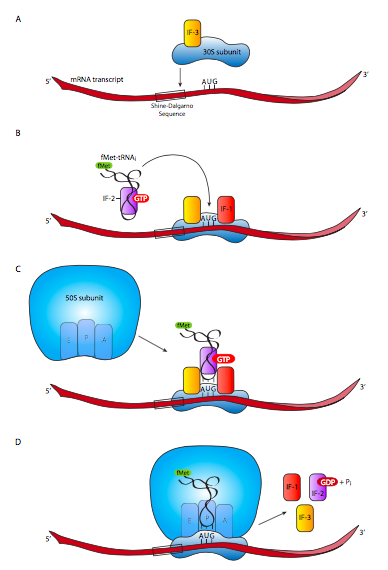
The 30S ribosomal subunit dissociates from the 50S ribosomal subunit if it was associated with one, and binds to intiation factors IF-1 and IF-3. IF-1 binds to the A site, where it prevents new aminoacyl-tRNA molecules from entering before the full ribosome is assembled. It also facilitates the assembly and stabilization of the initiation complex. IF-3 is required to allow the 30S subunit to bind to mRNA. Once this has occurred, IF-2-GTP arrives on scene, carrying with it the initiator aminoacyl-tRNA. This settles into the P site, which is positioned so that the anticodon of the tRNA settles over the AUG start codon of the mRNA. Hydrolysis of the GTP attached to IF-2 and release of all the initiation factors is needed to allow the 50S subunit to bind to the 30S subunit to form the full and fully functional ribosome. Because GTP hydrolysis was required, the joining of the subunits is irreversible spontaneously, and requires expenditure of energy upon termination of translation. Once the 50S subunit joins with the 30S subunit, the A site is ready to accept the next aminoacyl-tRNA.
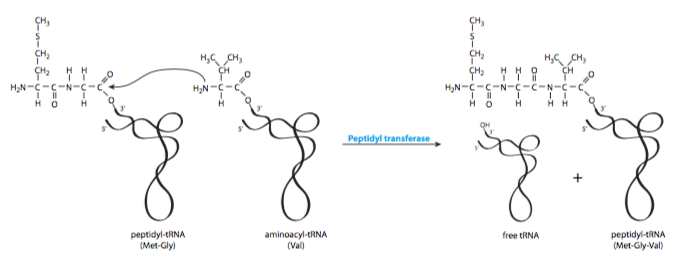
A common and understandable misconception is that the new amino acid brought to the ribosome is added onto the growing polypeptide chain. In fact, the mechanism is exactly the opposite: the polypeptide is added onto the new amino acid (Figure \(\PageIndex{4}\)). This begins from the second amino acid to be added to a new protein (Figure \(\PageIndex{5}\)). The rst amino acid, a methionine, you should recall, came in along with IF-2 and the initiator tRNA. The new aminoacyl-tRNA is escorted by EF-Tu, an elongation factor that carries a GTP. Once the aa-tRNA is in place, EF-Tu hydrolyzes the GTP and dissociates from the aminoacyl-tRNA and ribosome.
For a long time, there was a bit of mystery surrounding the simultaneous docking of two tRNA molecules on immediately adjacent codons of mRNA. Under normal conditions, there should not be enough room, since the tRNAs are fairly bulky and one should obstruct the other from reaching the mRNA to make a codon-anticodon match. The matter was finally cleared up in 2001 with x-ray crystallographic examinations showing a bend in the mRNA between the codon in the P slot and the codon in the A slot. The bend puts the two associated tRNAs at slightly different angles and thus creates just enough room for both to maintain basepair- ing hydrogen bonds with the mRNA. See Yusupov et al, Science 292 (5518): 883-896, 2001.
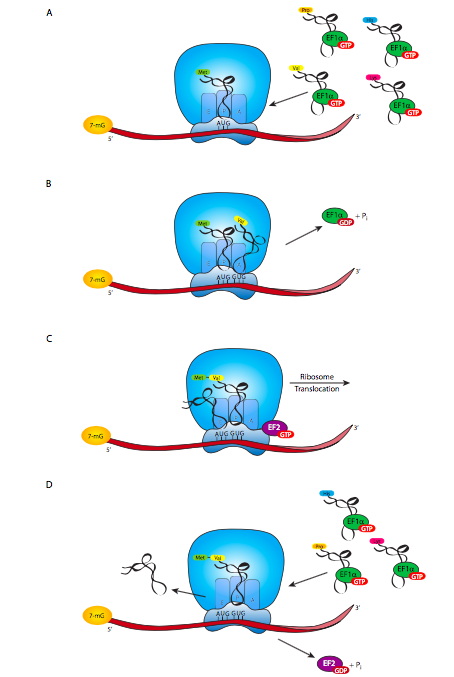
When a new aminoacyl-tRNA drops into the A slot of the ribosome, the anticodon is lined up with the codon of the mRNA. If there is no complementarity, the aminoacyl-tRNA soon floats back out of the slot to be replaced by another candidate. However, if there is complementarity (or something close enough, recalling the idea of wobble) then H-bonds form between the codon and anti-codon, the tRNA changes conformation, which shifts the conformation of EF-Tu, causing hydrolysis of GTP to GDP + Pi, and release from the aa-tRNA. The codon-anticodon interaction is stable long enough for the catalytic activity of the ribosome to hydrolyze the bond between fMet and the tRNAf in the P slot, and attach the fMet to the new amino acid with a peptide bond in the A slot. The new amino acid is still attached to its tRNA, and as this process occurs, the ribosome shifts position with respect to the mRNA and tRNAs. This puts the now- empty (no amino acid attached) tRNAf in the E slot, the tRNAaa in the P slot, attached to that aa which is bonded to Met, and the A slot is again open for a new tRNA to come in. The elongation factor EF-G binds near the A slot as soon as EF-Tu leaves, and is required for ribosomal translocation, providing energy for the process by hydrolyzing a GTP that it carries with it to the ribosome. From my students’ experiences, the best way to learn this seems to be to study the diagrams and see the movements of the molecules, filling in the mechanistic details in your mind. This process continues until the ribosome brings the A slot in line with a stop codon.
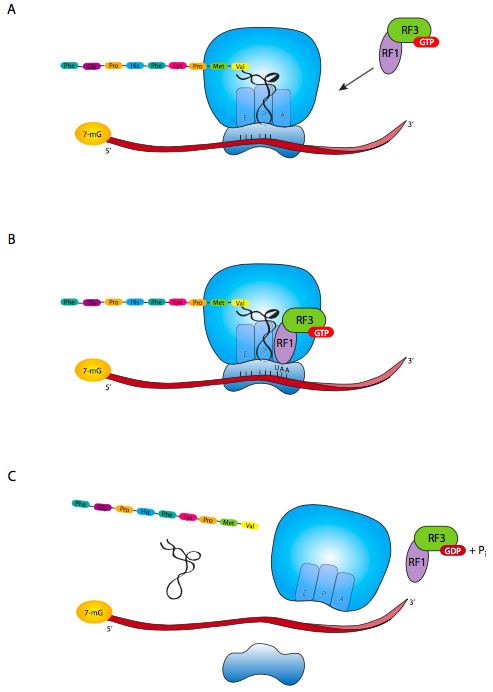
There is no tRNA with an anticodon for the stop codon. Instead, there is a set of release factors that t into the A site of the ribosome, bind to the stop codon, and activate the ribosome to cut the bond between the polypeptide chain and the last tRNA (Figure \(\PageIndex{6}\)). Depending on which stop codon is present either RF1 (recognizes UAA or UAG) or RF2 (for UAA or UGA) first enters the A slot. The RF1 or RF2 is complexed with RF3, which is involved in subsequent releasing of the RF complex from the A slot. This is necessary because once the polypeptide has been released from the ribosome, the mRNA must be released. Ribosome releasing factor (RRF) also binds in the A slot, which causes a conformational change in the ribosome releasing the previous and now empty tRNA. Finally, EF-G binds to RRF, and with an accompanying hydrolysis of GTP, causes dis- sociation of the ribosome into separate large and small subunits. Note that it is the combination of EF-G/RRF that causes dissociation; EF-G alone plays a different role in ribosome movement when it is not at the stop codon.


