5.1: Glycolysis
- Page ID
- 16114
Nearly all metabolic reactions are catalyzed by enzymes in order to keep up with the energy and material demands of the cell. In fact, the discussion of some of the metabolic processes in this chapter will almost seem to be laundry lists of enzymes. We will begin with one such list in describing the catabolism of the simple sugar, glucose, through the process of glycolysis.
Glycolysis
Whether the cell is prokaryotic or eukaryotic, one of its basic methods for generating usable energy is glycolysis. This process uses glucose, which is the most common energy source for most cells. However, glucose cannot be directly broken down to provide energy for the cell: glycolysis is a process that breaks it down in a series of reactions to create adenosine triphosphate (ATP), which is the most common energy “currency” of the cell. That is, ATP can release usable energy in a single reaction.
Glucose, being a 6-carbon sugar, has a large amount of potential energy stored in its bonds. However, since it is thermodynamically stable, it would take the investment of a lot of external energy to release the energy of glucose in one step (e.g. lighting it on fire to break it down into CO2 and H2O), and not only is it impossible for cells to generate that kind of energy at once, the cell has no mechanism to use all the energy released at one instant in time. Most of it would be wasted as excess heat. Instead, the cell uses enzymes to destabilize and break down the sugar through a series of conversions into intermediate compounds. The basic process and enzymes involved are as follows.
1. Glucose is phosphorylated by hexokinase to make Glucose-6-Phosphate. The enzyme is so named because it is a kinase (puts a phosphate group on) that acts on a hexose (six-carbon sugar). In this case, it places the phosphate on the 6-carbon of glucose. However, hexokinase can also phosphorylate other hexoses such as fructose and mannose (all in the D-conformation). There are two major reasons this is good for the cell. Since glucose concentration is higher inside the cell than outside, there is pressure for it to move back out of the cell. By converting it to G6P, it is no longer part of the glucose concentration gradient, and it has a charged phosphate group, making it nearly impossible to leak out of the membrane. The addition of the phosphate also increases the energy in the molecule, making it less thermodynamically stable, so that it can be broken down. This reaction requires the use of ATP as a phosphate donor and the energy needed to attach it. That is, energy is used in this step, not produced. Consider it an investment of energy though, since by the end of glycolysis, more ATP is produced than used.
Hexokinase requires ATP in the form of a complex (to the 2nd and 3rd phosphate groups) with a divalent cation, typically Mg2+ in vivo. ATP alone is actually a competitive inhibitor of hexokinase. The product, G6P, also functions as an inhibitor, thus providing some measure of feedback regulation. In fact, muscle cells using glycogen stores convert the glycogen directly to G6P, so hexokinase activity is very low in those cells.
2. Glucose-6-Phosphate is converted to Fructose-6-Phosphate by phosphoglucose isomerase. As the name implies, the isomerase simply rearranges the existing atoms within the G6P to make the F6P without removal or addition of any atoms.
3. Fructose-6-Phosphate is phosphorylated by phosphofructokinase (PFK) to Fructose- 1,6-bisphosphate. There is again an investment of an ATP to provide the phosphate group and the energy to attach it.
PFK is an important regulator of glycolysis. It is a tetrameric protein, and each subunit has two binding sites for ATP: one is the normal substrate site, the other is an inhibitory site such that binding of ATP lowers the enzyme’s affinity for F6P. ATP is not the only regulator of PFK activity: AMP is also a positive regulator of PFK, and can increase it up to 5-fold.
4. The Fructose-1,6-bisphosphate is cut in half by aldolase, yielding a molecule of dihy- droxyacetone phosphate and a molecule of glyceraldehyde-3-phosphate.
There are two classes of aldolases: class I are found in animals and plants, while class II are found in fungi and bacteria. Class I require no cofactors, but class II require a divalent cation (physiologically usually Fe2+ or Zn2+).
5. The G3P can participate in the next reaction, but the dihydroxyacetone phosphate, despite its similarity, cannot. So, it needs to be rearranged by triose phosphate isomerase, which converts it to another molecule of glyceraldehyde-3-phosphate.
Triose phosphate isomerase is a “perfect enzyme” that catalyzes the formation of product as fast as the enzyme and substrate can make contact in solution (i.e. rate is purely diffusion-limited).
6. Each of the two molecules of G3P generated from the glucose molecule now undergo oxidation catalyzed by glyceraldehyde-3-phosphate dehydrogenase (GAPDH) in the presence of NAD+ and inorganic phosphate (Pi). Each of these reactions produces 1,3-bisphosphoglycerate, which has a high-energy phosphate group, and NADH. NADH is a high energy electron carrier (electron comes from G3P). In eukaryotes with an
aerobic environment, this NADH will likely be used to help generate ATP through the tricarboxylic acid cycle (aka Krebs cycle or citric acid cycle). In anaerobic situations, the NADH will participate in fermentation for reasons discussed in the next section.
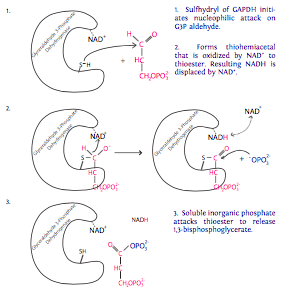
7. The phosphate group on the 1-carbon of 1,3-bisphosphoglycerate is transferred to ADP by phosphoglycerate kinase to make 3-phosphoglycerate and ATP (finally!). From the two molecules of G3P entering step 6, we get two molecules of ATP to provide energy for the cell in this step. Recalling the earlier investment of ATP (in steps 1 and 3), the reaction has only “broken even” at this point. 2 in, 2 out.
The name of the enzyme suggests that a phosphate is added to phosphoglycerate. This is not a mistake: remember that enzymes can catalyze reactions in either direction, depending on reaction conditions. Under conditions of high phosphoglycerate and ATP, phosphorylation of phosphoglycerate would occur. However, the physiological conditions are a relatively high concentration of the 1,3-bisphosphoglycerate in comparison to relatively low levels of phosphoglycerate thus driving the reaction “backwards” with respect to the naming of the enzyme.
8. The 3-phosphoglycerate is then rearranged by phosphoglycerate mutase to make 2-phosphoglycerate. This molecule has a higher free energy of hydrolysis than when the phosphate group is on the 3-carbon.
The action of phosphoglycerate mutase is not just the intramolecular phosphate group transfer that it seems to be at first glance. The enzyme must first be activated by phosphorylation, and it is the enzyme’s phosphate that is added to the 2-carbon of 3PG. The doubly-phosphorylated intermediate then transfers its 3-phosphate to the enzyme, and 2PG is released.
9. That energy is used to create ATP, as the 2-phosphoglycerate undergoes dehydration by enolase to make phosphoenolpyruvate (PEP).
PEP is made because hydrolysis of the phosphate from 2PG does not release enough energy to drive phosphorylation of ADP to ATP. PEP hydrolysis, on the other hand, releases significantly more than needed.
10. Pyruvate kinase then transfers a high energy phosphate group from PEP to ADP, producing an ATP for use by the cell, and pyruvate.
Pyruvate kinase requires not only divalent Mg2+ as with most other kinases, but also K+. The enzyme works in two steps: the ADP attacks the PEP phosphorus to make ATP and enolpyruvate. Enolpyruvate is then converted to its keto tautomer.
Keeping in mind the doubling of reactions from steps 6-10 (splitting of fructose-1,6- bisphosphate generates two G3P), the total usable energy production from glycolysis of a single molecule of glucose is 4 ATP and 2 NADH. However, the net ATP production is only 2 ATP if we remember the initial investment of two ATP in the early steps. Not really anything to write home about. Furthermore, although the NADH and pyruvate can participate in the tricarboxylic acid cycle in aerobic eukaryotic situations to generate a significant amount of ATP, in anaerobic situations, they do not produce usable energy.

Bidirectional arrows indicate enzymes used for both glycolysis and gluconeogenesis. Unidirectional arrows indicate enzymes that only function in glycolysis. *Note that reactions 6-10 are occurring in duplicate (two G3P from one glucose).
Thus anaerobic ATP production, i.e. glycolysis, is far less efficient at extracting energy from a glucose molecule than aerobic ATP production, which can generate approximately 38 ATP per glucose. On the other hand, when a lot of ATP must be generated quickly, glycolysis is the mechanism of choice, in cells such as the fast-twitch fibers of skeletal muscle. These cells actually have very few mitochondria because glycolysis can produce ATP at a much higher (up to 100 times) rate than oxidative phosphorylation. What happens to the pyruvate and NADH? In aerobically metabolizing cells, they go to the mitochondria for the TCA cycle and oxidative phosphorylation. In anaerobes, they undergo fermentation.
Note that the NADH produced by glycolysis in the cytoplasm does not directly participate in oxidative phosphorylation in the mitochondria since the inner mitochondrial membrane is impermeable to it, but it sends a “virtual equivalent” into the mitochondria via one of two pathways: the aspartate-malate shuttle combines malate-α-ketoglutarate antiports, aspartate-glutamate antiports, and metabolite interconversion by transaminase with malate dehydrogenase to oxidize NADH cytoplasmically and use the energy generated to reduce NAD+ in the mitochondrial matrix; the other pathway is a DHAP shuttle system, in which NADH is used to reduce dihydroxyacetone phosphate to glycerol-3-P using a cytoplasmic glycerol-3-phosphate dehydrogenase, and the cycling the DHAP to glycerol-3-P via a flavoprotein dehydrogenase embedded in the inner mitochondrial membrane. This flavoprotein dehydrogenase takes the electrons from glycerol-3-P to make FADH2, which can participate in the electron transport chain.
The DHAP or glycerophosphate shuttle is less efficient than the malate-aspartate shuttle, generating approximately 2 ATP vs 2.7 ATP per NADH. However, it can operate even when the concentration of cytoplasmic NADH is low, as happens in tissues/cells with a very high metabolic rate (including skeletal muscle and brain), while the malate-aspartate shuttle (prevalent in liver and heart) is sensitive to the relative concentration of NADH and NAD+.


