1.5: DNA Replication - Introduction to Prokaryotic replication
- Page ID
- 18123
1953
- The DNA model of Watson and Crick suggested how genetic information might be replicated: either strand of the duplex can be used as a template to replicate the sequence information.
- But, was the replication conservative (i.e. the original parental strands remain together after replication) or semi-conservative(one parental strand pairs with one newly synthesized strand)?
The answer for prokaryotic organisms (i.e. lack a true membrane bound nucleus and cellular organelles; e.g. bacteria) came from the 1958 experiment of Meselson and Stahl.
1958
The nitrogen in ammonium salts in culture broth is incorporated into DNA bases. The most common isotope of nitrogen is 14N. However, 15N ammonium salts (a heavier isotope) can also be obtained.
- DNA from E. coli cells grown with 15N ammonium salts will have a higher density than DNA grown in "normal" (14N) ammonium salts.
- Such DNA will migrate differently on cesium chloride (CsCl2) equilibrium density gradient centrifugation.
- The more dense DNA will migrate as a lower band (on this type of centrifugation the characteristic migration position is a function of density, and is independent of DNA length).
.png?revision=1&size=bestfit&width=495&height=327)
Figure 1.5.1: Density of 15N and 14N DNA
Meselson and Stahl reasoned that if they grew E. coli in 15N salts then switched media to 14N salts for additional rounds of replication, the mode of replication could be deduced from the density of the DNA.
.png?revision=1&size=bestfit&width=591&height=576)
Figure 1.5.2: Meselson and Stahl experiment
- After switching to the 14N media and allowing the cells to go through a round of replication a single band of intermediate density was observed (i.e. between 14N and 15N control DNA samples).
- After a second round of replication in 14N media two bands were present in approximately equimolar amounts; one was intermediate in density and the other migrated as purely 14N labeled DNA.
The results were consistent with a semi-conservative mode of replication for DNA. Evidence of semi-conservative replication of DNA has since been obtained with both plant and animal DNA.
DNA Replication in E. coli
E. coli DNA polymerase characteristics:
- Polymerase will only elongate an existing polynucleotide. It cannot initiate polynucleotide formation:
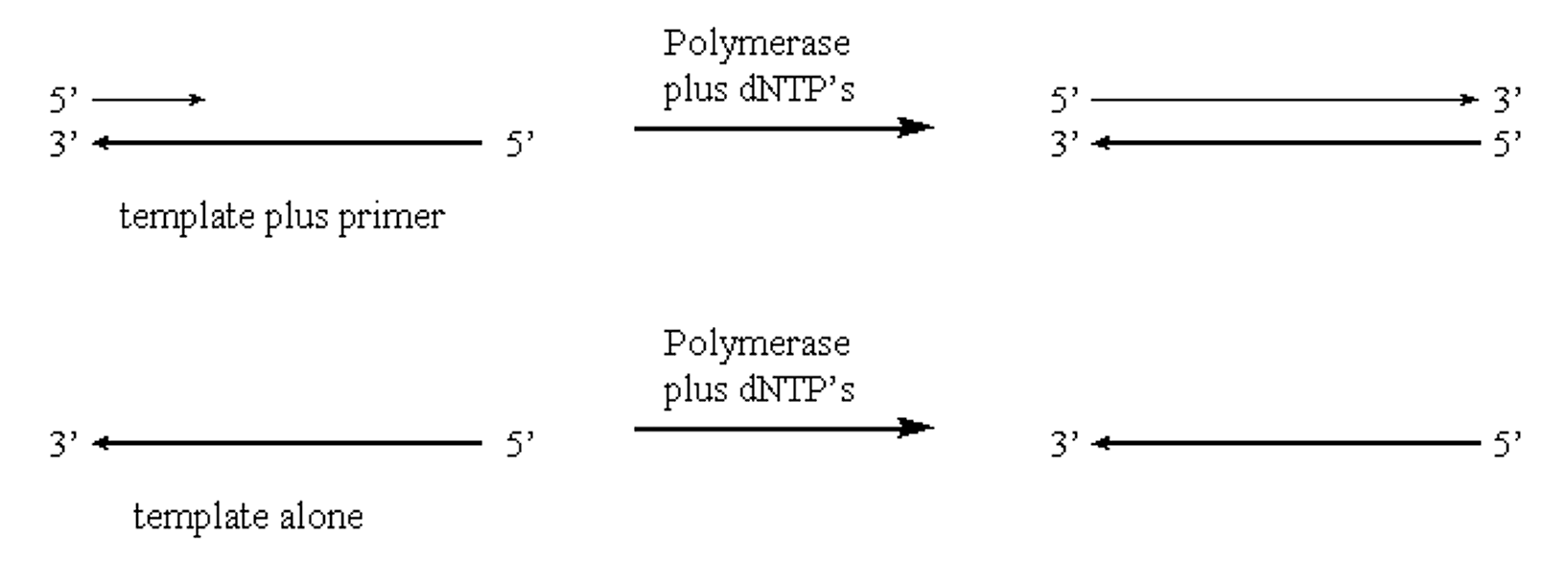.png?revision=1&size=bestfit&width=851&height=304)
Figure 1.5.3: DNA Polymerase activity
- Polymerase will catalyze polymerization of nucleotides only in one direction (5'>3') via a phosphodiester bond between a 3' hydroxyl and 5' phosphate group.
- DNA polymerase is unable to unwind duplex DNA to separate the two strands which need to be copied
E. coli genome is circular duplex DNA of approximately 4 x 106 base pairs (i.e. 4 Mb)
- The genome has a single origin of replication.
- DNA duplication in E. coli begins at a specific site in the DNA called "oriC".
OriC is a region of DNA approximately 240 nucleotides long.
- It contains repetitive 9-base pair and 13-base pair sequences (known as the '9-mer' and '13-mer' regions).
- These sequences are AT rich regions, which melt at lower temperatures than DNA containing GC pairs.
- These regions are postulated to help melt the DNA duplex in the oriC region for initiation of DNA replication.
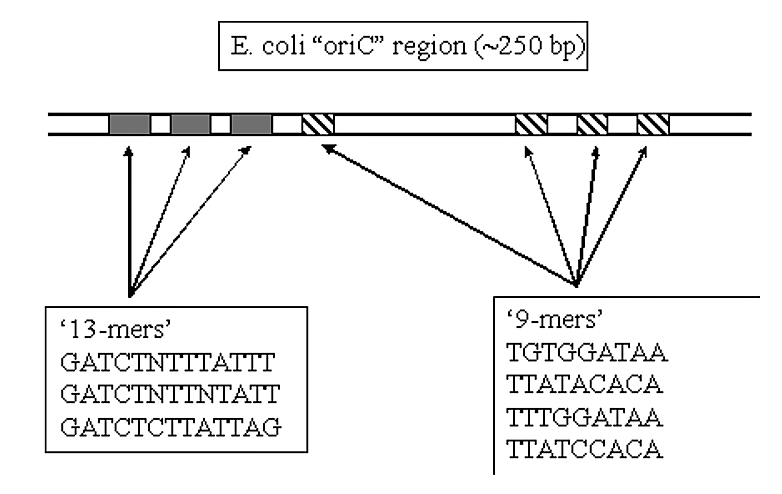.png?revision=1&size=bestfit&width=539&height=340)
Figure 1.5.4: oriC region
The dnaA gene product: (dnaA protein)
- Strains of E. coli with mutations in the dnaA gene were able to grow at 30 °C, but not at 39-42 °C.
- However, if DNA synthesis was begun at 30 °C, and then the temperature was shifted to 42 °C, DNA synthesis continued until the genome was replicated (and the cell divided), but no new initiationof DNA synthesis was possible.
Conclusion: Somehow the product of the dnaA gene (i.e. the dnaA protein) is required for initiation of DNA synthesis.
Studies of purified dnaA protein:
- dnaA protein binds to the '9-mer' region in oriC and forming a multimeric complex with 10-20 protein subunits (i.e. at a single oriC region there will be bound 10-20 dnaA protein molecules).
- Binding requires ATP.
- Further addition of ATP was observed to result in a melting and opening up of the DNA duplex in the oriC region. This was determined by addition of S1 nuclease (like mung bean, but will also cut DNA at the site of an internal nick), which resulted in cleavage of DNA at the site of oriC.
The dnaB gene product: (dnaB protein)
The protein encoded by the dnaB gene appears to be essential for DNA replication. The dnaB protein has been identified as ahelicase. A helicase moves along a DNA strand opening up the duplex to melt and separate the DNA strands.
- dnaB protein binds to the single stranded DNA in the general region of the oriC DNA segment.
- Binding requires ATP as well as the dnaC gene product (the dnaC protein).
- After helicase/dnaC binds to the DNA, the dnaC protein is released.
- Two helicases bind at the oriC region, one helicase on each strand of the DNA.
This stage represents the prepriming complex:
.png?revision=1&size=bestfit&width=601&height=425)
Figure 1.5.5: Prepriming complex
Separated strands in the oriC region are prevented from reannealing by the binding of single-stranded binding protein (ssb protein).
The dnaG gene protein:
The dnaG gene protein is called primase.
- Primase catalyzes synthesis of short RNA molecules that function as primers for DNA synthesis by E. coli DNA polymerase III(pol III).
- Primase binds to dnaB protein at oriC and forms a primosome.
- The primase within the primosome complex provides RNA primers for synthesis of both strands of duplex DNA.
- Primase lays down tracks of pppAC(N)7-10 (RNA).
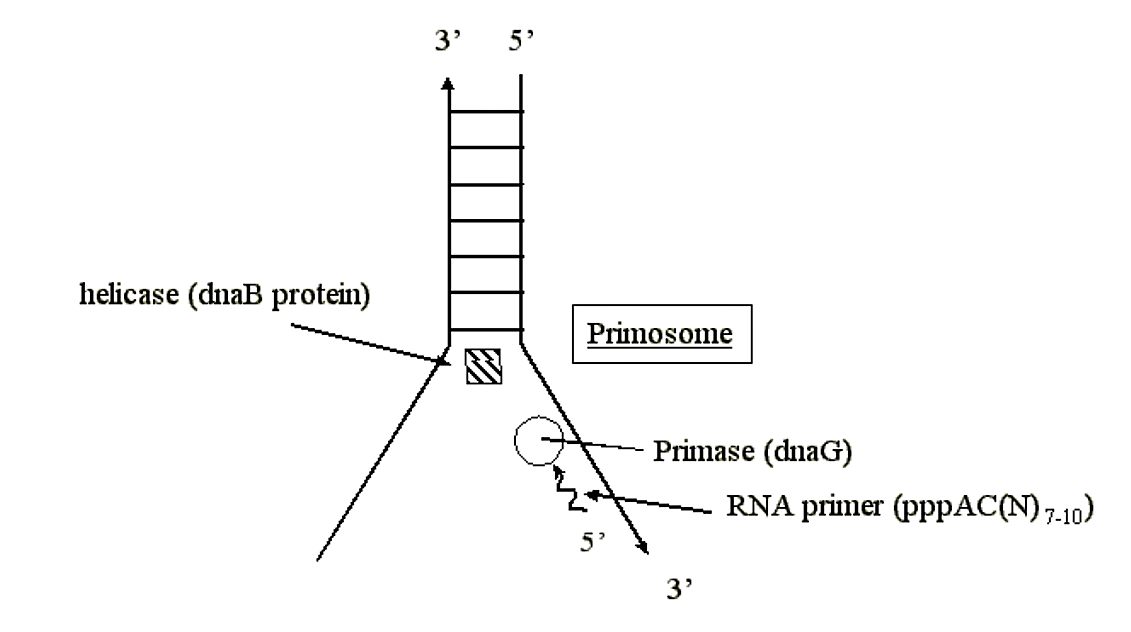.png?revision=1&size=bestfit&width=597&height=334)
Figure 1.5.6: Primase activity
- After synthesis of the 9-12 mer RNA primer, DNA Pol III holoenzyme enters the replication fork and is able to utilize the RNA as a primer for DNA synthesis.
- As the replication fork opens up, the leading strand synthesis can continue, but a gap develops in the lagging strand:
.png?revision=1&size=bestfit&width=754&height=417)
Figure 1.5.7: Lagging Strand
DNA Pol III is a large multicomplex enzyme (holoenzyme) which is somewhat dimeric in nature (there are two polymerase active sites). The two active polymerase sites in Pol III could actually function to synthesize both nacent strands at the fork. However, the synthesis of the lagging template strand would be in the opposited direction to the movement of the Pol III complex:
.png?revision=1&size=bestfit&width=743&height=430)
Figure 1.5.8: Pol III movement
Primase can bind to the Pol III complex, but the arrangement of the DNA strand as it passes through the Pol III/primase complex is quite unique. It forms a loop structure such that primase and the Pol III active site can accomplish discontinuous synthesis of the lagging template strand even though the general direction of the Pol III complex is opposite to the require direction of DNA synthesis:
.png?revision=1&size=bestfit&width=689&height=451)
Figure 1.5.9: Loop for Pol III activity
After primase makes another primer on the lagging template, the adjacent Pol III active site can extend the primer (incorporating dNTP's) by utilizing the same loop structure and feeding the template through in the direction shown.
.png?revision=1&size=bestfit&width=699&height=386)
Figure 1.5.10: Synthesis of lagging strand
The lagging strand loop cannot be fed through the Pol III complex forever, and after a nascent DNA strand is synthesized the loop is released and a new one is formed using the opened template DNA further up the fork:
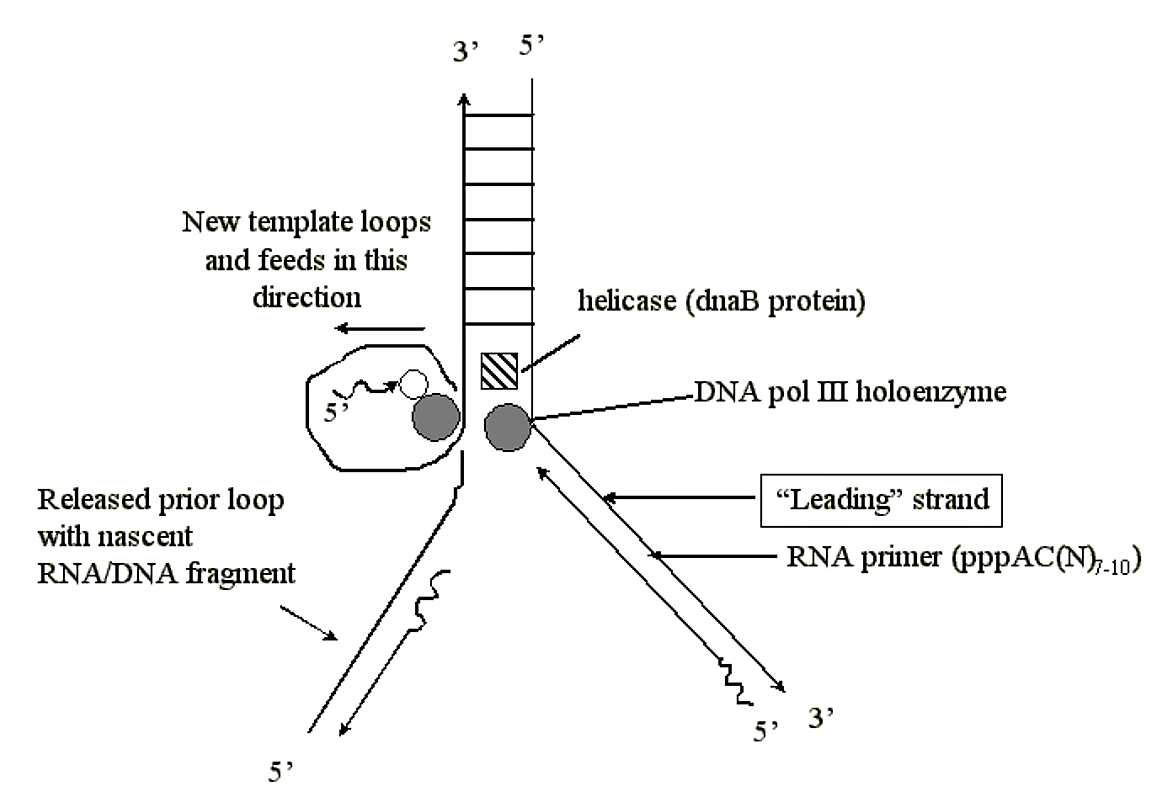.png?revision=1&size=bestfit&width=589&height=406)
Figure 1.5.11: Continuation of lagging strand synthesis
As synthesis continues:
- there will be a single continuous DNA strand on the leading strand
- there will be a series of short fragments on the lagging strand, containing both RNA and DNA, called Okazaki fragments:
.png?revision=1&size=bestfit&width=475&height=412)
Figure 1.5.12: Okazaki Fragments
How are these RNA/DNA fragments converted into one long continuous DNA strand? The RNA could be removed by a polymerase which has 5'->3' exonuclease activity, however, Pol III lacks this activity.
- DNA Pol I does have 5'->3' exonuclease activity
- it can extend the DNA synthesis via nick-translation.
- The nick-translation activity restults in degradation of the RNA primers.
- The end result is a series of "nicks" in the lagging strand, now 100% DNA:
.png?revision=1&size=bestfit&width=377&height=479)
Figure 1.5.13: Nicks in lagging strand
- DNA Pol I leaves and DNA ligase then joins these discontinuous DNA fragments to form a continuous DNA duplex on the lagging strand.
Summary of steps in E. coli DNA Synthesis
- dnaA protein melts duplex in oriC region.
- dnaB (helicase), along with dnaC and ATP binds to replication fork (dnaC protein exits).1 (Pre-priming complex)
- Single strand binding protein (ssb protein) binds to separated strands of DNA and prevents reannealing.
- Primase complexes with helicase, creates RNA primers (pppAC(N)7-10) on the strands of the open duplex2 (Primase+helicase constitute the Primosome).
- After making the RNA primers, DNA pol III holoenzyme comes in and extends the RNA primer (laying down dNTP's) on the leading strand.
- As the replication fork opens up (via helicase + ATP action) leading strand synthesis is an uninterrupted process, the lagging strand experiences a gap.
- The gap region of the lagging strand can wind around one active site unit of the Pol III complex, and bound Primase initiates an RNA primer in the gap region3.
- On the lagging strand, Pol III extends the RNA primer with dNTP's as the lagging template strand is looped through the Pol III complex
- After synthesis of a nascent fragment the lagging strand loop is released and the single strand region further up near the replication fork is subsequently looped through the Pol III complex.
- Steps 7-9 are repeated.
- Meanwhile, Pol I removes the RNA primer regions of the Okazaki fragments via 5' to 3' exonuclease activity ( nick translation
- Pol I exits and ligase joints the DNA fragments (on lagging strand).
Notes From Above
- Polymerases are unable to open up duplex DNA, thus the requirement for helicase
- Polymerases cannot replicate a DNA template in the absence of a primer (either DNA or RNA).
- Polymerases extend a polynucleotide in the 5' to 3' direction only. Gaps at the 5' end must be filled by "upstream" discontinuous synthesis.
|
|
DNA Pol I |
DNA Pol II |
DNA Pol III |
|---|---|---|---|
|
5'->3' Polymerase activity |
* |
* |
* |
|
3'->5' Exonuclease activity(proof reading) |
* |
* |
* |
|
5'->3' Exonuclease activity(nick translation) |
* |
||
|
Synthesis from: |
|||
|
Duplex DNA |
|||
|
Primed single strand |
* |
||
|
Primed single strand plus ssb protein |
* |
* |
|
|
Chain elongation rate(in vitro) bp/min |
600 |
? |
30,000 |
|
Molecules/cell |
400 |
? |
10-20 |
|
Mutation Lethal? |
* |
* |
Pol Functions:
- Pol I: gap filling during DNA synthesis and repair, removal of RNA primers
- Pol II: involved in DNA synthesis of damaged templates
- Pol III: functional polymerase at the replication fork
Pol III Structure and function
- A "holoenzyme" complex of 10 different polypeptides
- resultant molecular weight is greater than 600 KDa (i.e. it is a large complex).
- It is structurally an asymmetric dimer - it contains two copies of most of the polypeptides which comprise it, including two catalytic sites for nucleotide addition (i.e. polymerization).
The various protein subunits have a variety of functions:
- Subunits for polymerase activity: a, e, subunits
- Subunits to dimerize the core polymerase (t)
- Subunits to increase processivity (i.e. to increase the ability to synthesize long stretches w/o releasing from the DNA template): b subunits
- Subunits to bind b to DNA-primer substrate: (g, d, d', c, )
Termination of DNA replication
- Specific termination sites of DNA replication exist in E. coli.
- Termination involves the binding of the tus gene product (tus protein).
- This protein may act to prevent helicase from unwinding DNA (will therefore halt pol III and pol I action).
- DNA replication produces two interlocking rings which must be separated.
- This is accomplished via the enzyme topoisomerase.
ColE1 Plasmid
E. coli can contain a small extrachromosomal element called the ColE1 plasmid. This plasmid has the following general features:
- 6.4 Kb circular duplex DNA
- Autonomously replicating
- 10-15 copies per cell (i.e. per E. coli chromosome)
Although it is autonomously replicating, it does not contain an oriC type of sequence for initiation of replication, and does not undergo the same steps in replication.
- The plasmid produces (among other things) two RNA oligonucleotides (RNA I and RNA II)
- RNA II has complementarity to the ColE1 origin of replication, which contains an AT rich sequence
- The bound RNA II molecule can serve as a primer for polIII
- Since RNA II binds on one strand only, the replication of the ColE1 plasmid is unidirectional
- RNA I has complementarity with RNAII, and such a duplex RNA cannot serve as a replication primer, thus control of the plasmid copy number is achieved by the interaction between RNA I and RNA II.


