28.16: Signaling in Plants
- Page ID
- 15120
Introduction
Plants are obviously comprised of cells. Hence they must engage in cell signaling within and between cells. It is beyond the scope of this book to give a detailed description of cell signaling in plants. Instead, we focus on 5 key classic plant hormones, auxins, cytokinins, ethylene, gibberellins, and abscisic acid, which are produced by leaves, flowers, shoots, roots, or fruit, and see how they initiate signaling in plants. Finally, we would be remiss if we didn't include the profound signaling in plants initiated by light. Most of this section comes directly from a series of sources, with modifications and additions (mostly molecule models).
Auxins (3-indolebutyric acids derivatives) are regulators of growth and development and are found in actively growing parts of the plant (root, shoot, leaves) but mostly in the cell stem. Auxins facilitate the bending of plants toward the light, for example. They work in conjunction with other hormones like cytokinins. When auxins are higher than cytokinins, roots will form, while the opposite produces shoots. Auxins facilitate the elongation of cells, while cytokinins promote cell division and growth as well as wound repair. Gibberellins also are plant growth regulators and facilitate cell elongation. They also help in germination, elongation of the stem, fruit ripening, and flowering. Abscisic acid affects seed development and maturation and helps plants tolerate environmental or biotic stresses. It also inhibits growth and metabolism. Ethylene affects fruit ripening, organ abscission, and growth by restricting cell elongation.
We will focus on the hormones, their protein receptors, and how the hormone:receptor complex initiates some key events in the cell.
Auxin
Much of this section derives from Kou et al. Appl. Sci. 2022, 12(3), 1360; https://doi.org/10.3390/app12031360. Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Auxins, the first plant hormones discovered, regulate plant growth and development. The most common auxin is 3-indole acetic acid. Figure \(\PageIndex{1}\) shows the structures of naturally occurring auxins.
Figure \(\PageIndex{2}\) shows an interactive iCn3D model of auxin bound to its receptor, TIR1 ubiquitin ligase (2P1Q)
Figure \(\PageIndex{2}\): Auxin bound to its receptor TIR1 ubiquitin ligase (2P1Q) (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...6xSkEZ3aJA4gT6
Auxin (IAA) is shown in spacefill CPK colors along with an unexpected binding cofactor, inositol hexakisphosphate (IHP), shown in spacefill CPK colors. The peptide shown in light brown sticks is part of the protein Auxin-responsive protein IAA7, a member of a class of proteins called AUX/IAAs. These are short-lived transcriptional factors that function as repressors of early auxin response genes at low auxin concentrations.
The magenta subunit, TIR1 (transport inhibitor response 1), is part of the larger TIR1 complex, the SCF(TIR1) E3 ubiquitin ligase, of which only TIR1 is shown. Its mere name suggests that it is involved in the ubiquitinylation of a key protein involved in auxin activity, which will be targeted for proteolysis. That protein is the repressor protein IAA7 (an AUX/IAA protein).
Auxin binds in a hydrophobic pocket, which accounts for the binding of the other largely hydrophobic auxins shown in Figure \(\PageIndex{1}\). Note however that Arg 401 that forms a salt bridge (ion-ion interaction) with the carboxylate of 3-indole acetic acid
Figure \(\PageIndex{3}\) shows an interactive iCn3D model of auxin bound in the hydrophobic pocket of its receptor, TIR1 ubiquitin ligase (2P1Q).
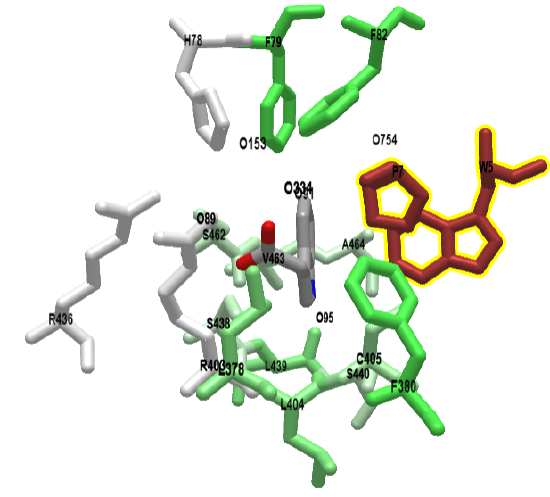
The green color represents nonpolar side chains. The brown side chains, Trp and Pro, are from the auxin-responsive protein IAA7 peptide. To reiterate, the protein IAA7 is a member of a class of proteins called AUX/IAAs which repress auxin activity. The IAA7 peptide packs over the auxin in the binding pocket.
Now we can see how auxin function to regulate gene transcription. First, we must introduce another protein family, the auxin response factors (ARFs). These are transcription factors that bind to a key DNA sequence, the auxin response element (AuxRE) in promoter sequences of auxin-activated genes. Once bound they can either activate or repress transcription from target genes. Auxin binds its receptor TIR1 enabling the binding of an AUX/IAAs (like IAA7) repressor and the binding of the complex to ARF. The TIR1 ubiquitin ligase activity of the complex ubiquitinylates the bound AUX/IAAs (like IAA7) repressor, targeting it for degradation, freeing the ARF to become active in the regulation of gene transcription in the nucleus.
ARFs are structurally similar, with most members containing three regions: DBD (DNA-binding domain), MR (middle region), and PB1 (Phox and Bem 1). Figure \(\PageIndex{4}\) shows a model of how auxin affects ARF transcriptional activity.
At low concentrations of auxin, the AUX/IAA repressor binds to the ARF transcription factor through their PB1 domains. The PB1 (Phox and Bem1) domain is about 80 amino acids in length. It acts as a protein binding module allowing heterodimerization or homo-oligomerization with proteins have also contained the PB1 domain.
The dimer of AUX/IAA and ARF recruits the co-repressor TPL (TOPLESS) to inhibit the ARF activity and the expression of auxin-responsive genes. When the concentration of auxin is increased, Aux/IAA binds to the SCF TIR1/AFB complex and is ubiquitinated and then degraded by 26S protease. The ARF transcription factors are released to activate the transcription of downstream genes. DBD, DNA-binding domain; MR, middle region; PB1, Phox, and Bem 1.
- At low concentrations of auxin, the AUX/IAA repressor binds to the ARF transcription factor and forms a dimer that recruits the co-repressor TPL (TOPLESS) to inhibit the ARF activity and the expression of auxin-responsive genes;
- When the concentration of auxin is increased, Aux/IAA binds to the auxin:SCF TIR1/AFB complex (remember that the auxin receptor is the TIR1 component of the complex) and is ubiquitinated by the TIR1, which is also a ubiquitin ligase;
- The ubiquitinated AUX/IAA protein is degraded by proteolysis by the 26S protease, allowing the ARF to become an active transcription factors
It appears that the MR domain of ARF determines whether it activates or inhibits transcription. If it is rich in proline, serine, and threonine, it acts as an inhibitor. If it is enriched in glutamine and leucine it acts as an activator. Some reports show that Aux/IAA and ARFs can form not only dimers but also larger complexes (oligomers), noting that oligomerization of Aux/IAA proteins may be essential for the inhibition of ARF proteins and only sufficient amounts of Aux/IAA proteins can exert the inhibitory effect of ARF proteins.
Figure \(\PageIndex{5}\) shows an interactive iCn3D model of the DNA binding domain of arabidopsis thaliana auxin response factor 1 (ARF1) in complex with auxin response element-like sequence ER7 (4LDX)
__protomor-like_sequence_ER7_(4LX).png?revision=1&size=bestfit&width=383&height=347)
The ARF protein must translocate to the nucleus to regulate gene transcription. ARF7 and ARF19 have been shown to form micron-sized aggregates in the cytoplasm. These have low responses to auxin. Aggregation occurs through PB1 domain interactions between ARFs as well as through intrinsically disordered regions. Mutation of a single lysine in the PB1 prevents aggregation and leads to morphological changes in the plant. This shows the importance of regulating not only transcription but also the translocation of proteins to the nucleus.
Figure \(\PageIndex{6}\) reviews the activation of ARFs and some of the genes affected by ARF.
Auxin promotes the formation of the TIR1/AFB Auxin/Indole-3-acetic acid inducible (Aux/IAA) co-receptor to promote the ubiquitylation and subsequent degradation of the Aux/IAA repressor. Aux/IAA degradation relieves repression of auxin response factor (ARF) transcription factors, allowing for auxin-responsive gene expression. One of the transcript families upregulated by auxin is the SAUR family. The small auxin up RNA (SAUR) proteins encoded by these transcripts have been suggested to play roles in multiple processes, one of which is interaction with and inhibition of members of the PP2C.D family of phosphatases, which act to regulate H+-ATPase activity. Further, indole-3-butyric acid response 5 (IBR5) and mitogen-activated protein kinase 12 (MPK12) have been implicated in regulating auxin-responsive gene transcription; this regulation is not through destabilization of the Aux/IAA repressors, suggesting a yet-to-be-discovered mechanism of regulating auxin-responsive gene expression. F
Cytokinins (CKs) and Ethylene (ET)
Much of this material derives from Bidon et al. Cells 2020, 9, 2526; doi:10.3390/cells9112526 . Creative Commons Attribution (CC BY) license. (http://creativecommons.org/licenses/by/4.0/).
Cytokinins (CKs) and ethylene (ET) are among the most ancient organic chemicals on Earth. The structure of a representative cytokinin (kinetin) and ethylene are shown in Figure \(\PageIndex{7}\).
A wide range of organisms including plants, algae, fungi, amoebae, and bacteria use these substances as signaling molecules to regulate cellular processes. Because of their ancestral origin and ubiquitous occurrence, CKs and ET are also considered to be ideal molecules for inter-kingdom communication. Their signal transduction pathways were first determined in plants and are related to the two-component systems of bacteria (which we explored in a previous section), using histidine kinases as primary sensors.
CKs share a common structure of N6-substituted adenine (see Figure \(\PageIndex{7}\)), with biological activities defined by the N6-substituents (isoprenoids or aromatic groups). They were originally described as the major hormones regulating cell division but are also implicated in the control of morphogenesis and embryogenesis and inhibition of senescence. Conversely, ET is a simple gas, often referred to as the senescence hormone in plants, acting to stimulate the senescence of leaves and petals as well as the ripening of fruits. Both CK and ET are also well known to orchestrate plant responses to many types of biotic and abiotic stresses.
Signaling pathways in plants are related to the two-component systems typically described in prokaryotes. CKs and ET are perceived by two types of membrane-bound histidine kinase receptors, CRE1 and ETR1 as shown in Figure \(\PageIndex{8}\).
Panel A shows the cytokinin signaling pathway. CKs in Arabidopsis primarily are recognized by dimerized receptors such as the CRE1 receptor via the cyclase/histidine kinase-associated sensing extracellular (CHASE) domain. CRE1 then auto-phosphorylates (histidine kinase (HK) activity) and immediately transfers its phosphate group to the conserved histidine of a protein belonging to the histidine-containing phosphotransfer (HPt) family. This small protein then acts as a cytoplasm-to-nucleus shuttle and in turn phosphorylates a type B response regulator, which, when activated, positively regulates the transcription of response genes to the CK signal.
Panel (B) shows the ET signaling pathway. Ethylene molecules are detected by ethylene receptors (labeled ETR1) with ethylene binding to the three transmembrane helices (shown in sky blue). The binding of ET to the dimerized ETR1 receptor downregulates its activity. In the absence of ET, ETR1 activates the serine/threonine kinase CTR1. The CTR1 protein then phosphorylates the EIN2 protein located in the ER membrane, leading to the proteolysis of EIN2. In the presence of ET, ETR1 activity is reduced, leading to less CTR1 activity; this leads to lower phosphorylation and accumulation of EIN2 protein and subsequent activation of the EIN3 and related transcription factors. EIN3 then positively regulates the transcription of ET signal response genes.
Panel (C) shows the domain structure of the Arabidopsis ET (ETR1) and CK (CRE1) receptors.
Mechanistically, the two pathways use fundamentally different families of downstream modules.
It is now known that bacteria also use CK and ET signaling, as described in Figure \(\PageIndex{9}\).
Let's look in more detail at CRE1, cytokinin response 1, the main cytokinin receptor in plants. Different computational programs often show different domain structures. Figure \(\PageIndex{10}\) shows the domain structure determined by Pfam.

Uniprot describes this domain structure, color-coded as in Figure \(\PageIndex{11}\)
- 131-149: transmembrane
- 200-382: Green Chase (Cyclases/Histidine kinases Associated Sensory Extracellular)
- 420-443 transmembrane
- 472-537: Red His Kinase A Phosphoaccepter domain
- 584-760 Blue HK kinase, DNAgyrase, HSP-like ATPase
- 786-920: Yellow Reg REsp 1
- 946-1071: Yellow Reg Regulator receiver domain
Figure \(\PageIndex{11}\) shows an interactive iCn3D model of Histidine kinase 4 - Cytokinin receptor 1 (CRE) from Arabidopsis thaliana (AlphaFold model - Q9C5U0). The coloring matches the Pfam domains shown in Figure \(\PageIndex{10}\).
_from%25C2%25A0Arabidopsis_thaliana_-_PfamDomain_organization_(Q9C5U0).png?revision=1&size=bestfit&width=373&height=175)
Figure \(\PageIndex{12}\) shows an interactive iCn3D model of Histidine kinase 4 - Cytokinin receptor 1 (CRE) from Arabidopsis thaliana (AlphaFold model) - Domain organization (Q9C5U0) that clearly shows the extracellular and intracellular domains.
_from%25C2%25A0Arabidopsis_thaliana_-_Domain_organization_(Q9C5U0).png?revision=1&size=bestfit&width=100&height=173)
The N-terminal methionine is in cyan spacefill and the C-terminal Ser is in spacefill. Two transmembrane helices are shown in dark gray spacefill (125-145) and light gray spacefill (430-450). These connect the extracellular domain (cyan, 146-429) and the two cytoplasmic domains (magenta 1-124, which is mostly disordered in the model, and 451-1080). This model does not reflect the relative disposition of the protein in the actual structure, but clearly shows the extracellular and cytoplasmic domains. The extracellular domain (cyan, 146-429) is the CHASE domain.
Here are the step involved in cytokinin signaling through its receptor (shown in Figure \(\PageIndex{8}\) :
- the cytokinin binds to the CHASE domain
- the receptor autophosphorylates a His in the HK domain
- a phosphotransfer from the pHis to an Asp in the Yellow Reg (Regulator) Receiver domain
- a phosphotransfer from pAsp to the His in the histidine-containing phosphotransfer protein (HPt)
- a final transfer from pHIs to an Asp in a response regulator (RR)
The MAPK cascade is activated in the cytokinin signaling pathway. Phosphorylated pRR can also regulate target gene transcription. Type-A RRs are negative regulators of cytokinin signaling. It also acts with phytochromes (discussed at the end of this section) to regulate red light signaling. Cytokinin receptors can bind synthetical chemicals that act as defoliants and herbicides.
Intermolecular interactions in the cytokinin signaling pathway leading to transcriptional effects are illustrated in Figure \(\PageIndex{13}\).
Gibberellin
Much of this material derives from Hedden, P., Sponsel, V. A Century of Gibberellin Research. J Plant Growth Regul 34, 740–760 (2015). https://doi.org/10.1007/s00344-015-9546-1. https://doi.org/10.1007/s00344-015-9546-1. Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/)
Gibberellin controls growth and development pathways in plants and fungi. They act in plants by removing growth limitation by promoting the degradation of the growth-inhibiting DELLA proteins which contain the Asp-Glu-Leu-Leu-Ala (DELLA) motif. The name gibberellin derives from the fungus Gibberella fujikuroi. There are many types of gibberellins, which are all diterpenoids. The structures of the main bioactive GAs in plants, GA1 and GA4, are shown below in Figure \(\PageIndex{14}\).
Gibberellin initiates signaling by binding to the nuclear gibberellin receptor. One such receptor is the Gibberellin Insensitive Dwarf1 (GID1). When bound, it leads to the proteolysis of another protein bound to it called a DELLA protein (an example is GAI), a transcriptional regulator that inhibits growth. The control of transcriptional activity by gibberellins is hence reminiscent of that of auxins.
Figure \(\PageIndex{15}\) shows an interactive iCn3D model of the gibberellin(GA3)- active gibberellin receptor GID1L1 bound to the DELLA domain of GAI (2ZSH).
_DELLA_recognition__gibberellin_receptor%25C2%25A0GID1L1_(2ZSH)v3.png?revision=1&size=bestfit&width=246&height=231)
The gibberellin receptor is gray and associated with the inner leaflet (blue) of the membrane. The plant hormone gibberellin A3 is bound to the receptor. Amino acid side chains in the receptor involved in the interactions with gibberellin A3 are shown in sticks, colored CPK. The DELLA protein GAI RAG protein is shown in cyan. The 3 amino acid motifs within it (DELLA, cyan spacefill; and VHYNP and LExLE, both magenta spacefill) are also shown. As with the auxin receptor, GID11A binds gibberellin in a deep pocket, which is covered by an N-terminal helix of the receptor. That helix recognizes and binds to the DELLA sequence in the DELLA transcription regulator protein.
Figure \(\PageIndex{16}\) shows the effects of a mutation that leads to deficiencies in gibberellin 1-3 (right-hand side).

The mechanism by which GAs promote growth is shown in Figure \(\PageIndex{17}\).
Binding of bioactive GA results in a conformational change in the GID1 receptor that promotes interaction with DELLA proteins. Recruitment of an F-box protein initiates ubiquitination of DELLA by an SCF E3 ubiquitin ligase targeting the DELLA for proteasomal degradation. Loss of DELLA relieves growth repression and suppresses other DELLA-mediated responses
Abscisic acid
Much of this material derives from Hewage et al. (2020). Advanced Science. https://doi.org/10.1002/advs.202001265. This is an open-access article under the terms of the Creative Commons Attribution License.
The phytohormone abscisic acid (ABA) is the best-known stress signaling molecule in plants As such it will be key as plants struggle to adapt to climate change. Its structure is shown in Figure \(\PageIndex{18}\).
ABA protects land plants from biotic and abiotic stresses. ABA receptors proteins (PYLs) contain a conserved pyrabactin resistance/pyrabactin resistance-like/regulatory domain (PYR/PYL/RCAR) that binds ABA and triggers a cascade of signaling events.
ABA has significant roles throughout a plant's life cycle. From the single-celled zygotic stage to the mature multicellular plant, plant developmental stages involve ABA. ABA allows germination only under optimum conditions and inhibits growth under stress conditions. The adult plant as well as the seedling experience biotic and abiotic stressors that vary in severity and persistence. ABA allows that plant to survive by inducing both short-term and long-term stress responses, including rapid and reversible stomatal closure, long-term growth inhibition, dormancy, senescence, and abscission. ABA is therefore both a developmental and a stress-signaling molecule with diverse roles, as shown in Figure \(\PageIndex{19}\).
ABA signaling in drought
Let's look at a specific example of ABA signaling in the presence of drought, stress that will expand as the world's climate changes due to the combustion of fossil fuels.
Insufficient levels of soil water can result in an imbalance of water between the cells and the outer environment. A resulting change in the cellular electrolyte content affects metabolism, resulting in an osmotic imbalance or stress. The osmotic stress thus leads to the accumulation of ABA in cells and triggers ABA signaling. The cellular pool of ABA is dramatically increased during drought. Biosynthesis, catabolism, conjugation, and transportation of ABA are coordinated to increase ABA levels. ABA rapidly regulates plant water levels by controlling stomata. Stomata, which are microscopic pores controlled by two highly differentiated epidermal cells (guard cells), have the primary role in regulating gas exchange between the air and plant. Open stomata allow CO2 to diffuse into the leaf mesophyll and reach the sites of photosynthesis. They also allow water vapor to exit from the plant interior to the atmosphere.
By allowing transpirational water loss, stomata allow the cooling of the plant and the managing of the interior water levels. Thus, stomata are essential regulators that connected the plant interior to the outside environment. Increases in osmotic pressure in guard cells lead to water uptake and then to cell expansion; as the cells expand, the pore opens because of differential thickenings of guard cell walls. Stomatal movements are regulated by numerous environmental signals such as light, plant growth regulators, pathogens, drought, cold, and nutrient status. Stomatal movement is the quickest response to ABA signaling. Therefore, the core ABA signaling is essential for guard cell function. The involvement of ABA signaling events in guard cell function is summarized in Figure \(\PageIndex{20}\).

In the absence of ABA, (left) ABA receptors (PYLs) are in ligand-free form. H+ATPase action pumps H+ ions outside of the plasma membrane. The SnRK2 protein kinases and the S-type anion channel SLAC1 are kept dephosphorylated by PP2Cs. The dephosphorylation state of SLAC1 prevents the nonspecific activation of S-type anion channels.
In the presence of ABA (right) PYLs bind to and inhibit PP2Cs. ABA inhibits H+ATPase activity, blocking the H+ pumping outside. The Ca2+-independent protein kinases (SnRK2s) are released from PP2C inhibition and activated by auto-phosphorylation. Ca2+-permeable cation (ICa) channels are released from PP2C-mediated inhibition, causing increases of ABA-responsive Ca2+ in cytosol leading to activate CPKs. The activated SnRK2s and CPKs phosphorylate SLAC1 The SnRK2.6/OST1 protein kinase phosphorylates and activates the R-type anion channel ALMT12/QUAC1. The K+ ions are effluxed via voltage-dependent outward K+ (K+out) channel GORK, causing a guard cell turgor decrease leading to stomatal closure. PYLs: ABA receptors; ABA: abscisic acid; PP2C: protein phosphatase 2C proteins; OST1: open stomata 1/SnRK 2.6 protein kinase; Ca/CPK: Ca2+/calcium dependent protein kinases; ICa2+: plasma membrane nonselective cation channel permeable to Ca2+ SLAC1: slow anion channel-associated 1 (SLAC1); QUAC: aluminum-activated malate transporter 12/quickly activating anion channel 1 (ALMT12/QUAC1); GORK: guard cell outward rectifying K+ channel (GORK); KAT1: K+ activated 1 potassium ion channel; A−: anions; K+: potassium ions.
The ABA receptor core complex
There are three main phases of ABA signaling: ABA synthesis/metabolism, long-distance transport, and ABA binding to its receptor. Downstream signaling ensues. through transcriptional activators/repressors and plasma membrane-located channel proteins. The main components of the core ABA signaling pathway are shown in Figure \(\PageIndex{21}\).

In the absence of ABA (A above), SnRK2 kinases are dephosphorylated by protein phosphatase 2C (PP2Cs). In the presence of ABA (B above) PP2Cs are inhibited by the complexes PYLs-ABA. Thus, the SnRK2 kinases are released and make a cascade of downstream transcription factors, NADPH transporters, and ion channels phosphorylate the transcription factors that induce ABA-responsive gene transcription, and ion channels act on the guard cells to bring about transpirational control.
ABA receptors (PYLs) bind ABA, PP2C, and protein kinases. The ABA:PYL complex binds PP2Cs, leading to conformational changes in the active sites of PP2Cs that inhibits the phosphatase. This in turn leads to the release of downstream protein kinases (SnRK2s) from PP2C-mediated inhibition. The SnRK2s undergo autophosphorylation to activate a series of ion channels, NADPH oxidases, and transcription factors via phosphorylation. This activates both short-term and long-term ABA responses such as stomatal closure and upregulation of ABA-dependent gene expression. MAPKKKs (MAPK3s) also activate SnRK2.6 by phosphorylating a specific site during salinity stress.
ABA binding regulates a double-negative regulatory system, in which the ABA receptor (PYLs) act as ABA receptors, PP2Cs as negative-regulatory coreceptors, and SnRK2s as negative regulators. In addition to the regulation by SnRK2 and PP2Cs, several post-translational modifications also regulate ABA signaling. Phosphorylation, dephosphorylation, ubiquitination, farnesylation, and sumoylation have been found to modulate ABA signaling by targeting core components (PYLs or PP2Cs) or other interacting proteins downstream.
ABA Receptors (PYLs)
PYLs are soluble proteins, and among the 14 PYLs in Arabidopsis, 13 functions as ABA receptors. All PYLs are known to share a dominant helix-grip structure. This characteristic motif consists of a seven-stranded antiparallel β-pleated sheet, which is flanked by two α helices. The β-pleated sheets enfold a long carboxy-terminal α-helix of PYLs. The apo-PYLs contain a sufficiently large hydrophobic pocket of 543˚A between the C-terminal helix and β sheet. The size of this pocket is estimated to be 480˚A in the ABA-bound state. The 23 pocket residues are highly conserved and are more hydrophobic than the other parts of PYLs. The interactions of ABA and PYL2 are shown in Figure \(\PageIndex{22}\).
Figure \(\PageIndex{22}\): The binding mode of ABA and PYL2 (PDBID: 3KDI) in A) 3D and B) 2D (redrawn).[211] In the 3D structure, the cartoon of PYL2 is colored in white (A). The important residues and ABA are shown in sticks with blue and yellow colors respectively. The H-bonds are marked with red dotted lines.
Figure \(\PageIndex{23}\) shows an interactive iCn3D model of Abscisic acid bound to the Abscisic Acid Receptor (PYL2) (pdbid: 3KDI)
%25C2%25A0_(pdbid_3KDI).png?revision=1&size=bestfit&width=312&height=236)
In the absence of ABA, the apo-PYL2 has a pocket surrounded by four surface loops. When ABA binds, one loop (CL2) closes onto the pocket, forming a PP2C binding site for the phosphatase ABI1 and ABI2. This blocks the active site of the phosphatase.
Figure \(\PageIndex{24}\) shows an interactive iCn3D model of ABA-bound PYL1 and the Protein Phosphatase 2C ABI1 (pdbid 3kdj)
.png?revision=1&size=bestfit&width=332&height=235)
The PP2C phosphatase (ABI1) is shown in gray with its active site highlighted in green spacefill. The ABA receptor PYL1 is shown in cyan with ABA shown in sticks, colored CPK. The CL2 loop of the ABA-bound PYL1 receptor is shown in red spacefill. It projects into the PP2C active site, inhibiting its activity.
Light Signaling through Phytochromes
Much of this material derives from Liu, Y., Jafari, F. & Wang, H. Integration of light and hormone signaling pathways in the regulation of plant shade avoidance syndrome. aBIOTECH 2, 131–145 (2021). https://doi.org/10.1007/s42994-021-00038-1. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/. get rid of this
Plants deal with competing plants in regions of high plant density by sensing changes in the intensity and wavelengths of light. Signaling leaves to responses (stem elongation, reduced branching, early flowering, etc) called shade avoidance syndrome (SAS). A photosensory system initiates signaling that alters gene transcription. In the SAS in plants in a large canopy, the upper leaves used red and blue light for photosynthesis. Multiple photoreceptors are used. Some transcription factors are also sensitive to light. For example, PIF3, a transcription factor, binds to light-responsive genes only when it binds to another transcription factor called Pr. Pr is resident in the cytoplasm but moves to the nucleus after altering conformation on absorbing red light.
Legris, M., Ince, Y.Ç. & Fankhauser, C. Molecular mechanisms underlying phytochrome-controlled morphogenesis in plants. Nat Commun 10, 5219 (2019). https://doi.org/10.1038/s41467-019-13045-0. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/. do
Phytochromes are present in bacteria, cyanobacteria, fungi, algae, and land plants. We will focus mostly on phytochromes in Arabidopsis. In land plants, phytochromes are red and far-red light receptors that exist in two forms. They are synthesized in the inactive Pr state, which upon light absorption converts to the active Pfr conformation. Pfr is inactivated upon far-red (FR) light absorption or through thermal relaxation, which depends on temperature. Phytochromes act as dimers, resulting in three possible phytochrome species: Pr–Pr, Pfr–Pr, and Pfr–Pfr. Pr and Pfr have different absorption maxima, but due to overlapping spectra both conformers are always present in the light while only prolonged darkness returns all phytochrome to Pr. Given that phytochrome responses depend on the proportion of Pfr conformers, signaling is influenced by a combination of light quantity, color, and temperature. These features of phytochromes are summarized in Figure \(\PageIndex{25}\).
Panel a shows factors that control phytochrome activity. Phytochromes exist in two conformations, Pr and Pfr, the latter being the active form. They exist as dimers so three species can be found. Each monomer can be activated by red light (R) and inactivated by far-red light (FR) or by thermal reversion, a process that depends on temperature (T). At least in the case of phyB, Pfr in heterodimers reverts much faster than that in homodimers, allowing phyB to perceive temperature both during the day and during the night.
Panel b shows plant phytochrome absorption spectra of the Pr and Pfr conformations. In dark-adapted seedlings, phytochromes are in the Pr form. Upon a saturating R pulse, due to overlapping absorption spectra of Pr and Pfr, only 87% of Pfr is achieved.
Panel c shows action spectra for phyA and phyB in the control of hypocotyl elongation. Fluence rate (number of particles passing per unit time) response curves are measured at different wavelengths and fluence rate that leads to 40% inhibition compared with dark control is determined. To specifically determine action spectra for phyA and phyB, for phyB the curve was performed with phyB-GFP/phyAphyB seedlings, and for phyA using phyB-5 seedlings. Values are relative to the response obtained at the most efficient wavelength in each case
Plant phytochrome structure
Plant phytochromes are dimeric, each monomer consisting of ~1150 amino acids. The chromophore, a linear tetrapyrrole named phytochromobilin (PΦB), whose structure is shown in Figure \(\PageIndex{26}\), is attached to the protein.
Figure \(\PageIndex{26}\): Structure of phytochromobilin (PΦB)
The domain structure of phytochromes is shown in Figure \(\PageIndex{27}\).
The apoprotein can be divided into the N-terminal PSM, which consists of the N-terminal extension (NTE), for which structural information remains scarce, and three structurally related domains Period/Arnt/SIM (PAS), cGMP phosphodiesterase/adenylyl cyclase/FhlA (GAF), and a phytochrome-specific domain (PHY) and a C-terminal module (CTM) comprising two PAS domains and a histidine kinase-related domain (HKRD). The chromophore is bound covalently to a conserved cysteine in the GAF domain, which has intrinsic chromophore lyase activity. Light perception triggers a Z to E isomerization around the C15–C16 double bond of PΦB, which leads to a cascade of structural modifications in the protein. Figure \(\PageIndex{28}\) shows an interactive iCn3D model of phytochrome (Deinococcus) Pfr form in the Photoactivated State (5C5K)
%25C2%25A0Pfr_form_in_the_Photoactivated_State_(5C5K).png?revision=1&size=bestfit&width=298&height=309)
The protein is shown in its active dimeric state. One chain is shown in secondary structure colors and the other in cyan. The chromophore (heme derivatives) are shown in spacefill in both subunits. Side chains surrounding the chromophore are shown in colored sticks in the cyan chain.
At first glance, the presence of the histidine kinase-related domain (HKRD) would seem to suggest that phytochromes transduce their signal through the C-terminal module (CTM). Although many bacterial and cyanobacterial phytochromes have a C-terminal histidine kinase domain and act as light-regulated histidine kinases, plant phytochromes are not histidine kinases, and their role as Ser/Thr kinases remains contentious. The photosensory module (PSM) fused to a nuclear localization signal and a dimerization sequence is sufficient to restore most phyB functions, pointing to key signaling functions of the PSM.
Major The major of the plant phytochrome CTM are dimerization, nuclear import, and localization to sub-nuclear structures known as photobodies. However, it was recently shown that the C-terminal part of phyB also engages in light-regulated interactions and regulation of PIF activity. Moreover, the activity of the CTM is controlled by post-translational modification with SUMOylation limiting the ability of active phyB to interact with downstream signaling targets thereby limiting light responses. In addition, the CTM modulates active (Pfr) phytochrome levels with the HKRD inhibiting the Pr–Pfr photoconversion while the PAS–PAS promotes thermal reversion. Hence, while the division of plant phytochromes into PSM and a CTM helps describe the molecule, both parts of the photoreceptor contribute to the regulation of active Pfr levels and downstream signaling activities.
Figure \(\PageIndex{29}\) shows a simplified mechanism for phytochrome control of transcription factors in different light environments.
Panel a shows the response below the soil surface during growth in partial or a complete absence of light (called etiolated growth). For simplicity, we consider that phytochromes remain inactive (Pr) below the soil surface, which results in the accumulation of transcription factors PIFs, EIN3, and ARFs and subsequent induction of etiolation and auxin response genes. The COP1/SPA ubiquitin E3 ligase accumulates in dark and leads to proteasome-mediated degradation of HY5, a transcription factor that suppresses the expression of genes required for etiolation and induces expression of genes required for de-etiolation.
Pane b shows changes that occur when light intensity increases (de-etiolation). Light perception activates phytochromes (Pfr) which promote de-etiolation by directly inhibiting PIFs and EIN3, and indirectly inhibiting ARFs by stabilizing Aux/IAA proteins. The Pfr form of either phyA or phyB interacts with SPA proteins, resulting in the inhibition of COP1/SPA. This results in the stabilization of HY5 leading to the induction of de-etiolation-related gene expression and repression of etiolation genes.
Pane c shows de-etiolated plant in response to shade (reduced R/FR). Low R/FR in shade reduces the fraction of active phytochrome (Pfr/Ptot). PIFs accumulate and induce growth-promoting gene expression. In addition, PIFs induce a negative feedback loop exemplified by HFR1 expression. HFR1 (and other HLH proteins) binds to PIFs forming non-DNA-binding heterodimers. COP1/SPA is also involved in this loop by leading HFR1 to proteasome-mediated degradation. Arrows indicate positive regulation, blunt-ended arrows indicate negative regulation and dotted-lined arrows indicate nucleo-cytoplasmatic relocalization
Figure \(\PageIndex{30}\) summarizes how phytochromes affect transcription.
Panel a (top to bottom) shows sequential steps by which Pfr inhibits PIFs. Top: PfrA interacts with PIF1 and PIF3 while PfrB interacts with PIF1–PIF8. Middle left: for PIF1, 3, and 4 phytochrome inhibits DNA binding. Middle right: Interaction with Pfr leads to the phosphorylation of PIFs. Many kinases have been found to phosphorylate PIFs (see text) with PPKs phosphorylating PIFs in response to light. Bottom: after light-induced phosphorylation, PIF3 is degraded by LRBs and EBFs with phyB co-degradation occurring in the LRB-mediated process (left, center), phosphorylated PIF7 interacts with 14-3-3 proteins and remains in the cytoplasm (right).
Panel b shows other mechanisms of transcriptional control by phytochromes. Left: PfrA and PfrB interact with SPA and inhibit the COP1/SPA complex. Center: PfrB interacts with EIN3 to promote ERF-mediated EIN3 degradation. Right: PfrA and PfrB interact with Aux/IAA to prevent their degradation by SCFTIR1/AFB.
Panel c shows Patterns of PIF abundance depending on the developmental state and growth conditions. In etiolated seedlings PIFs accumulate to high levels, promoting etiolated growth. Upon light exposure, PIFs are rapidly degraded in a phytochrome-dependent manner, with half-lives of ~5 min for PIF1 and PIF5, and ~10 min for PIF3 and PIF4 (left). In contrast, in light-grown seedlings PIFs are under strong transcriptional control, allowing accumulation of the protein even in conditions when phytochrome activity is predicted to be high (right), SD (short days), LD (long days)




v2.png?revision=1&size=bestfit&width=302&height=290)