28.6: Phosphatases
- Page ID
- 14993
Introduction
Protein kinases phosphorylate proteins in a process that can either activate or inhibit the target protein's activities. To control signaling processes, the activities altered by protein phosphorylation can be readily reversed by dephosphorylation of the Ser-, Thr- or Tyr-phosphoesters by simple hydrolysis. These reactions are catalyzed by protein phosphatases. Some of these phosphatases also cleave phosphates from lipids as well.
There are three main families of phosphatases, phospho-Tyr phosphatases (PTP), the phospho-Ser/Thr phosphatases, and those that cleave both. Of all phosphorylation sites, most (86%) are on serine, 12% involve threonine and about 2% are on tyrosine. They can also be categorized by molecular sizes, inhibitors, divalent cation requirements, etc. In contrast to kinases which differ in the structure of their catalytic domains, many protein phosphatases (PPs, also abbreviated Ppp for Protein phosphatases) gain specificity by binding protein cofactors which facilitate translocation and binding to specific phosphoproteins. The active phosphatase hence often consists of a complex of the phosphatase catalytic subunit and a regulatory subunit. Regulatory subunits for Tyr phosphatases may contain a SH2 domain allowing binding of the binary complex to autophosphorylated membrane receptor Tyr kinases.
We'll consider examples of all four families. They recognize target proteins through protein:protein interactions and specific binding site motifs. There are over 10,000 pSer and p-Thr sites on target proteins, so targeting specific sites must be quite nuanced.
Serine/Threonine Phosphatases
Important Ser/Thr phosphatases (PPs for Protein Phosphatases) include:
- Protein phosphatase 1 (PP-1 or Ppp1) - This is the most abundant PPP in humans. Different regulatory subunits target this to the liver glycogen particles (GL subunit), striated muscle glycogen, and sarcoplasmic reticulum (GM subunit) or smooth muscle fibers (M subunit). It is also present in the nucleus where it is presumably involved in the regulation of transcription factors. It is also involved in RNA splicing and signaling at neural synapses.
- Protein phosphatase 2A (PP-2A or Ppp2) - is a trimer with catalytic, regulatory, and scaffolding (also regulatory) structural subunits. It is found mainly in the cytoplasm and is involved in a myriad of cellular processes.
- Protein phosphatase 2B (PP-2B or Ppp3) - also called calcineurin or Ca2+/Calmodulin dependent protein phosphatase - It consists of a catalytic subunit (calcineurin A) and a regulatory, calcium-binding subunit (calcineurin B). It is inhibited by the complex of the immunosuppressant cyclosporin and FK506 with immunophilins. PP2B regulates PKA and PKC
- (PP2C) -
The catalytic subunits of PP1, 2A, and 2B share a great deal of amino acid homology, and based on this homology, belong to one family. PP2C belongs to another family. PPs are often categorized into three other families including, phosphoprotein phosphatases (PPPs) and metal-dependent protein phosphatases (PPMs). There are about 30 catalytic PP subunits (many fold fewer than Ser/Thr Kinases). They gain specificity by binding numerous modulatory regulatory subunits.
As with other proteins, the names given to the proteins when discovered often do not reflect an organization scheme that would name different members based on structural similarities. PP-1, 2A, and 2B are better named Ppp1, Ppp2, and Ppp3 which denote members of the Protein PP (PPP) family. PP-2C would be named Ppm1 as the first member of the PPN family. All PPPs have three short sequence motifs that bind divalent cations.
In addition, older names for the given PPs referred to both the catalytic subunit and the dimer with the regulatory subunits. For clarity, the name of the catalytic protein phosphatase 1 is PP1c, and its regulatory subunits as RIPPOs, regulatory interactors of protein phosphatase one.
Protein Phosphate-1 (PP-1):
PP-1 is involved in many signaling pathways that control cell division, protein synthesis, etc. It catalyzes most serine–threonine dephosphorylation in cells. It is perhaps best known for its regulation of glycogen mobilization. Insulin signals the well-fed state in healthy people and promotes glucose uptake through the GPCR insulin receptor which we discussed in Chapter 12.4. Under these condition excess glucose is used to elongate glycogen, our main carbohydrate energy storage polymer. In contrast, the starving or low energy state is signaled by the hormone glucagon. You would expect that signaling pathways activated in the presence of insulin would promote glycogen synthesis and inhibit glycogen breakdown. PP-1 is a key factor in the regulation of both processes:
- Insulin activation of glycogen synthesis - PP1 dephosphorylates glycogen synthase (the enzyme that synthesizes glycogen) and in the process ACTIVATES it.
- Insulin inhibition of glycogen breakdown - PPI dephosphorylates two key enzymes involved in glycogen breakdown, phosphorylase kinase and glycogen phosphorylase a (with a pSer14), and in the process INHIBITS them.
PP1c interacts with many different regulatory subunits (RIPPOs) forming unique heterodimers. The regulatory subunits also bind potential substrates for PP1c and help localize the enzyme. at sites. The regulatory subunit involved in the PP1c effect on glycogen is called the glycogen-targeting subunit, GM. There are 7 such regulatory subunits involved in glycogen metabolism. GM (RGL) is expressed in muscles and GL in the liver. All of the regulatory subunits (unfortunately called G-subunits) have a conserved RVxF amino acid sequence which interacts with specific sites on the catalytic subunit typically distal to the active phosphatase site. Binding through the RVxF sequence does not affect the active site of the PP-1c. The binding of the regulatory subunit to the PP-1c can also occur outside of the canonical RVxF sequence. The regulatory subunits also have starch binding domain (SBD), also called the carbohydrate-binding module (CBM21). The subunits are often highly disordered until they are bound.
Figure \(\PageIndex{1}\) shows an interactive iCn3D model of protein phosphatase 1 (PP1) bound to the muscle glycogen-targeting subunit (Gm) and microcystin (6DNO) and the toxin microcystin.
_-muscle_glycogen-targeting_subunit_(Gm)_-microcystin6DNO.png?revision=1&size=bestfit&width=286&height=282)
PP-1 alpha catalytic chain is shown in gray. Its active site, where it binds Ser/Thr phosphorylated proteins is shown in magenta. Bound in that site is the toxin microcystin. The small chain shown in cyan is protein phosphatase 1 regulatory subunit 3A. PP-1 binds to its regulatory sequence to a 65RVxF68 sequence common on many regulatory PP-1 subunits. This subunit contains a serine 67 which is phosphorylation by protein kinase A. This inhibits the binding of the catalytic and regulatory subunits. (PKA) of the “x” residue in the GM RVxF motif, Ser67GM, inhibits PP1 binding (16). Val79 and Lys80 on GM also form a motif that binds a corresponding pocket in the catalytic subunit.
Note the pi-cation (red) and pi-stack (blue) interactions from Phe 68 of the GM 65RVSF68 motif of the regulatory subunit to the PPI catalytic subunit. Microcystin is 7-mer peptide ring that has 5 noncanonical amino acids and 2 regular ones. A covalent bond from Cys273 of PP1 (labeled in the above iCn3D model) to the methyl-dehydroalanine (Mdha) of the toxin forms. Microcystins are produced by toxic cyanobacteria and are very toxic and lethal, especially to animals including humans that drink water contaminated with the cyanobacteria. They will pose a greater threat in a warming world from climate change. They also bind to PP-2A.
Another example of a regulatory subunit for PP1 is the PP1 nuclear target subunit (PNUTS). The activity of the complex in the nucleus regulates the phosphorylation state o many proteins involved in the cell cycle including p53, a tumor suppressor in many tumors. It also regulated chromatin structure and RNA processing. As with the muscle glycogen-targeting subunit (Gm), PNUTS is intrinsically disordered when not bound to PP1 and is very extended when bound. The catalytic subunits of PP1c and PP2A have an acidic, hydrophobic, and C-terminal grove. PNUTS binds one of the substrate groves at an arginine subsite, but not the active site.
So far we can say the specificity of the binding of PPP is determined to some degree by the regulatory subunits. We turn to the contributions of the catalytic specificity of PP-1c when we compare it to PP-2Ac below.
Mechanism of PP1-A and PP-2A and B
The catalytical subunits of PP-1 and PP-2 are very homologous with nearly identical key active site side chains. The site contains two Mn2+ ions very near each other as shown in Figure \(\PageIndex{2}\).
The figure crudely shows each Mn2+ is octahedrally coordinated to side chains and one water oxygen for each metal ion. Water 2 (or more likely OH-) and an aspartate D92 oxygen bridge the two ions. The phosphate of the target Ser-OPO32- or Thr-OPO32-, or an inhibitor such as tungstate, also bridges the metal ion.
Figure \(\PageIndex{3}\) shows an interactive iCn3D model of Human Protein Phosphatase 1 Active Site Residue (4mov) which shows the same active site residues
.png?revision=1&size=bestfit&width=426&height=354)
The metals singly or in combination probably reduce the pKa of bound water to produce the deprotonated hydroxide, which engages in an SN2 attack on the phosphate. Hence the metal ions act as an electrostatic catalyst.
The subtle differences in the active site and the three groups contribute to the specificity of the PPPs.
Comparison of the catalytic subunits of PP1A the PP2A
As mentioned above, substrate specificity is altered by subtle changes in the active site and three groves of PPPs. Figure \(\PageIndex{4}\) shows the acidic groves for PP1c and PP2Ac.
The color is the electrostatic surface potential with red indicating negative and blue (notably absent) positive. The acidic groove is stronger in PP1c. The asterixis * shows the catalytic cleft containing two Mn2+ ions. Hoermann et al. Nature Communication | (2020) 11:3583 | https://doi.org/10.1038/s41467-020-17334-x. Creative Commons Attribution. 4.0 International License, http://creativecommons.org/licenses/by/4.0/.
The negative acidic grove is highly enriched in negatively charged side chains. Figure \(\PageIndex{5}\) shows the actual amino acids contributing to the negative electrostatic potential in aligned PP1c and PP2Ac.
The orange spheres show the location of acidic side chains. PP1c is shown in blue/black and PP2Ac in red/gray.
The stronger acidic groove in PP1Ac gives it great preferences for pSer/pThr-protein targets with basic motifs than PP2Ac in a fashion that is independent of the bound regulator subunit. In contrast, PP2A needs to interact with regulatory subunits with more acidic composition to target basic motifs in protein targets. These features for PP1c and PP2Ac are compared in Figure \(\PageIndex{6}\).
Panel (d) shows that the holoenzyme for PP1 has a great preference for positive basic motifs than the holoenzyme for PP2A, which needs to associate with a negatively charged regulatory subunit for activity towards target proteins with basic motifs.
Panel (e) also shows that the catalytic subunits of both PP1 and PP2A prefer p-Thr protein targets compared to p-Velocity vs substrate graphs show these effects as well. e Both, PP1 and PP2A holoenzymes have a preference for pT due to higher catalytic efficiency of their respective catalytic subunits towards pT over pS. Hoermann et al. ibid
Figure \(\PageIndex{7}\) shows an interactive iCn3D model of the protein phosphatase 2A catalytic subunit in complex with a larger regulatory subunit and bound to the phosphatase inhibitor and tumor promoter okadaic acid (2IE4). The toxin, found in sponges and shellfish, is produced by dinoflagellates.
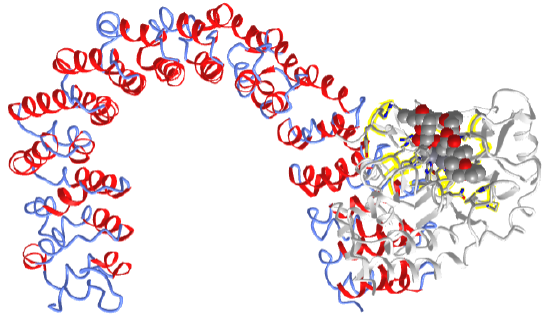
The regulatory subunit is shown in secondary structure color. This scaffolding protein is shaped like a horseshoe. The phosphatase inhibitor okadaic acids are shown in spacefill bound to the catalytic subunit shown in gray. The side chains of the catalytic subunit interacting with the 2 Mn2+ ions are shown in CPK-colored sticks (zoom in to see them). On binding the catalytic subunit, the scaffolding regulatory subunit is quite flexible and adaptable in interacting with other proteins.
Protein Phosphate 2B: Calcineurin (CN)
Calcineurin (CN), or PP2B, is depended on Ca2+. It is involved in the development, immune signaling, and heart function. It consists of a catalytic site (CNA) and a calcium-binding regulatory subunit CNB so it is another example of PPP heterodimers. CNA has a catalytic domain and domains that bind CNB (the regulatory subunit), calmodulin (CAM, a calcium-binding protein that we will explore more in the next chapter section), and an autoinhibitory domain that blocks the active site. On Ca2+ release from internal organelles, the ion binds to both CNB and also CAM. These events cause conformational changes that release the bound autoinhibitor.
As with the other phosphatases, much effort has been made to determine how CN interacts with specific pSer- and p-Thr sites on targets. We'll focus on one, the integral membrane Na+/H exchanger 1 (NHE1). This protein is itself regulated by Ca2+ ions and by phosphorylation by kinases we have previously studied, the MAPK ERK1/2 and the JNK kinase. Erk2 phosphorylates NHE1 at 6 Ser/Thr side chains in the recognition sequences name [S/T]P11. Several different phosphatases, including CN, can regulate NHE1 activity through direct dephosphorylation.
CN binds short linear motifs (SLiMs) named PxIxIT and the LxVP that are found in interacting partners including regulatory subunits as well as inhibitors and substrates. As we saw above, the regulatory subunits of PP1A and PP2A are highly disordered. Likewise, SLiMs are on intrinsically disordered regions as well as interacting proteins.
- PxIxIT binds to the catalytic domain of CNA22. It also enables interaction between CN and NHE1.
- LxVP binds to a cleft between the CNA and CNB, which is only available in the active form of the protein.
CN doesn't dephosphorylate multiple nearby p-Ser side chains of NHE1 (pS363, pS723, and pS726) since they are close to the NHE1-PxIxIT interaction sit,e which sterically restricts their binding to the active site. However the 3 other phospo S/T sites on NHE1 (pS771, pS785, and the actual target site (pT779) are far enough away from the NHE1's PxIxIT site so they can interact with the CN active site. Making the T779S mutation shows that dephosphorylation of their phosphorylated version shows a faster rate with pT779 and a slower, yet reasonable rate for pS779. Therefore other specificity factors are in play. A newly discovered very short 4-amino acid site motif in NHE1 including pS779 appears to be a source of selectivity. This TxxP motif in NHE1 is 779TPAP782. Such short recognition motifs are different than the selection of substrates by PP1 which involved multiple domain binding interactions and steric restrictions imposed by them.
Figure \(\PageIndex{8}\) shows a model of the structure of the NHE1 exchanger (left panel a) and the calcineurin CNA/CNB complex.
jd
Figure \(\PageIndex{8}\): Docking motifs mediate the interaction of NHE1ct with CN
The motifs present in NHE1 (LxVP and PxIxIT) in the intrinsically disordered tail of NHE are indicated in the left panel and their corresponding binding sites in the CNA dimer are shown in corresponding colors. The calmodulin binding site is also shown. Erk2 phosphorylation sites in NHE1 are shown are listed in panel C along with the consensus motif sequences (PxIxIT in purple, LxVP in orange, and TRAP (uncolored)). Hendus Altenburger et al. Nature Communication (2019) 10:3489 | https://doi.org/10.1038/s41467-019-11391-7Creative Commons Attribution 4.0 International License. http://creativecommons.org/ licenses/by/4.0/.
Figure \(\PageIndex{9}\) shows an interactive iCn3D model of calcineurin (PP2B) complex bound to a peptide from the Na+ /H+ -exchanger 1 (6NUC)
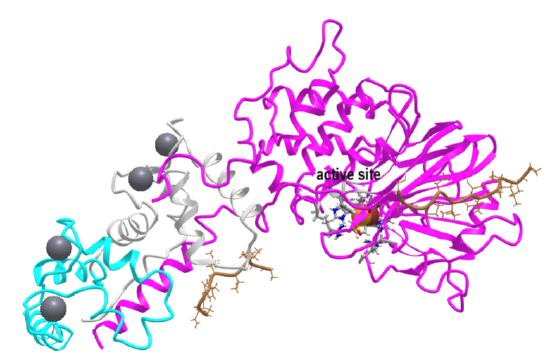
Protein Phosphatase 2C
The protein phosphatase 2C (PP2C) is a member of a family of metal-dependent protein phosphatases sometimes abbreviated as PPMs. (Of course, the other phosphatases we discussed above are also metal-dependent.) The required Mg2+ or Mn2+. PP2C is a monomeric enzyme with at least four isoforms in humans. In humans, there are at least 17 members. One is unfortunately named protein phosphatase 1D but also referred to as PPM1D, PP2Cδ, or Wip1). It is involved in heterochromatin silencing and the cycle. Mutations in its gene can accordingly give rise to tumors.
PP2Cs in humans have a binuclear metal cluster which reduces the pKa of water, producing OH- for SN2 attack on the phosphorus in the pSer or pThr in target phosphorylated proteins. Figure \(\PageIndex{10}\) describes binding interactions around the binuclear site in PP2Cα.
Figure \(\PageIndex{10}\): Binding interactions around the binuclear site in PP2Cα. after Pan et al. Sci Rep 5, 8560 (2015). https://doi.org/10.1038/srep08560htt...cles/srep08560
The metal binding site with many water molecules Interactions with the metals is very different than the PP1 and PP2 active site shown in Figure \(\PageIndex{2}\).
If Cd is bound at the M1 site, the activity of the enzyme is blocked so it is required for catalysis. Making the mutations affecting M2 (D38A and D38K) suggests that M2 is involved in binding the phosphate of the substrate, and also stabilizes the transition state and the leaving group in the reaction. H62 probably acts as a general acid catalyst.
Not all PPCs require both metal ions. The plant hormone abscisic acid regulates stress responses in plants. When it binds to a particular receptor called PYL1 (alternative name PYR1-like protein 1), the receptor interacts with a PP-2C called ABI1 (also called Absisic acid-insensitive 1). Figure \(\PageIndex{11}\) shows an interactive iCn3D model of the ternary complex of Abscisic acid, PYL1 and ABI1 (phospholipase 2C) (3KDJ)
%25C2%25A03KDJ.png?revision=1&size=bestfit&width=459&height=305)
The catalytic subunit, which in contrast to PP1, PP2A, and PP2B, has only 1 Mn2+ ion, is shown in gray with amino acids side chains interacting with the metal ion labeled. The cyan subunit is the receptor of abscisic acid, which is shown in spacefill.
Protein Tyrosine Phosphatases
Protein Tyr phosphatases (PTPs) consist of receptor-like (transmembrane) and intracellular Tyr phosphatases. They more resemble tyrosine kinases in their complexity than the Ser/Thr phosphatases. There are about 100 PTPs in the genome, a number similar to the number of protein tyrosine kinases. PTPs have an active site Cys in a CX5R-(S/T) motif with an active site Cys nucleophile and an Arg in the phosphate binding (P) loop. Some examples we will discuss include:
- Low molecular weight PTPase - These have roles in the metabolism and differentiation of cells. They have a molecular weight of 18,000 and have an active site CX5R-(S/T) motif, where the C (Cys) is an active site nucleophile.
- PTP1B - dephosphorylates many cell surface receptors (insulin, EGF, PDGF) that have been phosphorylated on Tyr residues. Its main activity seems to dephosphorylate nascent receptors in the endoplasmic reticulum before they get to the final cell membrane destination.
- Tyrosine phosphatase nonreceptor type 11, ptpn11, commonly called SHP2
Figure \(\PageIndex{12}\) shows the protein tyrosine phosphatase (PTP) superfamily.
In contrast to the active site of the Ser/Thr phosphatases like PP-1, PP-2A and PP-2B, the active sites of protein tyrosine phosphatases (PTPs) do not have a bimetal ion cluster in the active site. Rather they all have an active site cysteine that acts as a nucleophilic catalyst in the hydrolysis of the p-Try phosphoester bond. The active site PTP domain is found in all of the proteins, so all use similar catalytic mechanisms shown in Figure \(\PageIndex{13}\).
The phosphotyrosine side chain of the target phosphoprotein binds in the phosphate-binding P-loop (H/VCxxxxxRS/T), which contains the nucleophilic Cys 12 and Arg 18 that stabilizes the charge on the phosphate. The Asp in the WPD loop positioned across from the nucleophilic Cys 12 acts as a general acid. Since the active site is nearly identical in the PTPs, it has been hard to design drugs that bind to the active site but that are also selective for specific PTPs.
Low molecular weight protein tyrosine phosphatase - LMW-PTP
This protein tyrosine phosphatase is the simplest of all in structure. It has the phosphate-binding P-loop (12CxxxxxR18) with the nucleophilic Cys 12 and Arg 18 that stabilizes the charge on the phosphate. It does not have the conserved WPD loop but deploys Asp 129 across from Cys 12 as a general acid. This enzyme exists as two main isozymes, A and B. Figure \(\PageIndex{14}\) shows an interactive iCn3D model of human low molecular weight protein tyrosine phosphatase bound to sulfate (1xww)
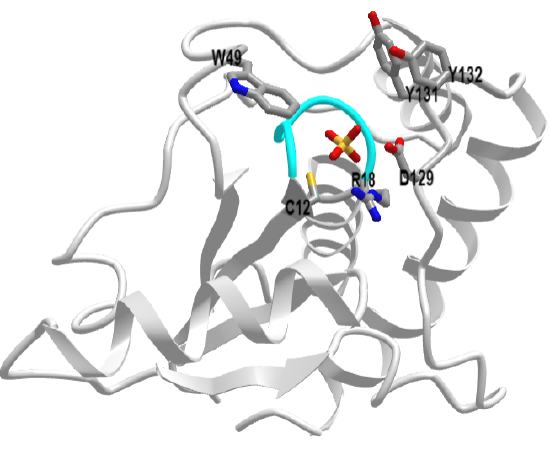
The active site is deep as shown in Figure \(\PageIndex{15}\).
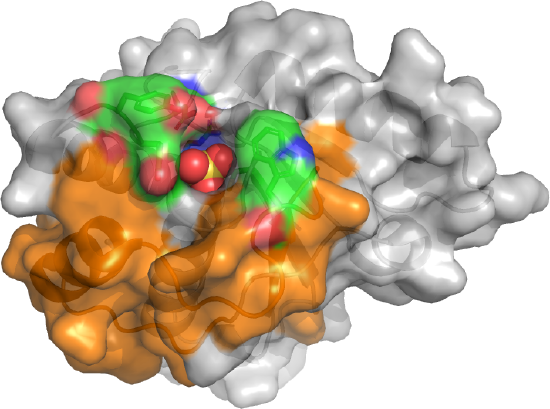
The sulfate, a mimetic for the phosphate on the p-Try protein target, is deeply buried. The CPK-colored surface (green red, blue) around the sulfate are the side chains of Tyrosine 131 and 132 as well as Trp 49. Y131 and Y132 are part of a loop containing Asp 129, the general acid. This loop is analogous to the WPD loop. The three aromatic amino acids on top of the pocket make it deep enough that pSer and pThr side chains can't reach the active site nucleophile Cys 12. Trp 49 is also in a variable loop (34 amino acids, shown as an orange surface) that differentiates two of the major isozymes, the A and B forms, and contributes to substrate binding specificity.
Tyrosine-protein phosphatase non-receptor type 1, also known as PTPN1 or PTP1B PTP1B
PTP-1B regulates the endoplasmic reticulum unfolded protein response and is involved in insulin JAK/STAT and HER2 (ErbB2) signaling. It has a full-length form (MW 50,000) and a C-terminal shortened form (37,000)
Given the difficulty in targeting the active site which is common in all PTPs, efforts have concentrated on the development of allosteric inhibitors that bind to exosites removed from the active site. An example is trodusquemine (MSI-1436) used in the treatment of obesity and type 2 diabetes. It binds much more tightly to the full-length form.
Figure \(\PageIndex{16}\) shows an interactive iCn3D model of the human Protein Tyrosine Phosphatase 1B (1-301) in complex with the inhibitor OTA (5K9W).
Figure \(\PageIndex{16}\): Human Protein Tyrosine Phosphatase 1B (1-301) in complex with the inhibitor OTA (5K9W). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...fTurfTTFiW2b47
The P-loop is shown in magenta, the WPD loop in cyan, and the substrate binding loop (SBL), which allows entry of pTyr but not pSer and pThr, in blue. The key side chain in the P-loop (Cys 215 and Arg 221) as well as the catalytic general acid (Asp 181) are shown in sticks and labeled. The inhibitor is shown in spacefill, and CPK colors. The movement of the WPD loop is rate-limiting for the hydrolysis of P-Tyr esters. On binding, the WPD starts to close, and in the process Arg 221 moves to form salt bridges with the phosphate. Full closure of the WPD follows, which positions Asp 181 for general acid catalysis. Key interactions of PTP-1B phosphoprotein in insulin and cytokine signaling, are shown in Figure \(\PageIndex{17}\).:
Figure \(\PageIndex{17}\): Protein tyrosine phosphatase 1B (PTP1B) and its effects on signaling. Maja Köhn ACS Cent. Sci. 2020, 6, 4, 467–477. Publication Date: March 13, 2020. https://doi.org/10.1021/acscentsci.9b00909. This is an open-access article published under an ACS Author Choice License, which permits
copying and redistribution of the article or any adaptations for non-commercial purposes.
Panel (A, left) shows how PTP1B dephosphorylates the insulin receptor and the insulin receptor substrate (IRS), which we have explored in a previous section. Panel (A, right) show its activity in the JAK/STAT pathway, which we have all seen previously. One cytokine receptor that it regulates is the leptin receptor. The hormone leptin, released from fat cells (adipocytes) is a key regulator of lipid metabolism. Pane B shows the structures of key inhibitors of PTP-1B.
Protein tyrosine phosphatase nonreceptor type 11 (ptpn11) also known as SHP2 (SH2-domain containing phosphatase-2)
This is an example of another phosphatase in which a mutation leads to cancer. It is downstream and activated by most receptor tyrosine kinases (RTKs) involved in the activation of the MAPK pathway with its ultimate links into the nucleus and activation of gene transcription.
Figure \(\PageIndex{18}\) shows an interactive iCn3D model of Non-receptor Protein Tyrosine Phosphatase SHP2 in Complex with Allosteric Inhibitor Pyrazolo-pyrimidinone 5 (6MDB)
.png?revision=1&size=bestfit&width=409&height=294)
The phosphatase domain is shown in gray. The N- and C-terminal SH2 domains are shown in green and blue, respectively. The allosteric inhibitor is shown in spacefill and CPK colors. The P-loop in the catalytic domain is shown in red with the Cys 459 (active site nucleophile) and R465 (stabilizer of phosphate in the complex) shown in sticks, CPK colors, and labeled. The bound inhibitor is especially interesting as it binds at an allosteric site. As mentioned above, it is very difficult to design specific inhibitors that target just one PTP given their common active sites and mechanisms. Figure \(\PageIndex{19}\) shows multiple features of SHP2.
Figure \(\PageIndex{19}\):. SH2-domain containing phosphatase-2 SHP2. Köhn ibid.
Panel (A) shows how SHP2 recruited to phosphorylated RTKs activates the MAPK pathway. The dotted line indicates multiple steps. Upon receptor activation, SHP2 is recruited in different ways to activate the MAPK pathway.
Panel (B) shows that in the inactive state, the N-terminal SH2 domain (green) blocks access to the active site. When the N- and C-terminal domains bind pY residues in a single pY-protein, two pY-proteins, or pY residues on its C-terminal tail, a conformational change ensues opening the active site (5EHR).
Panel (C) shows how another allosteric inhibitor keeps the protein in a closed state (5EHR).
Panel (D) shows how SHP2 can decrease T-cell responses through the MHC:Tcell receptor (TCR) complex. Tumor cells express a ligand called PD-L1, which binds to the PD1 receptor on the T cell surface. After binding, SHP2 is recruited to PD1, decreasing T cell activation. This is not a good thing since it inhibits the immune response to the cancer cell.
Dual Specificity Phosphatases (DUSPs)
Another important phosphatase is phosphatidylinositol 3,4,5-trisphosphate 3-phosphatase and dual-specificity protein phosphatase PTEN (phosphatase and tensin homolog). Dual-specificity protein phosphatase hydrolyzes pTyr- as well as pSer- and pThr- phosphoesters in target proteins. They don't require divalent metal cations and are closer in structure to protein tyrosine phosphatases. They have an active site cysteine in a P-loop also containing arginine. In addition, they are lipid phosphatases, removing phosphate from the inositol ring from phosphatidyl inositol derivatives. These both impact many signaling pathways. Its activity as a lipid phosphatase makes it a tumor suppressor protein as it inhibits the PI3K-AKT/PKB signaling pathway by dephosphorylating phosphoinositides. Hence it modulates both AKT and mTor pathways.
The domain structure of PTEN is shown in Figure \(\PageIndex{20}\).
The C2 domain enables phospholipid binding. Multiple post-translational modification sites are indicated. The PEST motif is sequence rich in proline (P), glutamic acid (E), serine (S), and threonine (T) and bounded by positively charged amino acids (Lys, Arg, or His) that act as signals for protein degradation. The PDZ domain, often found at the C-terminal of signaling proteins, acts as a scaffolding site for interaction with other signaling proteins. In the next chapter section, we will consider redox signaling, for which PTEN is a great example. Disulfide formation (in a more oxidizing environment) between the nucleophilic Cys 124 and a nearby Cys 71 (figure above) inhibits PTEN phosphatase activity.
Figure \(\PageIndex{21}\) shows an interactive iCn3D model of an AlphaFold computational model of full-length human PTEN (Uniprot P60484).
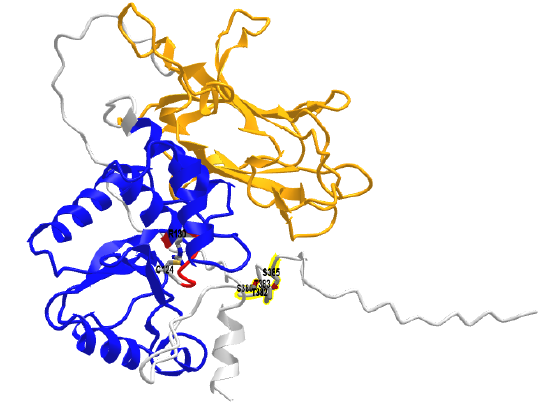
The phosphatase (PTPase) domain is shown in blue and the C2 domain is in orange. The P-loop is in red with the active site Cys 124 and R130 in colored sticks and labeled. The backbone of the highly extended intrinsically disordered C-terminus region is shown in gray. It contains the clustered residues Ser 380, Thr 382, Thr 383, and Ser 385 (shown in colored sticks and labeled) that are sites for phosphorylation by activated kinases.
iFigure \(\PageIndex{22}\) shows key molecules dephosphorylated by PTEN, including the lipid PIP3, and Thr 308 and Ser 473 on AKT.
When PTEN dephosphorylates Akt1, it inhibits AKT activity and effectively antagonizes the main PTEN-PIP3-PDK1-Akt pathway. PTEN is considered a tumor suppressor for this reason. Mutations in PTEN hence are associated with cancer.




_in_complex_with_the_inhibitor_OTA_(5K9W).png?revision=1&size=bestfit&height=300)