3.5: Antibodies in Quantitation and In Vivo Detection
- Page ID
- 64709
Written by ?? Needs work
We will describe three different uses of antibodies for the detection and quantitation of analytes.
Enzyme-Linked Immunosorbent Assay (ELISA)
(work derived from the Human Atlas Project)
Since the very first use of antibodies for the detection of antigens, many different technologies have been developed that make use of the antibodies' capability to bind to other molecules. During the 1950s, Yalow and Berson developed a method where radioactivity is used to determine the amount of an analyte in a solution. This 'radioimmunoassay' (RIA), for which Yarlow received the Nobel prize in 1977, was a very sensitive method for the detection of hormones but using radioactivity for antigen detection is not safe and suitable for general use. Hence, an alternative procedure was developed by linking enzymes to antibodies instead of a radioactive molecule, and by adhering molecules to surfaces. This is the basis of the widely-used "enzyme-linked immunosorbent assay" (ELISA). Many variants of experimental procedures have been developed, and it is common to build assays using more than one antibody to detect a target of interest (see Figure 1). To further enhance the possibilities offered by the immunoassay format, applications based on microarrays have been developed which allow the measurement of more than one analyte in a single reaction chamber (see below). Different detection methods are described in Figure \(\PageIndex{1}\).
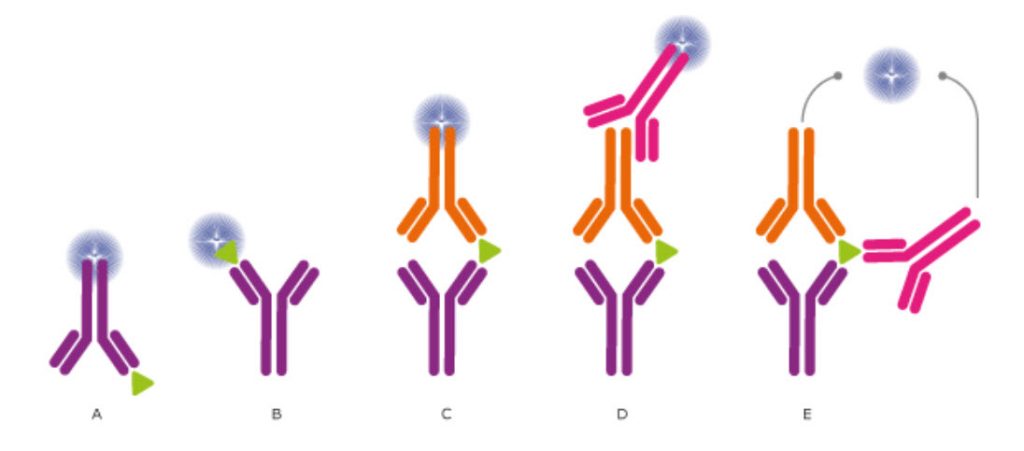
Figure \(\PageIndex{1}\): Different detection methods for ELISA and Other Immunossays. (CC BY-SA 3.0; The Human Atlas Project)
In ELISA assays, the antibodies may (A) detect an immobilized antigen, (B) capture a labeled antigen, (C) capture an unlabeled antigen and use a second, labeled antibody to detect the captured antigen, or (D) use a third antibody for detection, or even use two antibodies for detection (E). Direct labeling of the antibody or antigen as in (A), (B), and (C) is the simplest and fastest method for detection. Using a secondary antibody as a detection method, as shown in (D) and (E), will further increase the sensitivity and selectivity of the analysis. The method used in (D) also allows greater flexibility, whereas method (E) further increases the specificity, as three antibodies must bind the antigen in order to produce a reporter molecule. Out of the presented assays, the most commonly used concepts are shown in (C) and (D). We will describe how ELISA data is used to determine concentrations of analytes in Chapter 5.
A new era in immunoassays started with the development of a technology called microarrays. The term microarray most commonly describes the ordered organization of small-volume droplets that have dried on a small surface area. The reaction dimensions are miniaturized so that many assays can be performed in multiple samples in parallel. Glass slides can be used and robotic pipettors can deposit very small drops of liquid (1 nL = 10-9 L on the glass surface in an ordered fashion, with the spots having sizes of around 0.15 mm. Another common technique for multiplexing is to use even smaller and color-coded particles (diameter of 0.005 mm). These particles can be coated with antibodies to fish out the analyte from the solution.
Microarray assays are used for parallel analysis of DNA and RNA molecules. In addition, multiplexed techniques are used to determine many proteins simultaneously, and to study post-translation modifications such as phosphorylation. Another example is the analysis of antibodies circulating in the blood of patients. Protein microarrays can reveal the interactions of ligands with the whole protein or larger protein fragments, while peptide microarrays are used to detect small peptides (epitopes) of the proteins that bind antibodies. A typical epitope mapping result is shown in Figure \(\PageIndex{2}\) (Edfors et al., 2014).
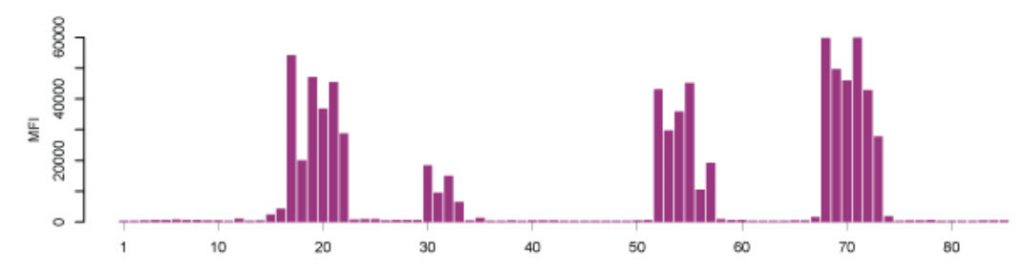
Figure \(\PageIndex{2}\): Epitope Mapping of Polyclonal Antibodies. Polyclonal antibodies binding to a peptide array where the result displays four distinct linear epitopes and the consecutive overlapping peptides which are bound. X-axis: peptides, Y-axis: mean fluorescence intensity (MFI). (Edfors et al., 2014) Image from The Human Atlas Project
Synthesizing millions of overlapping peptides with only one amino acid residue shift on such arrays enables the mapping of antibody binding regions at high resolution. This gives a very detailed analysis of the linear (continuous) epitopes recognized by an antibody.
Immunohistochemistry - Detecting Proteins in Vivo
(work derived from the Human Atlas Project)
Immunohistochemistry (IHC) is a powerful microscopy-based technique for visualizing cellular components, for instance, proteins or other macromolecules in tissue samples. The strength of IHC is the intuitive visual output that reveals the existence and localization of the target protein in the context of different cell types, biological states, and/or subcellular localization within complex tissues.
The IHC technique was invented during the 1940s (Coons, Creech, & Jones, 1941) and is routinely used as an important tool in health care and pathology for e.g. diagnostic purposes or to stratify patients for optimized treatment regimes. IHC is also widely used in research where molecules of interest are analyzed to study their roles in both healthy and diseased cells and tissues on the molecular, cellular or tissue level. There are many different ways to perform visualization of targets in tissues using IHC or IHC-based methods, and numerous protocols exist for different applications and assays. Even though IHC is generally a robust and established method, new assays often need careful optimization depending on the tissue or on the properties of the target protein, binder-molecule and/or reporter system.
The classical IHC assay is illustrated in Figure \(\PageIndex{3}\) and involves the detection of epitopes expressed by a single protein target within a tissue sample using a "primary antibody" capable of binding those epitopes with high specificity. After the epitope-antibody binding event, a "secondary antibody" capable of binding the primary antibody with high specificity is added. The secondary antibody is coupled to a reporter molecule and after the antibody-antibody binding event, a chemical substrate is added which reacts with the reporter molecule to produce a colored precipitate at the site of the whole epitope-antibody complex.
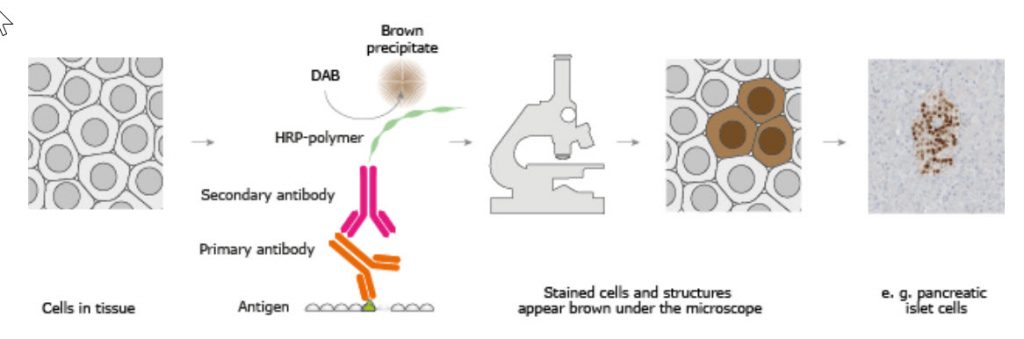
Figure \(\PageIndex{3}\): The Basic Principle of Immunohistochemistry. mage from The Human Atlas Project
In the schematic illustration, a formalin-fixed paraffin-embedded tissue section is stained using a primary antibody directed toward a specific protein target. A solution containing the primary antibody is added to the tissue section and the antibodies are allowed some time to find and bind to their target. After this step, unbound and surplus antibodies are washed away and the secondary antibody is added. The secondary antibody, which carries a linker molecule with horseradish peroxidase (HRP) enzymes, is also allowed some time to bind to the primary antibody, followed by another washing step. After this, 3,3' Diaminobenzidine (DAB) is added. The HRP enzyme transforms the DAB substrate into a brownish precipitate that is deposited in the tissue at the site of the reaction, thus producing a visual representation of where the primary antibody first bound its target.
Tissue preparation
The tissue plays a central role in the experiment and it is important that it is processed so that epitopes and proper morphology is preserved. The most common processing for IHC is to prepare formalin-fixed paraffin-embedded (FFPE) tissue blocks. The purpose of formalin fixation is to produce chemical cross-linking of proteins within the tissue. This terminates all cellular processes and freezes the cellular components at the place and in the conformation at the time of fixation and also to prevent degradation. After adequate fixation, the tissue is further processed and ultimately embedded in paraffin blocks, which are then sectioned into thin slices (usually 4-10µm) using a microtome. The sections are transferred to glass slides and allowed to adhere prior to further processing.
Other methods for fixation besides formalin are sometimes used. These include other types of aldehydes or using different alcohol solutions. The best choice of fixative is very much dependent on the assay. A common alternative to FFPE is to prepare frozen tissue samples. In this case, the tissue is embedded in a cryoprotective medium and frozen, and fixation is performed post-sectioning. Frozen tissues are sectioned in cryostats and have the advantage of short processing times and of better preservation of sensitive epitopes, but can often be inferior to FFPE tissues in terms of preserving histological morphology.
Antigen (epitope) retrieval
A concern associated with cross-linking fixatives like formalin or the length of time spent in the fixative medium is the masking of epitopes, which can obstruct the primary antibody from binding to its target. Especially with FFPE samples, there is often a need to revert some of the chemical crosslinking and "retrieve" the epitopes before proceeding to the actual IHC. There are several antigen retrieval protocols available and the main strategies include treating the tissue slide with heat, digestive enzymes, detergents, or combinations thereof. The most common method for antigen retrieval in FFPE samples is to pressure-boil the tissue slides in an acidic citrate buffer for around 15-20 minutes.
Antibody binding
The quality and specificity of the binding molecule are crucial for any IHC based technique, and the choice of binder can directly affect the outcome, reliability, and possibly also the interpretation of the assay. Antibodies are by far the most common type of binding-molecule used for IHC, and although most antibodies are able to adequately detect the correct molecule of interest, they may also vary greatly in their specificity for their intended target. Antibodies with high specificity are therefore more reliable when interpreting "on-target" binding, since they produce little or no "off-target" binding or "background". Antibodies that are less specific can produce more off-target binding, and the resulting background will possibly interfere with the correct interpretation of the true on-target signals. There are two main types of antibodies; polyclonal antibodies which are a heterogeneous mix of antibodies that bind different epitopes on the target and monoclonal antibodies which bind the same epitope. Polyclonal antibodies are often very potent due to their ability to detect and bind multiple epitopes on the same target. However, the epitopes they bind are often poorly defined, and with multiple and varying epitope-specificities comes the increased likelihood of off-target binding events and background noise. However, the potency of polyclonal antibodies can be advantageous since the concentration of binding events around the on-target molecule usually outweighs potential background noise. A drawback is that polyclonal antibodies are usually limited resources since they are derived from animal sera. Monoclonal antibodies, by contrast, have more continuity since they can be produced in hybridoma cell lines. Monoclonal antibodies are also often well-defined in terms of epitope binding, but can still generate results that are hard to interpret if the specificity is low or if the target epitope is present in low abundance.
Careful optimization and titration of antibody concentration for each assay are needed, since the result is dependent not only on the antibody's specificity and affinity for the target, but also on the concentration and availability of on-target and potential off-target epitopes present in the sample. Adding too many antibodies to the sample will increase the number of possible low-affinity off-target binding events once the on-target epitope(s) are saturated with binders. By lowering the antibody concentration, off-target binding events become rarer as they usually have lower affinity than on-target binding events. The risk when attempting to reduce background while using a low-affinity antibody is that the on-target signals are concomitantly weakened to the point of providing a false negative result.
Other types of binder molecules sometimes used in IHC-based techniques include affibodies, peptides, antibody fragments or other small molecules.
Detection systems
The whole purpose of performing IHC is to obtain a visual representation of where the target can be found within the experimental tissue and preferably and also gain information about the target's expression pattern among heterogeneous cell populations and/or subcellular sites. This is exemplified in Figure \(\PageIndex{4}\), which illustrates how different antibodies are used to visualize different cellular or tissue compartments within a complex tissue. To visualize the target-antibody interaction, some kind of detection system that produces an observable stain or signal is needed. The most common method for introducing a detection system to the experiment is to use a secondary antibody that carries a pre-bound reporter molecule, i.e. enzyme or fluorophore. Secondary antibodies are usually targeted specifically towards antibody molecules from different animal species. For example, if the primary antibody is raised in a rabbit, then the secondary antibody must be raised in another animal and targeted specifically towards rabbit antibodies.
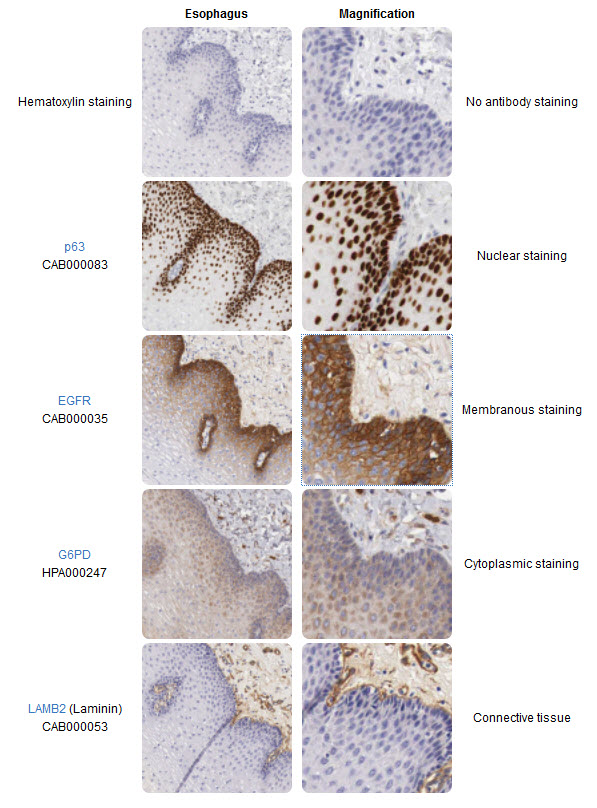
The right column shows a magnification of the corresponding images in the left column. In the IHC image, consecutive sections of a human esophagus stained using four different antibodies allow for direct comparison of different protein expression patterns within the tissue and within subcellular compartments. The top images are only counterstained for hematoxylin for comparison. The p63 antibody stains cell nuclei in a population of cells that reside in the basal part of the esophageal epithelium. The EGFR (Epidermal growth factor receptor) antibody appears to stain the same cell population as p63, but stains cellular membranes instead of nuclei. The G6PD (Glucose-6-phosphate dehydrogenase) antibody stains the cytoplasm of a wider repertoire of esophageal epithelial cells and also cells residing in the connective tissue. The Laminin (LAMB2) antibody stains only cells and structures in the connective tissue underlying the esophagus.
Image from The Human Atlas Project
For FFPE tissue samples the most common detection method is to use enzymatic reactions to generate a colored precipitate at the site of antibody binding. Secondary antibodies with an attached enzyme, e.g. horseradish peroxidase (HRP) or alkaline phosphatase (AP), are capable of converting chromogens like 3,3' Diaminobenzidine (DAB) or 5-bromo-4-chloro-3-indolyl phosphate/ p-nitroblue tetrazolium chloride (BCIP/NBT) into brown or bluish precipitates that are deposited in the tissue at the site of the reaction. Chromogenic stains are observable using light microscopy and are usually very stable over long periods of time, which is beneficial if the experiment needs to be archived or reviewed at a later time point.
For frozen tissue sections, it is more common to use fluorophore-linked secondary antibodies that emit a specific color (usually green, red, or blue) when excited by the correct wavelengths of light. Moreover, fluorophores are usually not stable for long periods of time. However, the benefit of using fluorophores is that they provide an easy method for performing double-labeling experiments where several antibodies directed toward multiple targets are assayed in the same sample. The secondary antibodies need to be targeted towards different primary antibodies and also to be coupled to different fluorophores. The different secondary antibodies are then observed separately by exciting them sequentially with different wavelengths of light. These different excitation results are saved as separate images (or color channels) and may later be overlaid to infer protein co-localizations etc.
Using reporter-carrying secondary antibodies for detection is in itself an amplification step since several secondary antibodies are able to bind a single primary antibody, but sometimes further amplification steps are desired to increase the signal and sensitivity of the experiment. In such cases, the secondary antibody may instead carry "linker molecules", for instance biotin polymers, which are able to recruit a larger number of reporter molecules in subsequent steps. This strategy for amplifying signals is useful for both enzymatic and fluorescent detection methods.
Counterstaining
Immunohistochemical staining using chromogens offers benefits from having a counterstain applied that enhances the contrast and facilitates the observation of histological features. The most common type of counterstain used for FFPE samples is hematoxylin, which stains cellular cytoplasm with a pale bluish color, and stain cell nuclei in a darker bluish nuance. Fluorescent stainings are usually not counterstained with hematoxylin, since the detection method is not based on light microscopy. Instead, the most common way to obtain counterstaining for fluorescence is to label cell nuclei by adding fluorescent dyes that bind nucleic acids.. After the actual immunohistochemical reaction, the only remaining steps are to use a coverslip to seal and protect the sample and for long-term storage. The most common way is to "glue" the coverslip to the sample using commercially available purpose-made resins.
 Add figure
Add figure
Image from NIH ImageJ-Programmpaket
Specific examples
IHC is widely used in both research and clinical practice. The Human Protein Atlas (HPA) project is a prime example of how high-throughput IHC is used to achieve large-scale mapping of the human proteome in a multitude of tissues, cancers, and cells. In the HPA project, a streamlined in-house large-scale antibody production chain facilitates the generation of specific antibodies, which after passing basic characterization and validation regimes, are used to systematically stain tissue microarrays containing hundreds of tissue cores within a single experiment. The system for IHC employed by HPA relies heavily on the standardization of protocols and automatization using machines, but the evaluation of the optimal titration for each antibody is performed manually before the antibody is approved for staining on the full set of tissues. Each stained tissue core is annotated with respect to immunohistochemical staining in tissues and cell types, and thereafter published as a high-resolution image on the web portal to be freely viewed by anyone.
In clinical practice, IHC is mainly used within pathology to aid physicians to evaluate tissue specimens with respect to healthy and or diseased states, to set diagnoses, and to define the molecular subtype of different types of cancer. A specific example where IHC is used diagnostically is when pathologists are presented with a metastatic tumor sample and the tissue origin of the primary tumor is unknown. In these cases, pathologists use a panel of different antibodies that target tissue-specific proteins, such as prostate-specific antigen for prostate cancer, and estrogen receptor for gynecological cancers, or cytokeratin 20 for gastrointestinal cancers (Gremel et al., 2014). Once a broad classification is made, additional tissue-specific antibodies are used to further pinpoint the origin of the primary tumor. This information is useful for choosing the best or most appropriate strategy for drug therapy and/or to locate the primary tumor for radiation therapy and/or surgery.
References
Molnar, C. and Gair, J. (2013) Antibodies. Chapter in Concepts in Biology, Published by B.C. Open Textbook Project. Available at: https://opentextbc.ca/biology/chapter/23-3-antibodies/
The Human Atlas Project. (2019) Methods. Available at: https://www.proteinatlas.org/learn/method
Uhlén M et al, 2015. Tissue-based map of the human proteome. Science
PubMed: 25613900 DOI: 10.1126/science.1260419
Thul PJ et al, 2017. A subcellular map of the human proteome. Science.
PubMed: 28495876 DOI: 10.1126/science.aal3321
Uhlen M et al, 2017. A pathology atlas of the human cancer transcriptome. Science.
PubMed: 28818916 DOI: 10.1126/science.aan2507
Ahern, K. and Rajagopal, I. (2019) Biochemistry Free and Easy. Published by Libretexts. Available at: https://bio.libretexts.org/Bookshelves/Biochemistry/Book%3A_Biochemistry_Free_and_Easy_(Ahern_and_Rajagopal)/09%3A_Techniques/9.04%3A_Gel_Exclusion_Chromatography.
Magdeldin, S. (2012) Gel Electrophoresis - Principles and Basics. Published by InTech under Creative Commons Attribution 3.0. Available at: https://pdfs.semanticscholar.org/4b93/70ac3946cec6e12c369679c4178a5ef38e61.pdf
Structural Biochemistry/Proteins/X-ray Crystallography. (2018, November 19). Wikibooks, The Free Textbook Project. Retrieved 15:40, August 17, 2019 from en.wikibooks.org/w/index.php?title=Structural_Biochemistry/Proteins/X-ray_Crystallography&oldid=3488057.
UCD: Biophysics 200A (2019) "NMR Spectroscopy vs X-ray Crystallography", Chapter published in Current Techniques in Biophysics. Published by Libretexts and available at: https://phys.libretexts.org/Courses/University_of_California_Davis/UCD%3A_Biophysics_200A_-_Current_Techniques_in_Biophysics/NMR_Spectroscopy_vs._X-ray_Crystallography
Wikipedia contributors. (2019, June 27). Protein purification. In Wikipedia, The Free Encyclopedia. Retrieved 23:32, July 28, 2019, from en.Wikipedia.org/w/index.php?title=Protein_purification&oldid=903657925
Wikipedia contributors. (2019, February 15). Fast protein liquid chromatography. In Wikipedia, The Free Encyclopedia. Retrieved 17:14, August 15, 2019, from en.Wikipedia.org/w/index.php?title=Fast_protein_liquid_chromatography&oldid=883530035
Wikipedia contributors. (2019, July 9). Protein mass spectrometry. In Wikipedia, The Free Encyclopedia. Retrieved 15:27, August 16, 2019, from en.Wikipedia.org/w/index.php?title=Protein_mass_spectrometry&oldid=905547289
Wikipedia contributors. (2019, July 8). Peptide synthesis. In Wikipedia, The Free Encyclopedia. Retrieved 06:13, August 17, 2019, from en.Wikipedia.org/w/index.php?title=Peptide_synthesis&oldid=905401648