20.5: CO2 uptake - C4 and CAM Pathways
- Page ID
- 41602
The source for the organization and some of the text derives from Sindayigaya and Longhini. https://www.peoi.org/Courses/Courses...chem/biochem18 CC - https://creativecommons.org/licenses...sa/3.0/deed.en
The C4 Pathway
Photorespiration, caused by the oxygenase activity of RuBisCo/Oxygenase and the ensuring glycolate salvage (C2) pathway, significantly diminishes the efficiency of photosynthesis in C3 plants. Rest assured, some plants have found a way around the problem by adding a few steps before RuBisCo. The altered pathway is called the C4 pathway and plants that use it are called C4 plants. The initial steps involve a temporary capture of CO2 in the form of HCO3-, into a 4C, not 3C sugar. Plants that use the C4 pathway include maize, sorghum, sugar cane, and many tropic plants. An overview of the pathways emphasizing the steps that precede RuBisCo is shown in Figure \(\PageIndex{1}\).
Experimental evidence shows that radiolabeled 14CO2 is captured into the 4C molecule, oxaloacetate (OAA), a citric acid cycle and gluconeogenic intermediate, through the enzyme phosphoenolpyruvate carboxylase, which uses HCO3- as a substrate. The reaction takes place in mesophyll cells. OAA can be reduced by NADPH using malate dehydrogenase, or converted to aspartate through a transamination (not shown in the figure). Malate moves in the bundle sheath cell where it is decarboxylated by the NADP-malic enzyme to pyruvate. Pyruvate can move back into the mesophyll cell and be converted to phosphoenolpyruvate (PEP) and then back to OAA by the enzymes pyruvate phosphate dikinase and PEP carboxylase. CO2 from the decarboxylation of malate is delivered as a substrate to RuBisCo.
Pyruvate dikinase is used in bacteria, protozoa, C4 plants, and another type, Crassulacean, discussed below. Its non-plant function is to produce ATP, in a fashion similar to pyruvate kinase. The net reaction is:
ATP + phosphate + pyruvate = AMP + PPi + H+ + phosphoenolpyruvate
Figure \(\PageIndex{2}\) shows a simplified mechanism for the reaction
The net reaction shows that in C4 plants, two molecules are phosphorylated by ATP. One is pyruvate and the other is inorganic phosphate (Pi). Hence the name dikinase. The reverse reaction of ATP synthesis occurs in bacteria and protozoan. There are two phosphorylated intermediates, an Enz-His-P, and an Enz-His-PPi. These served as "activated" phosphate carriers in the phosphotransfer reactions to pyruvate and to Pi, respectively. If PPi is not hydrolyzed to 2Pi, as illustrated in the top left of Figure 2 above, the reaction is fully reversible.
In C4 plants there are three reactions
- Pyr + E-His-P ↔ PEP + E-His
- E-His + ATP ↔ E-His-PP .AMP (. indicates a noncovalent interaction)
- E-His-PP .AMP + Pi ↔ E-His-P + AMP + PPi
When PPi is hydrolyzed, the net input of ATP to phosphorylated pyruvate is two ATP equivalents.
In the next step, PEP is carboxylated in a carbon capture reaction by PEP carboxylase, which as mentioned above used HCO3- as a substrate, not CO2 per se. PEP carboxylase also does not have a competing oxidase activity. The product is malate, which releases locally high "saturating" concentrations of CO2 in the bundle sheath cells, which significantly suppresses the oxygenase activity of RuBisCo.
Pyruvate phosphate dikinase undergoes a very large change in a conformational change in domain organization during the catalytic cycle. The two kinase activities are located at different sites in the enzyme. The phosphorylation of Pi occurs in the N-terminal domain, while the phosphorylation of pyruvate is in the C-terminal domain. The center domain that links to the N- and C-terminal domains by associated "tethers" is the site of the catalytic histidine involved in phosphotransfer. A swiveling of domains occurs to allow sequential phosphotransfers.
The Pfam domain structure for the protein is shown in Figure \(\PageIndex{3}\).

Figure \(\PageIndex{4}\) shows an interactive iCn3D model of an AlphaFold predicted model of chloroplast pyruvate, phosphate dikinase from Flaveria brownii (Brown's yellowtops) (Q39734)
Figure \(\PageIndex{4}\): AlphaFold predicted model of chloroplast pyruvate, phosphate dikinase from Flaveria brownii (Brown's yellowtops) (Q39734). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...VB6Aywuc2bLD38
The green represents the N-terminal NBD, the red is the central domain with the catalytic His (side chain shown as CPK-colored spheres and labeled), and the blue the PEP/Pyr PBD).
Figure \(\PageIndex{5}\) illustrates the conformation changes that occur in the central domain (yellow in this figure) and the N-terminal nucleotide-binding domain (NBD, green) in various ligand-bound states.
Panel a–c show movement of the central domain (yellow). In (a) it swivels to face the PBD domain (PDB 5JVL/C) and in (c) it faces the NBD domain (TbPPDK (PDB 2X0S)). In (b) it is in an intermediate position.
Panels d-f show the movement of the NBD (the three subdomains are depicted by three different greens). Pane (d) is the state without bound nucleotide (PDB 5JVJ/A), while panel (f) shows the fully closed, nucleotide-bound state (PDB 5JVL/A). Panel (e) shows a semi-closed, nucleotide-bound state (PDB 5JVL/C). Minges, A., Ciupka, D., Winkler, C. et al. Structural intermediates and directionality of the swiveling motion of Pyruvate Phosphate Dikinase. Sci Rep 7, 45389 (2017). https://doi.org/10.1038/srep45389. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/
Figure \(\PageIndex{6}\) shows interactions of bound substrates in different conformation states of PPDK.
Figure \(\PageIndex{6}\): Substrate binding sites of FtPPDK.
Panel (a) shows the semi-closed state of the PEP binding site (PDB 5JVL/A) with the catalytic H456 (yellow) pointing away from PEP.
Panel (b) shows the closed state of the PEP binding site (PDB 5JVL/C) showing interactions between PEP and surrounding residues, including the catalytic H456 (yellow).
Panel (c) shows the closed state of the nucleotide-binding site of 5JVL/D occupied with 2'-Br-dAppNHp, a nonhydrolyzable ATP analog. Interacting residues are highlighted. Minges, A et al. ibid.
After CO2 is delivered from malate in the bundle sheath cell using RuBisCo, the rest of the reactions are the same as in the C3 pathways.
Once CO2 is fixed into 3-phosphoglycerate in the bundle-sheath cells, the other reactions of the Calvin cycle take place exactly as described earlier. Overall the C4 pathways require more ATP. A molecule of PEP is required for each CO2 fixed in the C4 pathway which takes the equivalent of two ATPs. So for each CO2 fixed in the C4 pathway, it takes 5 ATPs compared to 3 ATPs in the C3 pathway. As mentioned above the affinity (estimated from the KM) of CO2 for RuBisCo decrease with increasing temperature, which decreases the energetic efficiency of carbon capture. At higher temperatures (28-30 C), the extra energy cost for the C4 pathway balances out with the extra energy cost for the C3 pathway at higher temperatures.
Carbonic Anhydrases
We first encountered carbonic anhydrase when we discussed its mechanism in Chapter x.xx. We'll now discuss its function and activity in the C4 pathway in some detail. Given that we are facing a climate crisis due to the increasing levels of CO2 in the atmosphere arising from the burning of fossil fuels, removing CO2 from the atmosphere, a process called for climate purposes carbon sequestration, and understanding the role of carbonic anhydrase in CO2 sequestration becomes even more important.
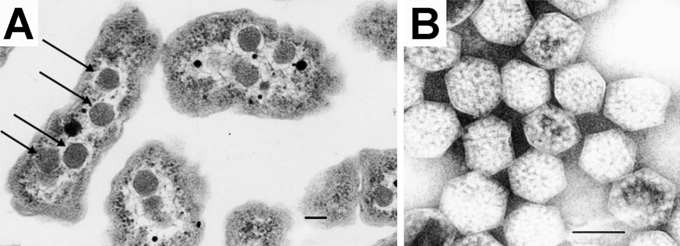
Plants have three different genes for carbonic anhydrase (α, β, and γ). Each can be differentially spliced so there many different isoforms of this protein in plants. They are most abundant in the chloroplast, cytoplasm, and in mitochondria and they have many additional roles outside of fixing CO2 in C4 (and also CAM) plants. In autotrophic (make their own "food") bacteria (such as cyanobacteria, also known as blue-green algae), there are no internal organelles. However, there are carboxysomes, which are protein-bounded vesicles (much like a bacteriophage head), which contain in their internal compartment not nucleic acid but RuBisCo and carbonic anhydrase. The carbonic anhydrase converts HCO3- to CO2 for reaction with RuBisCo. The carboxysome hence concentrates the CO2-producing and fixing enzymes for photosynthesis.
Figure \(\PageIndex{7}\):
 F
F
In C3 plants, CO2 (aq), that is dissolved CO2, is the actual substrate for RuBisCo so available HCO3- is converted to CO2 by carbonic anhydrase. In C4 and CAM plants, CO2 (aq) is first converted to bicarbonate by carbonic anhydrase. HCO3- (aq) is then used as the actual substrate for the "carbon fixation" step. Hence carbonic anhydrase has roles in C3, C4, and CAM plants.
All of the carbonic anhydrases have a Zn+2 at the active site. The alpha form, the most prominent in plants, was discovered in erythrocytes and is typically active as a monomer. It has one large 10-strand beta sheet surrounded by 7 alpha helices. The Zn ion is tetrahedrally coordinated by 3 histidine side chains and water. Gamma carbonic anhydrase is a trimer with three active sites at the interface between pairwise monomers with the Zn-coordinating histidine side chains from two different subunits.
Beta carbonic anhydrase in plants is typically an octamer of identical subunits. The Zn ion is coordinated by two cysteines, one histidine, and water. The monomer has 4 beta strands in a beta-sheet surrounded by alpha helices. An additional beta-strand is involved in monomer interactions. As the active site is in the interface of two subunits, the functional biological unit is the dimer, but a tetramer and even an octamer are typically formed. The substrate binding groups have a one-to-one correspondence with the functional groups in the alpha-carbonic anhydrase active site, with the corresponding residues being closely superimposable by a mirror plane. Therefore, despite differing folds, alpha- and beta-carbonic anhydrases have converged upon a very similar active site design and are likely to share a common mechanism.
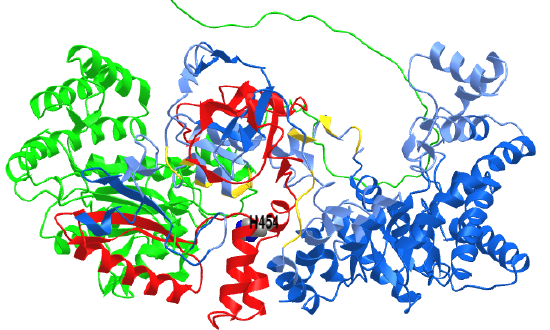
Figure \(\PageIndex{8}\) shows an interactive iCn3D model of beta-carbonic anhydrase from Pisum sativum (pea) with bound acetate (1EKJ).
Figure \(\PageIndex{8}\): Beta-carbonic anhydrase from Pisum sativum (pea) with bound acetate (1EKJ). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...VZTEJPy1hPxK27
For the sake of simplicity, only two subunits (A - gray and B - magenta) of the octamer are shown. Acetate, a proxy for HCO3-2, is shown in spacefill binding between the two subunits in CPK-colored spheres. The side chains involved in Zn2+ binding, Cys 160, His 220, and Cys 223 (unlabeled) are on the A chain. The amino acids that bind acetate are distributed between the A chain (Asp 162, Gly 224, Val 184) and the B chain (Gln 151, Phe 169, and Tyr 205) and are labeled. The same groups are involved in substrate binding in alpha carbonic anhydrase but in a mirror image orientation (but with the normal L-amino acids).
Beta carbonic anhydrase has high levels of expression in leaves and is found in chloroplasts, mitochondria, and the cytoplasm. Many plants have it in the cytoplasm and chloroplasts.
- β-carbonic anhydrase (βCA) in C3 plants: Most of βCA in leaves is in chloroplasts in mesophyll cells and may comprise up to 2% of leave protein. Yet studies have shown that you can delete the gene for it with minimal effect on the maximal rate of photosynthesis. However, plant development was affected so by interference the enzyme is probably most needed to produce HCO3- for biosynthesis.
- β-carbonic anhydrase (βCA) in C4 plants: Most of the βCA is found in the cytoplasm of mesophyll cells. There is catalyzes the first reaction of the C4 pathway, CO2 (aq) to HCO3- (aq). Mitochondrial (βCA) and γCA probably function to fix CO2 arising from oxidative respiration.
Crassulacean acid metabolism (CAM) pathway
Plants that encounter the chronic stress of low water availability have evolved yet another pathway to adapt to low water conditions. Stomata in C3 plants are open during the day to allow carbon capture from CO2, but they can close when water is limited. This obviously will inhibit plant growth. In the CAM pathway, the stomata stay open at night, allowing carbon capture at a time when water loss through the stomata would be lower. The incoming CO2 is fixed through carbonic anhydrase and then a series of several enzymes to form malic acid, which is transported for storage and use during the light in vacuoles. This CAM pathway is described in Figure \(\PageIndex{9}\).
The proteins and intermediates in the CAM pathways are ALMT9, aluminum-activated malate transporter; CA, carbonic anhydrase; MDH, malate dehydrogenase; OAA, oxaloacetate; ME, malic enzyme (NAD or NADP); P, phosphate; PEPC, phosphoenolpyruvate carboxylase; PEPCK, PEP carboxykinase; PPCK, PEPC kinase; PPDK, pyruvate, phosphate dikinase.
During the day, malic acid moves back into the cytoplasm where it is decarboxylated by malic enzyme and releases locally high CO2 concentrations for use by RuBisCo in the C3 cycle. In plants that use CAM, a series of other changes occur, including leaf structure and additional regulatory processes that coordinate metabolic genes.
Figure \(\PageIndex{10}\)s shows more details of the CAM cycle.
Thermodynamic comparison between the C3/C4 pathways and the Otto cycle
It is interesting to compare the thermodynamic efficiencies of plants and internal combustion engines that are governed by the thermodynamic Otto cycle. In the cycle, chemical energy in the form of gasoline and O2 are converted to thermal energy which is converted into mechanical energy. Both photosynthesis (the conversion of the energy of photons into chemical energy) have limited efficiencies.
- In internal combustion engines, power is limited in part by air uptake and by different efficiencies when running at non-constant speeds (like stop-and-go).
- In the C3 pathway, efficiency is limited by the oxygenase activity of RuBisCo/Oxygenase (given the much higher concentration of atmospheric O2 compared to CO2) and the ensuring photorespiration pathway.
- Water availability also plays a role as the net reaction of photosynthesis and glucose production, in simplified form, is 6CO2 + 6H2O → 6C(H2O6) so in high heat and low humidity the process efficiency decreases
Figure \(\PageIndex{11}\) shows a comparison of three stages, the storage component, the basic cycle, and a concentration mechanism, in photosynthesis and the internal combustion engine (ICE).
A concentrating mechanism in C4 plants and turbocharged cars provides concentrated CO2 and oxygen, respectively, to the core cycle (upper row). A storage mechanism in CAM plants allows carbon dioxide to be stored as malic acid at night and then passed to the Calvin cycle during the day, while a storage mechanism in hybrid electric vehicles (HEVs) allows energy to be stored in the battery during braking and then passed to the motor to power the drivetrain in parallel with the engine (bottom row).




_with_bound_acetate_(1EKJ).png?revision=1&size=bestfit&width=481&height=328)