17.1: Digestion, Mobilization, and Transport of Fats
- Page ID
- 15025
Introduction
In this chapter, we will discuss the breakdown of fats to produce ATP. Most of the available chemical energy stored in fats is in the form of highly reduced fatty acids. One form of fatty acid-containing lipids comes from our diet, which includes triacylglycerols (TAGs) and membrane lipids. Fatty acids, mostly in the form of TAGs, are moved in the circulation in the form of large lipid-carrying vesicles called lipoproteins. The lipids can be imported into cells for storage and energy use.
Another source of fatty acids comes from those synthesized within cells from the small molecule acetyl-CoA. Fatty acids are synthesized by an enzyme complex called fatty acid synthase. This enzyme is found most prevalently in adipose (fat) tissue and the liver. In addition, it is significantly expressed in the brain, lungs and mammary gland.
TAGs, stored in lipid droplets, are found in most cells. The major tissue used for TAG storage is adipose (fat) tissue, whose volume consists mostly of lipid droplet(s). Given the large mass of muscle tissue, there is also a considerable amount of TAGs stored as small lipid droplets in muscle cells. However, skeletal muscle cells don't synthesize fatty acids. They have the genes for fatty acid synthase but do not transcribe it into RNA so no enzyme is made. They can however import them for catabolism. Muscle TAGs can be oxidized for energy, especially during endurance excercise.
TAGs are also stored in the liver in lipid droplets. The liver also assembles lipoproteins, which are released by the liver. Excess TAGs are stored in the liver in various diseases including alcoholism and also in nonalcoholic fatty liver disease (NAFLD), which can progress into nonalcoholic steatohepatitis (NASH), a much worse disease.
There are two major forms of triacylglycerol-storing fat tissues, white adipose tissue (WAT) and brown adipose tissue (BAT). The more abundant WAT store triacylglycerols in one large lipid droplet in the cell and release fatty acid in processes controlled by the hormones insulin and epinephrine. This simple role can mask the fact that adipose tissue is a major player in the endocrine system and is involved in cell signaling and systematic control of metabolism. Adipose tissue releases the key hormones leptin and adipisin, which in analogy to the hormones and signaling agents released by immune cells (cytokines, lymphokines), can be called adipokines. They also secrete other adipokines including tumor necrosis factor α (TNF-α), adiponectin, and resistin.
In contrast, BAT is specialized not to store and release fatty acids. but rather to oxidize fatty acids in ways that maximize heat production, preventing hypothermia. They have multiple smaller lipid droplets, displaying a larger surface area for lipolysis, the hydrolytic cleavage of fatty acids from the TAGs. A particular mitochondrial protein, uncoupling protein 1 (UCP1), is expressed in brown but not white adipocytes, allowing a "futile" metabolic cycle leading to dissipation of heat instead of ATP synthesis. The relative abundance of white and brown adipocytes is critical in diseases like obesity and type 2 diabetes. BAT tissue is especially important in small animals (and in newborns) for thermoregulation. For smaller organisms, the surface area to volume ratio is greater than the ratio for larger animals, allowing more heat loss. The ratio of surface area AS per volume V for a sphere is given by:
\[\dfrac{\mathrm{SA}_{\text {sphere }}}{\mathrm{V}_{\text {sphere }}}=\dfrac{4 \pi \mathrm{r}^{2}}{\left(\dfrac{4}{3}\right) \pi \mathrm{r}^{3}} \nonumber \]
Let's assume an average large adipocyte is a sphere of diameter 100 uM. Compare this to a large sphere with a 100 times greater diameter (10,000 uM). The smaller sphere has a 1/100 of the diameter but a surface area/volume ratio 100 times greater than the large sphere.
An intermediate type of fat tissue consists of "bright" adipocytes. White adipocytes can be coaxed to differentiate into bright and brown cells, which could be an obesity treatment.
In this chapter section, we will follow the fate of fatty acids from dietary lipids which are cleaved from TAGs, loaded into chylomicrons, a lipoprotein assembled in the small intestine, secreted into the circulation, and taken up by the liver. The liver can store the incoming fatty acids in TAGs or release them back into the circulation in the form of another lipoprotein, very low-density lipoproteins (VLDL). Circulating VLDL can exchange lipids with other circulating lipoproteins. Lipoproteins deliver fatty acids to cells after interaction with the cell surface of target cells and either cleavage of TAGs by cell membrane-associated enzyme lipase, followed by fatty acid uptake, or by endocytosis of lipoproteins into the cells.
Lipoproteins
Before we look in more detail at the individual steps in lipids processing, let's look at the different lipoproteins, the large vesicular structures that allow the transport of fats, very insoluble molecules, in the circulation. Unlike normal liposomes or vesicles that have a lipid bilayer surrounding an interior aqueous compartment, lipoproteins have only a single monolayer of phospholipids encapsulating a nonaqueous interior filled with TAGs, cholesterol, and cholesterol esters. The protein part of the lipoprotein consists of one or several proteins bound on the outside of the particle. The proteins help solubilize the lipoprotein, confine its size, and prevent aggregation of the lipoproteins, which would be a health risk. The structure of a typical lipoprotein is shown in Figure \(\PageIndex{1}\).
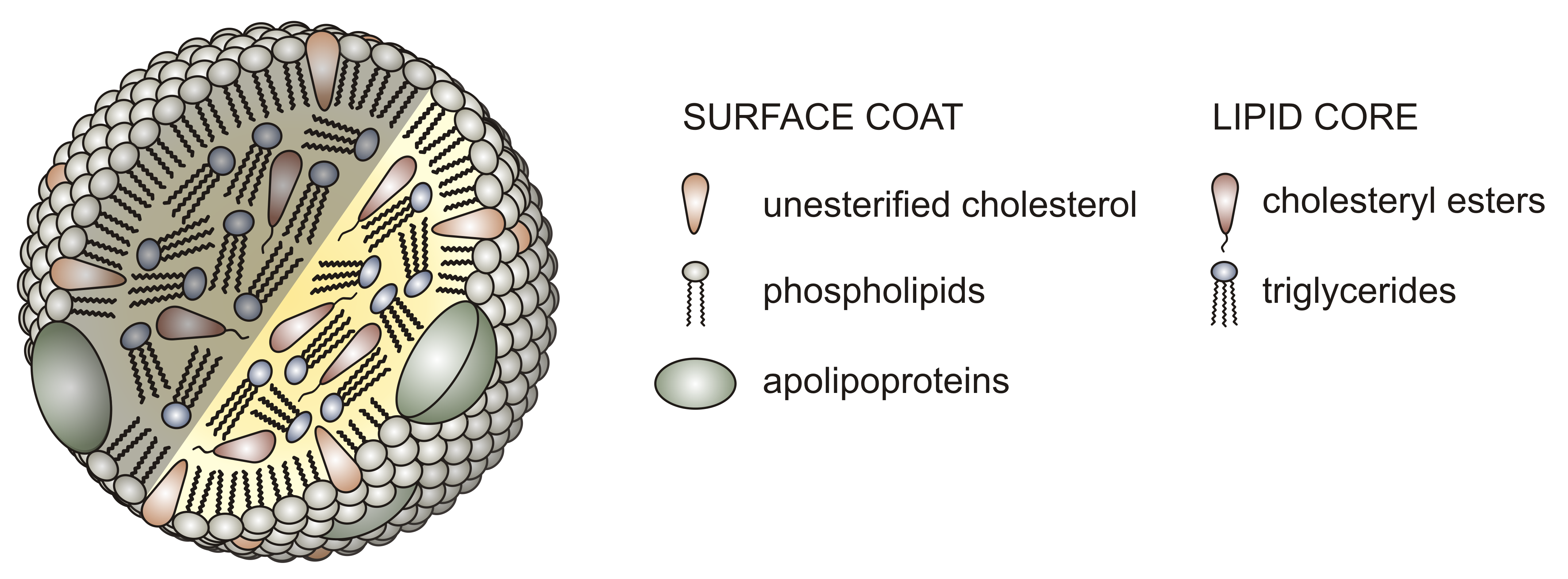
Lipoproteins are classified based on density. The lowest density chylomicrons are the largest with the most lipids (mostly TAGs) in their interior compartment. Very large density lipoproteins (VLDL), intermediate density (IDL), low density (LDL) and high density (HDL) have decreasing size, less encapsulated lipids, and increasing density. The relative sizes are shown in Figure \(\PageIndex{2}\).
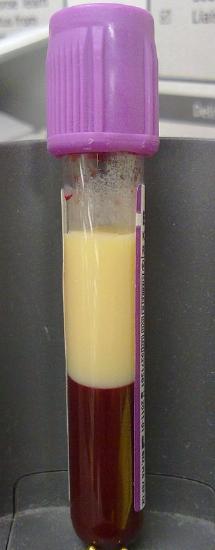
Lipoproteins (except chylomicrons) could be classified as nanoparticles, which typically vary in size from 1-100 nm. Larger lipoproteins as well as chylomicrons form emulsions in the blood, much as milk (also cloudy) is an emulsion of lipid/protein particles. The serum of people with high levels of lipids (hyperlipidemia) can look milky white, especially after eating foods rich in TAGs, when levels of chylomicrons are very high. Figure \(\PageIndex{3}\) shows the blood of a patient with hyperlipidemia after the addition of EDTA (which binds Ca2+ and prevents clotting) that has settled (without centrifugation). The milky white plasma on top (lower density) most likely has high concentrations of chylomicrons and/or LDL. The lower layer contains mostly red blood cells.
No x-ray structures of lipoproteins are available. However, a structure of a nascent HDL particle (3k2s) has been determined by small-angle neutron scattering. Figure \(\PageIndex{4}\) shows an interactive iCn3D model of it.
The major protein in HDL, a lipoprotein that protects against cardiovascular disease, is apolipoprotein A-I (apoA-I). Figure \(\PageIndex{4}\) shows that it adopts an antiparallel double superhelix as it wraps around the nascent HDL. The more hydrophobic surfaces of apo A-I are oriented inward allowing interactions with hydrophobic lipids in the core. It is probably prototypical for nascent lipoproteins. It will give you an idea of how proteins wrap around the outside of the particle. Mature lipoproteins are most likely spherical. This nascent HDL in the model contains 200 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholines (POPC) molecules, 20 cholesterol, and a single copy of apolipoprotein A-I (apoA-I).
Figure \(\PageIndex{5}\) below shows a cartoon image of VLDL, assembled from lipids synthesized/taken up by and released from the liver, and chylomicrons, assembled from dietary fats and released from enterocytes in the small intestine (size of the lipoproteins is not to scale). Note that VLDL has one copy of Apo B-100 while chylomicrons have one copy of Apo B-48.
All lipoproteins, except HDL, are members of the Beta-lipoprotein family as they contain an apo-B protein. The liver synthesizes apo B100, which becomes a permanent part of VLDL (i.e it is not exchangeable with other lipoproteins) and its metabolic derivatives so any lipoprotein containing apo B100 arose from the liver. Other proteins on lipoproteins are exchangeable. In contrast, enterocytes in the small intestine produce apo B-48 (48% the size of apo B100) so this protein marks the lipoproteins (chylomicrons and chylomicron remnants) that were assembled in the small intestine. Apo B100 has over 4500 amino acids and a molecular mass of 555K. The gene for the intestinal apo B48 is the same as for apo B100 except that it has a premature stop codon which leads to the shorter truncated apo B-48.
The apoproteins bind to specific receptors on cells which may allow the uptake of the lipoprotein. For example, the LDL receptors bind to the apo B-100 protein on a region removed from the apo B48 protein of chylomicrons. It also binds ApoE, which is found mostly predominately on HDL and VLDL but some are present in LDL. The LDL receptors have also been called the ApoB/ApoE receptor.
Over 90% of the apoB-containing particles in circulation are LDL. In addition, chylomicrons are present in circulation only after eating. Some apoproteins can act as cofactors and inhibitors for lipoprotein processing.
High concentrations of LDL are associated with increased cardiovascular risk. Chylomicron levels, given their transient and lower concentration levels, do not pose a health risk unless the enzyme required to remove fatty acids from them, lipoprotein lipase, is missing or defective, or if another apoprotein component, apo CII, which mediates the interaction with lipoprotein lipase, is missing. LDL-C (a term used to describe the total cholesterol in LDL particles which is routinely measured in clinical labs) can be lowed by a healthy diet centered around plant food). Drugs like statins, which decrease endogenous cholesterol synthesis, also remarkably lower LDL levels and decrease cardiovascular risk.
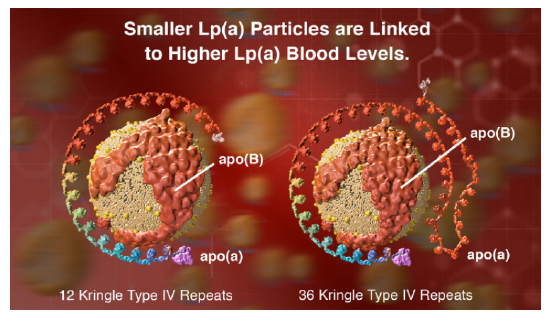
However, another protein, lipoprotein, also called Lp(a) or LP little a is an independent cardiovascular risk factor. Its blood concentration is regulated by genetics and not by diet. These particles contain, in addition to apo B100, apo (a), a protein that has a very unique repeating structure (up to 40 times) called a kringle, which is also found in some proteins involved in the blood coagulation system. People whose genes encode apo (a) with the fewest number of kringles express lots of that protein and their Lp(a) particles are smaller. This confers a greater cardiovascular risk compared to those expressing proteins with a large number of kringle domains. Figure \(\PageIndex{6}\) (from Amgen) shows models of Lp(a) with different numbers of repeating kringle (kinked) domains.
From a biochemical perspective, it is interesting to explore the differences in apolipoprotein binding to a single-leaflet encapsulated lipid nanoparticles compared to the interaction of peripheral and integral membrane proteins with intact bilayers (which we studied in Chapter 12.1). As mentioned above, the more nonpolar surfaces of apo AI in HDL are oriented inward toward the nonpolar lipid core. Presumably, apo B proteins in chylomicrons and LDL also wrap around the entire lipid surface.
The major organizing scaffolding protein of HDL is apoA-I (see iCn3D model above), It presumably plays a role similar to apoB in chylomicrons and LDL, but it is exchangeable. (Note: ApoA-I is also found in chylomicrons.) It is also a cofactor for the enzyme lecithin:cholesterol acyl transferase (LCAT), which effectively converts free cholesterol in the single bilayer into esterified cholesterol esters within HDL. In its apo-form, it also interacts with the cell surface transporter ATP-binding cassette A1 (ABCA1), which plays a role in the assembly of HDL particles. HDL also has apo C and apo E proteins, all of which are exchangeable.
It must be difficult to determine the structure of lipoproteins given their heterogeneity and size. The apoproteins have hydrophobic surfaces that promote self-association and aggregation. Apolipoproteins in the A, C, and E classes have repeating amphiphilic helices which imbed to some degree in the lipid particles. In addition, the proteins have a significant disorder and can adopt many bound conformations.
Figure \(\PageIndex{7}\) shows an interactive iCn3D model of the AlphaFold predicted structure of human ApoA (P06727)
The blue cartoon color represents high certainty in the AlphFold predicted structure while yellow to orange represents low certainty. The hydrophobic side chains are shown as sticks and help illustrate the amphiphilic nature of the structure.
Figure \(\PageIndex{8}\) shows an interactive iCn3D model of the AlphaFold predicted structure of human ApoE (P02649)
The blue cartoon color represents high certainty in the AlphFold predicted structure while yellow to orange represents low certainty. The hydrophobic side chains are shown as sticks and help illustrate the amphiphilic nature of the structure.
The exchangeable apolipoproteins have similar genetic sequences (four exons and three introns), as well as similar amino acid sequences. They have 11-mer amino acid tandem repeats and some (A-I, A-IV) have 22-mer tandem repeats. These repeats form amphiphilic helices as determined by sequence analysis. The first amino acid in the amphiphilic helix is often positively charged and a negative one is often found in the middle. Proline, a helix breaker, is often, but not always found between the helices.
Figure \(\PageIndex{9}\)s shows the primary sequence of apoA-I. An 11-mer repeat is shown in yellow highlight. The other highlighted stretches (different colors) are 22-mer repeats. Note that the repeats are not of identical sequences but rather of sequences that can form amphiphilic helices (i.e. secondary structure repeats).
The bottom part of Figure \(\PageIndex{9}\)shows a helical wheel projection (using Heliquest) of the red-highlighted 22-mer repeat. The arrow shows the hydrophobic moment with the arrowhead pointing to the more nonpolar face. The particular amphiphilic helix shown may or may not facilitate the binding of the bound conformation of the protein.
It follows that the relative areas of the hydrophilic and hydrophobic faces in the amphipathic helixes influence the lipid-associating properties of the exchangeable apolipoproteins. Another factor that might influence the lipid-binding ability of exchangeable apolipoproteins and which has not been studied in detail so far is the arrangement of tandem repeating amphipathic helixes with respect to one another.
Actual amphiphilic helices would bind to the membrane in a parallel fashion with the nonpolar face anchoring the protein to the lipid surface. Other experimental techniques are used to determine how a peptide or protein that can form amphiphilic helices interact with the lipid surface. These include site-directed mutagenesis studies coupled with spectroscopic (CD, fluorescence) and binding assay methods (using liposomes).
The properties of a membrane-bound amphiphilic helix are affected by the exact size and distribution of the polar/charged and nonpolar side chains. On binding, they sense or cause membrane curvature, interact with specific lipids, and stabilize specific membrane conformations (such as spherical for lipoproteins). Figure \(\PageIndex{10}\) shows how different proteins with amphiphilic membranes interact with membrane surfaces.
A key point to note is the large conformational changes that occur as the protein or parts of it go from the free, more disordered state, to the bound state with lipid-associating amphiphilic helices. The following proteins are depicted in the figure.
- The peroxisomal membrane protein Pex11 amphiphilic helix distorts the membrane;
- ARF1 is a small G protein in which only the GTP form localizes and binds through an amphiphilic helix to the membrane;
- The ALPS motif of the golgin GMAP-210 binds to only highly curved vesicles;
- The yeast transcriptional repressor Opi1 binds to the endoplasmic reticulum (ER) membrane in part through an amphiphilic helix;
- The heat shock protein Hsp12 has a long amphiphilic helix which helps stabilize the membrane;
- The extremely long amphiphilic helix of perilipin 4 coats lipid droplets and stabilizes even if there is a lack of phospholipids.
Dietary uptake and release into the circulation
Now how are the lipid nanoparticles assembled? We'll start with dietary lipids in the form of TAGs, glycerophospholipids, and cholesterol esters. The figure below shows key steps which are described in Figure \(\PageIndex{11}\)
Here are some key steps depicted in the figure:
- hydrolysis (lipolysis) of TAGs by pancreatic lipase, cholesterol esters (CE) by cholesterol esterase, and glycerophospholipids (GP)by phospholipase A2 in the lumen of the intestine. These enzymes interact at the interface of the lipid substrates and aqueous surroundings;
- the resulting products, which include free fatty acids (FA), 2-monoacylglycerol (MG), free cholesterol (FC), and lyso-glycerolphospholipids (lyso-GP), aggregate with the help of bile salts to form emulsions (like oil drops in water), which can be taken up by diffusion or possibly endocytosis when present in high amounts. Alternatively, membrane transporters (like FABPs and other proteins) can move them into the cell by facilitated diffusion;
- cytoplasmic transporters like fatty acid binding proteins move the lipolysis product to the ER where free fatty acids are reesterified. The enzymes involved include mono- and diacylglycerol acyltransferases (MGAT, DGAT) and sterol O-acyltransferase 2, also known as acyl-coenzyme A:cholesterol acyltransferase (ACAT-2). Multiple enzymes are involved in the resynthesis of glycerophospholipids);
- Apo B-48 is synthesized by ribosomes bound to the ER and interacts with a heterodimer of microsomal triglyceride transfer protein large subunit two (MTP) and protein disulfide isomerase (PDI). This facilitates the folding of apo B48 and loading of lipids using MTP into pre-chylomicrons;
- pre-chylomicron vesicles move to the Golgi with the help of Sar1b, a small G-protein (and GTPase) where the particle assembles to the full chylomicron, which is released from the cells as the mature large lipid nanoparticle.
An intriguing feature of lipases is that they work at the interface between the aqueous and nonaqueous (in this case lipid nanoparticle) environments. Let's briefly consider the mechanism of hydrolysis of TAGs by equine pancreatic mechanism. This enzyme utilizes the same mechanism we have seen earlier for the hydrolysis of a peptide bond by serine proteases. A catalytic triad of Asp 176, His 263 and Ser 152 as a nucleophilic catalyst is shown in the partial reaction displayed in Figure \(\PageIndex{12}\).
An acyl-Ser intermediate forms in step B (above), after the collapse of an oxyanion intermediate in step A, to form the product diacylglycerol. In the second half of the reaction (not shown completely), water, in a hydrolysis reaction, cleaves the acyl-Ser intermediate to reform the active enzyme as it releases the free fatty acid, R3CO2H. Other lipases also employ the same catalytic triad.
Figure \(\PageIndex{13}\) shows an expanded diagram showing the flow and fate of lipoproteins.
Chylomicrons interact with lipoprotein lipase (LPL), which also uses an Asp-His-Ser catalytic triad, to cleave fatty acid esters, which allows the delivery of free fatty acids to adipose cells. The adipocytes can also undergo de novo fatty acid synthesis. Fatty acids (FA) can also be produced by lipase-mediated lipolysis of stored TGs. Any of these free fatty acids (FA) in the adipocyte has two fates. They can be reesterified to glycerol to form TAGs (TG in the figure) or be exported from the cell and bind to a plasma carrier protein and transported to the liver, where it can be taken up by a variety of membrane proteins importers shown in the figure in yellow boxes. There, as in adipose cells, they can be reesterified to form TAG stores, which can then be packaged into VLDL particles for export. The fatty acid delivered (or synthesized) could also be used for ATP production through the citric acid cycle and oxidative phosphorylation.
VLDL in circulation can undergo lipolysis by lipoprotein lipase to produce fatty acids for uptake in "extrahepatic" tissue (bottom right of the diagram). As fats are removed from VLDL, their density increases as it forms IDL and LDL, which could be considered VLDL "remnants". VLDL is very enriched in TAGs, but after metabolic processing, the resulting LDL is depleted in TAGs and enriched in cholesterol/cholesterol esters. LDL (not shown in the above figure) can be taken up (endocytosed) by the liver and other cells after binding to LDL receptors, which recognize apo-B100 and other apoproteins. This allows the delivery of predominately cholesterol and cholesterol esters to tissues.
How do adipocytes and hepatocytes determine if free fatty acids should be esterifed for storage or released for energy use by other tissue? We'll discuss that in a subsequent section but the short answer is that in healthy fasting and exercise states, hormones (glucagon, epinephrine) will activate lipolysis in the liver and adipose cells, while in the fed state, insulin will promote storage of fatty acids as triacylglycerols.
Adipose cells don't assemble and release lipoproteins. Instead they release free fatty acids in the circulation which are carried by albumin, the major serum/plasma protein in the blood. The iCn3D Figure \(\PageIndex{14}\) shows an interactive iCn3D model of the complex of human serum albumin (HSA) binding seven 20:4Δ5,8,11,14 - arachidonic acids (1gnj ).
Given the multiple binding sites for fatty acids in albumin, it should come as no surprise that albumin also binds a host of small drugs, including medicinal drugs and toxins such as warfarin (blood thinner), diazepam, ibuprofen, indomethacin, and amantadine. These appear to bind preferentially at two major drug binding sites. This binding is probably helpful in delivering drugs through the circulation but potentially not useful if they aren't delivered to appropriate target tissue.
We discussed the structure of micelles which are spherical assemblies of single-chain amphiphiles that act as detergents. Oil from your clothes can enter the nonpolar interior of the detergent micelle and effectively solubilize the nonpolar molecule in the micelle, which are effectively nanoparticles with a diameter of 5-15 nm. You should hence not be surprised to discover that lipoproteins can also carry fat-soluble vitamins, steroid-like endocrine-disrupting substances, and drugs.
Lipoprotein lipase
The enzyme that breaks now TAGs in circulating chylomicrons and VLDL is lipoprotein lipase (LPL). It is a soluble protein secreted by adipocytes and muscle cells but is made by many cell types. It works at the luminal side of blood vessel endothelial cells and is recruited to that membrane surface by binding to the glycosylphosphatidylinositol-anchored high-density lipoprotein-binding protein 1 (GPIHBP1) as well as the proteoglycan heparan sulfate at the cell surface.
What is so interesting is that GPIHBP1 is only synthesized by endothelial cells. When lipoprotein lipase is secreted from cells, it binds to the extracellular matrix heparan sulfate but dissociates on the cleavage of heparan sulfate by heparinases. GPIHBP1 is highly acidic with an intrinsically disordered N-terminal domain containing a sulfated tyrosine and is highly enriched in glutamates and aspartates, which are often sequential in the sequence. Here is the single-letter sequence for amino acids 25-50 of the human version of GPIHBP1: EEEEEDEDHGPDDYDEEDEDEVEEEE. This sequence would have similar electrostatic and binding properties to the highly negatively charged heparan sulfate to which it also binds.
LPL also binds Ca2+ which stabilizes the active dimeric form of the protein. Its enzymatic activity is activated by apoC-II. Like pancreatic lipase, it employs a Ser-132, Asp-156, and His-241 triad in its hydrolytic action on TAGs/
Figure \(\PageIndex{15}\)l below shows an interactive iCn3D model of LPL in complex with GPIHBP1, shown in brown (6E7K). The calcium ion is shown (grey spacefill) as well as the catalytic triad (labeled, sticks, CPK colors). The highly negatively charged stretch of amino acids in GPIHBP1 was not present in the crystal structure.
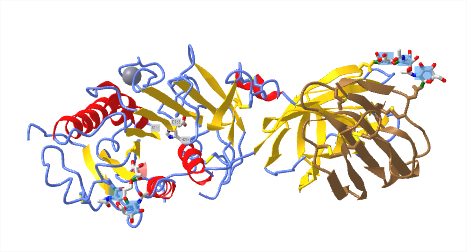
LDL: Receptor and Uptake
Lipoproteins are taken up into cells through receptor-mediated processes. Let's focus generically on the LDL receptor, the major carrier of cholesterol, given its role in cardiovascular disease. It is found in the cell membranes in most tissues. It is has many domain repeats, as illustrated in the Figure \(\PageIndex{16}\) calculated by SMART.
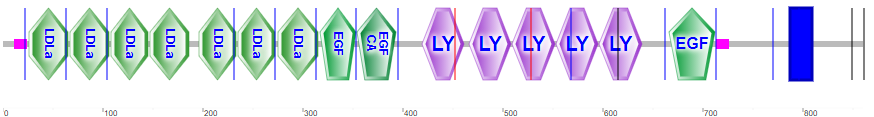
They include the N-terminal region cysteine-rich LDLa domains, which bind LDL, epidermal growth factor domains, LY (or LDLb) domains, and a transmembrane domain (blue rectangle).
Figure \(\PageIndex{17}\)s shows an interactive iCn3D model of the extracellular domain of the LDL receptor (1n7d).
Four tandem LY (LDLb) domains are shown in cyan, LDLa domains are shown in magenta and the EGF domain is shown in dark orange. Glycans are shown in symbolic nomenclature for glycans. Zoom into the structure to see the two disulfide bonds in each LDLa domain as well as the Ca2+ ions that stabilize the domains.
LDL binds its receptor at a broad binding interface with multiple LDLa domains. This may account for the fact the lipid nanoparticles with apo B100 or apo E can bind to it. The binding triggers a series of signaling events that lead to internalization by endocytosis of the receptor in pits coated with the protein clathrin. These eventually fuse with lysosomes where they are degraded and cholesterol delivered to the cell. The steps are described in Figure \(\PageIndex{18}\).
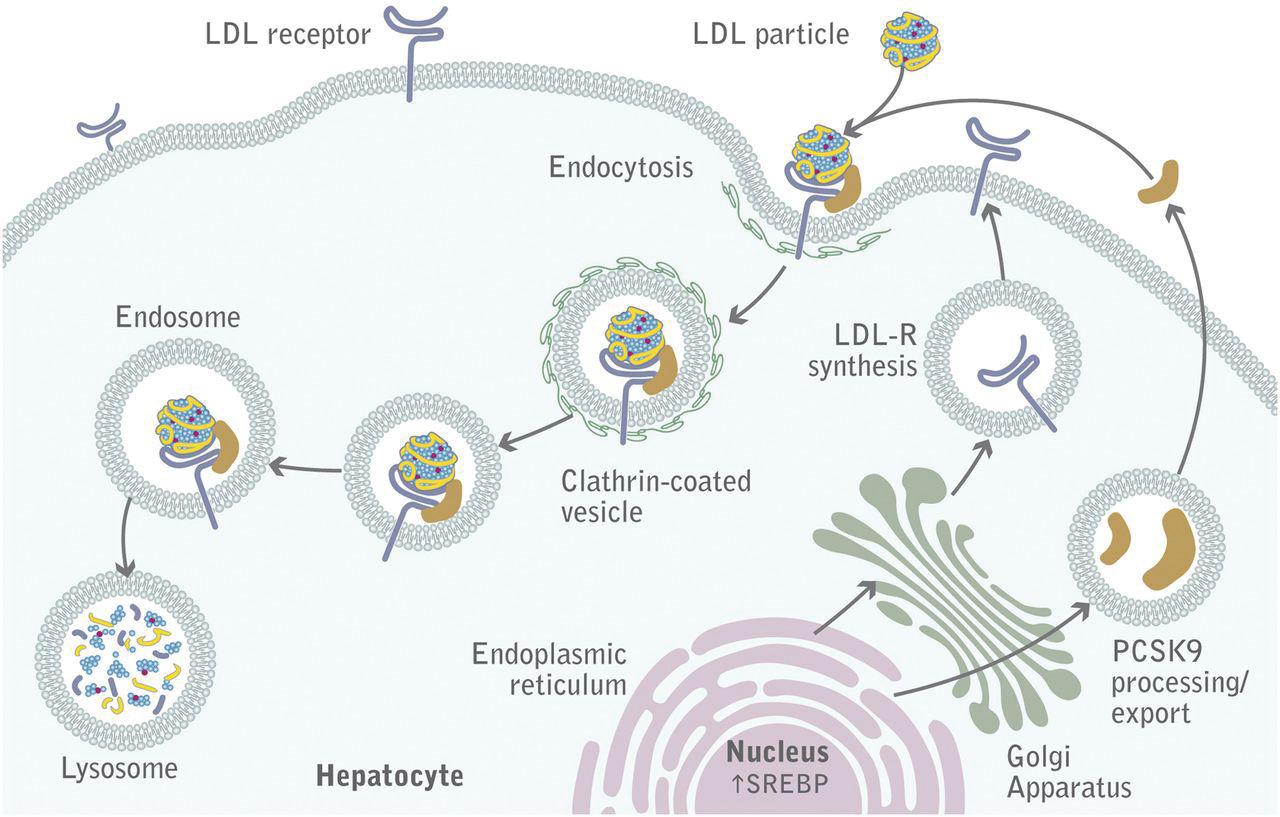
The LDL receptor survives lysosomal degradation and along with newly made receptors is delivered to the plasma membrane continually. A key protein, proprotein convertase subtilisin kexin type 9 (PCSK9), a serine protease secreted by the liver, promotes enzymatic degradation of the receptor and prevents its recycling to the membrane. It also binds to VLDLR and apolipoprotein E receptors and promotes their degradation as well. Its action reduces LDL clearance from the blood, increasing cardiovascular risk, so inhibitors of its action might be potent drugs to decrease circulating LDL.
The LDL receptor is just one member of the LDL receptor family. Other members of the family are illustrated in Figure \(\PageIndex{19}\).
These include the LDL receptor (abbreviated Ldlr in Figure \(\PageIndex{17}\), as well as the VLDL receptor (Vldlr), apolipoprotein E2 receptor (Apoe2), and LDL receptor-related proteins (Lrp)1-4. These also have a NPxY-motif (asterisk in the cytoplasmic domain) and a YWTD/β-propeller domain. Given the similarity in domain structure for the LDL family of receptors, the conformational flexibility of the apolipoproteins (at least free in solution), and similar structures for the exchangeable apolipoproteins, it shouldn't be surprising that the LDL receptor would interact with different classes of lipoproteins, albeit with different affinities.
As mentioned previously, apoE is found most abundantly on HDL and VLDL/chylomicrons and their remnants. It serves as a ligand that binds to members of the LDL receptors family (remember that LDL generally binds the LDL receptor through apo B100).
Apolipoprotein E has three major variants (alleles) named ε2, ε3, and ε4 (also called ApoE2, 3, and 4). ApoE3 is the most prevalent. ApoE4 is found in only 15% of people but more than 50% with Alzheimer's Disease (AD), so it's a risk factor for this disease. AD affects the brain, which also contains up to 30% of the cholesterol in the body, so aberrations to cholesterol transport and uptake in the brain are not unexpected in neurodegenerative diseases like AD.
ApoE is secreted by brain microglia (immune) cells and astrocytes (specialized glial cells). It assembles lipids into lipoproteins (HDL-like) and becomes the major vehicle for binding to and importing into neurons in a process initiated by the apoE receptor. The major apoE receptor for clearance of lipoproteins in the brain is sortilin (SorLA in Figure \(\PageIndex{17}\)).
AD is characterized by the accumulation of a toxic amyloid prion protein called amyloid beta (Aβ). It is derived from selective but abnormal proteolysis of the neural integral membrane protein amyloid precursor protein (APP). Aβ aggregates to form insoluble neurotoxic extracellular Aβ amyloid plaques. The process in normal and diseased cells is shown in Figure \(\PageIndex{20}\).
The figure shows normal (left) and aberrant processing of APP and the family of proteases (secretases) involved. While the LDLR doesn't appear to bind to APP or influence its proteolytic processing, it does bind Aβ. LRP1 is much bigger than LDLR, binds a multitude of ligands, and can be cleaved with the same enzymes as APP. Its expressed in the liver and especially in the brain and can regulate the removal of Aβ.
Immune cells in the brain, called microglial, remove Aβ plaques (which are extracellular) by phagocytosis. ApoE4 increases the inflammatory response (as measured by cytokine release) of the microglia (a good thing if the responses prevents infection or rids Aβ plaques) but also inhibits their ability to phagocytose the Aβ plaques and their metabolic activity.
An additional note: Having one allele ApoE4 allele appears to increase the risk of severe COVID-19 five times while being homozygous for E4 leads to a 17-fold increased risk of severe disease.
Scavenger Receptors
Patients with homozygous familial hypercholesterolemia (FH) have very high levels of LDL derived from defects in binding and update. Patients display fatty acids streaks under vessel endothelial cells which morph into calcified plaques and lesions filled with fat. Monocytes/macrophages, which migrate to sites of vascular injury, take up LDL and eventually differentiate into foam cells filled with lipids. Somehow, they have receptors that can bind and internalize LDL when the "normal" LDL receptor can't. Brown and Goldstein found that a specific chemical modification of LDL, acetylation, was necessary for the rapid uptake of "modified" LDL into macrophage receptors. These receptors are now called scavenger receptors (SRs).
There is a large family of scavenger receptors. It consists of classes A-J proteins that share functional but not sequence homology. They are found on macrophages and endothelium. They bind to and help remove "damage" signals including damage-associated molecular patterns (DAMPs) and chemical species chemically modified by reactive oxygen species. The ligands are often polyanions, end-stage glycans, and extracellular matrix proteins. One such example is oxidized-LDL (produced in vivo or by chemical oxidative modification with malondialdehyde), which binds to the same scavenger receptor, SR-A1, also called Macrophage scavenger receptor type I, as acetylated-LDL. Figure \(\PageIndex{21}\) shows the domain structure of the SR family.
SR-A1/MSR1 not only binds acetylated and oxidized LDL but also β-amyloid (42), heat shock proteins (43), and PAMPs from some bacteria and viruses.
It's very difficult, given the ever-increasing amount of "omic" data (genomic, proteomic, lipidomics, interactomics, metabolomics), for readers and authors alike, to conceptualize all of the possible combinations of interactions among biological molecules. For visual learners and perhaps everyone else, it's extremely useful to portray information on structures and interactions visually. An example using STRING, a database of known and predicted protein interactions, for the domain structure and protein:protein interactions of SR-A1/MSR1, is shown in Figure \(\PageIndex{22}\).
Note the interactions with apolipoproteins, apoB, apoE, and apoA1. The right hand side of the figure also shows interactions with collagen alpha-2(IV) chain (COL4A2), which is found in the extracellular matrix.
HDL metabolism: The Good Cholesterol
High levels of LDL (and Lp(a)) pose a cardiovascular risk. In contrast, high levels of HDL and apo A-I are cardioprotective. HDL is involved in "reverse" cholesterol transport as it is taken up by the liver and sent to the intestines for elimination from the body. We have shown earlier in Figure 2 that HDL exists as many variants, reflecting the assembly and remodeling of HDL by enzymes and lipid transfer proteins. Figure \(\PageIndex{23}\) shows the lifecycle of HDL.
Secreted apo A-1 accretes lipids in the circulation through the transport and delivery of phospholipids and cholesterol from cell membranes by the ATP-binding cassette transporters (ABC) A1 and G1. Another protein, the scavenger receptor BI (SR-BI), a polytopic integral membrane protein, is also involved. It acts as a receptor for a variety of "lipid" ligands including phospholipids, cholesterol esters, and phosphatidylserine (an outer membrane marker for cell apoptosis) as well as lipoproteins such as HDL.
Other proteins are involved as well in both the assembly but especially in the remodeling of HDL. The lipolytic enzyme lecithin-cholesterol acyltransferase (LCAT) removes a fatty acid from phospholipids and adds it to free cholesterol in the HDL to form cholesterol esters. Two major lipid transfer proteins, phospholipid transfer protein (PLTP) and cholesterol ester transfer protein (CETP), move lipids between HDLs and other lipoproteins. Cholesterol ester transport protein is made and secreted from the liver. It appears to exchange cholesterol esters from HDL for the return of TAGs from VLDL. Other enzymes (lipoprotein lipase and hepatic lipase are also involved in forming free fatty acids.
In the final step, HDL can deliver cholesterol ester and cholesterol to the liver through binding to liver scavenger receptor BI (SR-BI) mediated by apo A-I. The protein is expressed by the liver, adrenal gland, endothelial cells, macrophages, and many other tissues. It appears that HDL is not primarily taken up by the liver by endocytosis. In contrast, LDL is taken up by endocytosis mediated by the LDL receptor. SR-BI facilitates the transfer of cholesterol esters from bound HDL to the liver cell.
Unlike the widespread use of statins, which reduce LDL-C concentrations and clinically reduce cardiovascular disease risk, drugs (fibrates, niacin, inhibitors of cholesterol ester transfer protein - CETP), which raise HDL levels, don't seem to lead to significant decreases in cardiovascular risk. The cholesterol delivered in excess to macrophages can lead to the formation of foam cells under the endothelial layer. Foam cells are proinflammatory and convert in time to cholesterol plaques. In contrast, HDL-C does appear to have this direct atherogenic effect.




.png?revision=1)
.png?revision=1&size=bestfit&height=270)