15.2: Glycogenesis
- Page ID
- 15014
Introduction
The process of forming glycogen is called glycogenesis and it requires the activity of six enzymes as illustrated in Figure \(\PageIndex{1}\). We have already discussed several including hexokinase which phosphorylates the 6'-OH of glucose and phosphoglucomutase which converts glucose-6-phosphate to the glucose-1-phosphate isomer. In this section, we will discuss the remaining four enzymes and their role in glycogen biosynthesis. They are Glycogen Synthase, UDP-Glucose Pyrophosphorylase (preferred name UTP-glucose-1-phosphate uridylyltransferase), Glycogenin, and Glycogen Branching Enzyme.
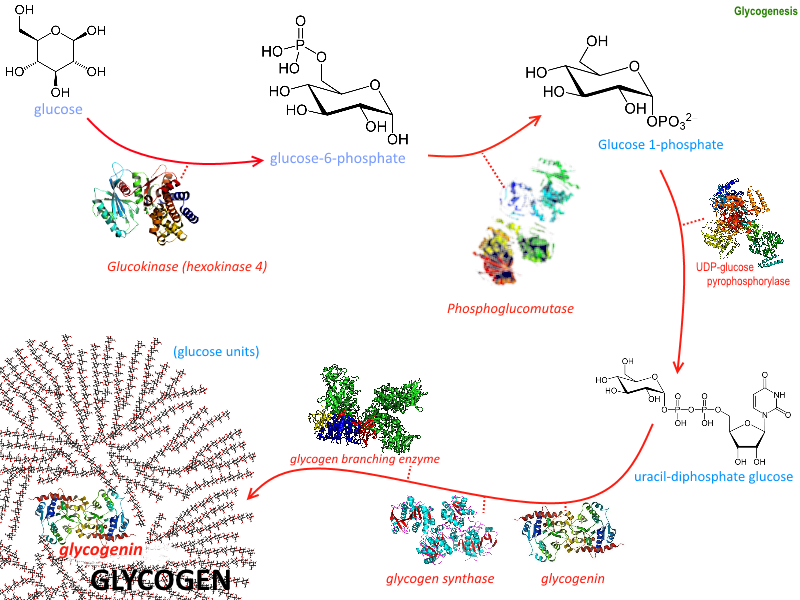
Given the importance of these enzymes in the synthesis of the second main energy storage molecule, we must probe the enzymes in detail.
Glycogen synthase (GS) is a key enzyme and its activity is highly regulated. In Chapter 15.1, we have already explored how insulin signaling upregulates the activity of this enzyme by inhibiting phosphorylation by GSK-3. Other effectors include the allosteric binding of glucose-6-phosphate, which also increases the activity of the GS. In a later section, we will also see that the hormone glucagon can also regulate the activity of the GS through protein kinase A (PKA) in a fashion that decreases glycogen synthesis and increases glycogen breakdown.
In the glycogenesis pathway, GS is responsible for building the majority of the main alpha 1 → 4 chain glucose acetal linkages. The GS does require a primer of 4 -6 glucose residues linked together by alpha 1 → 4 bonds to begin synthesis. Since GS can only form alpha 1 → 4 linkages in the main chain, it CANNOT create the alpha 1 --> 6 branches inherent to the core structure of glycogen.
To build the glycogen main chain, GS uses the glycogen primer and glucose that has been activated through covalent attachment to uridine diphosphate (UDP) at the 1-position. Upon completion of one round of synthesis, the 1 position of the incoming UDP-glucose is covalently attached to the 4-position of the nascent glycogen molecule, releasing the UDP as a leaving group.
\[\text { Glycogen }_{(n)}+\text { UDP-glucose } \rightarrow \text { Glycogen }_{(n+1)}+\text { UDP } \nonumber \]
UTP--glucose-1-phosphate uridylyltransferase (or UDP-Glucose Pyrophosphorylase)
The formation of the UDP-glucose required for the synthesis of the main chain of glycogen is mediated by UTP-glucose-1-phosphate uridylyltransferase (preferred name), which is also called UDP-glucose pyrophosphorylase (GalU or UGPase; EC 2.7.7.9). UGPase catalyzes the reversible reaction of glucose 1-phosphate and UTP into UDP-glucose and inorganic pyrophosphate (PPi) (Figure \(\PageIndex{2}\)). Enzymes of the UGPase family are ubiquitous and can be found in the tree of life.
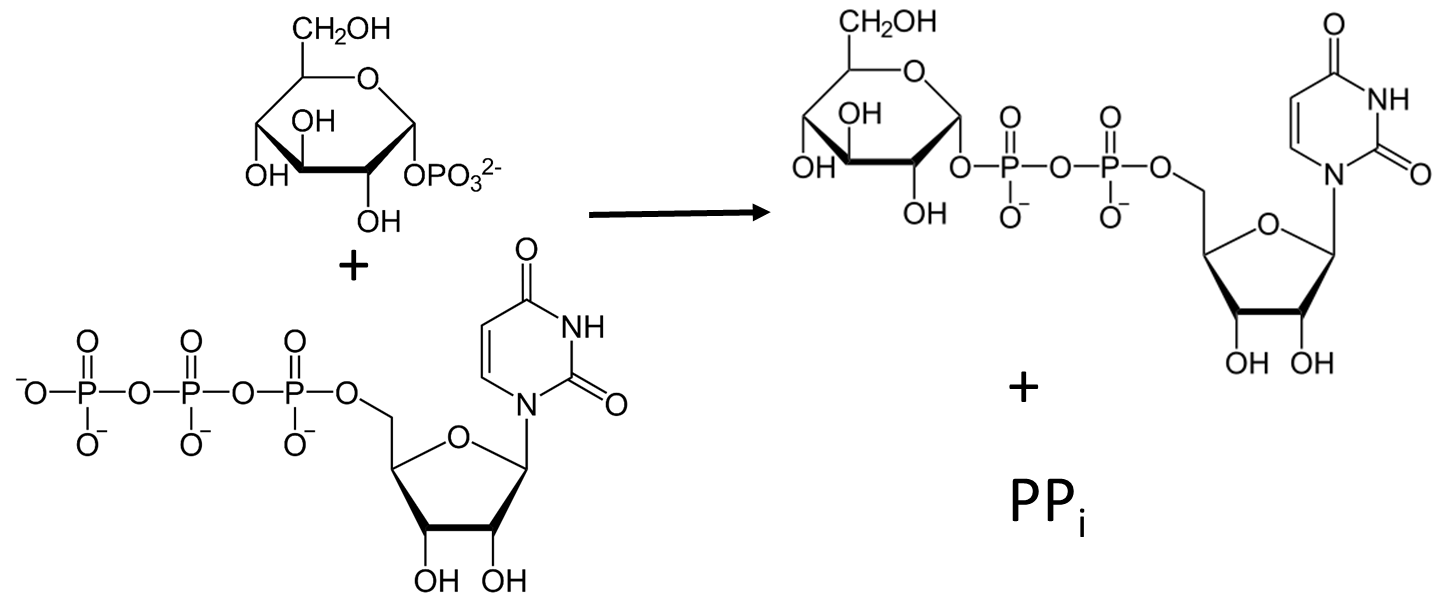
UDP-glucose is an activated form of glucose used in the synthesis of other glycans including sucrose, cellulose, start, and glycogen, and the glycan parts of glycoproteins, glycolipids, and proteoglycans. Hence it is a key metabolite, which gives more importance to understanding UGPase.
Like many other nucleotidyl transferases, UGPase requires divalent cations to promote the reaction (Figure \(\PageIndex{3}\)). In most cases, magnesium ions are employed. The reaction mechanism follows a sequential bi-bi-mechanism starting with the binding of UTP to the active site, in presence of a magnesium ion, followed by the binding of glucose 1-phosphate. The octahedral coordination sphere of the magnesium positions the substrates in the right way and enables the nucleophilic attack of glucose 1-phosphate on UTP. A lysine, an aspartate, and several water molecules within the active site help to stabilize the position of the substrates and cofactor for the proper nucleophilic attack of the phosphoryl oxygen of glucose 1-phosphate towards the α-phosphorous atom of UTP. Finally, PPi is released from the UGPase/Mg2+/UDP-glucose complex. UDP-Glucose then dissociates from the complex restoring the active site of the enzyme for another round of synthesis.
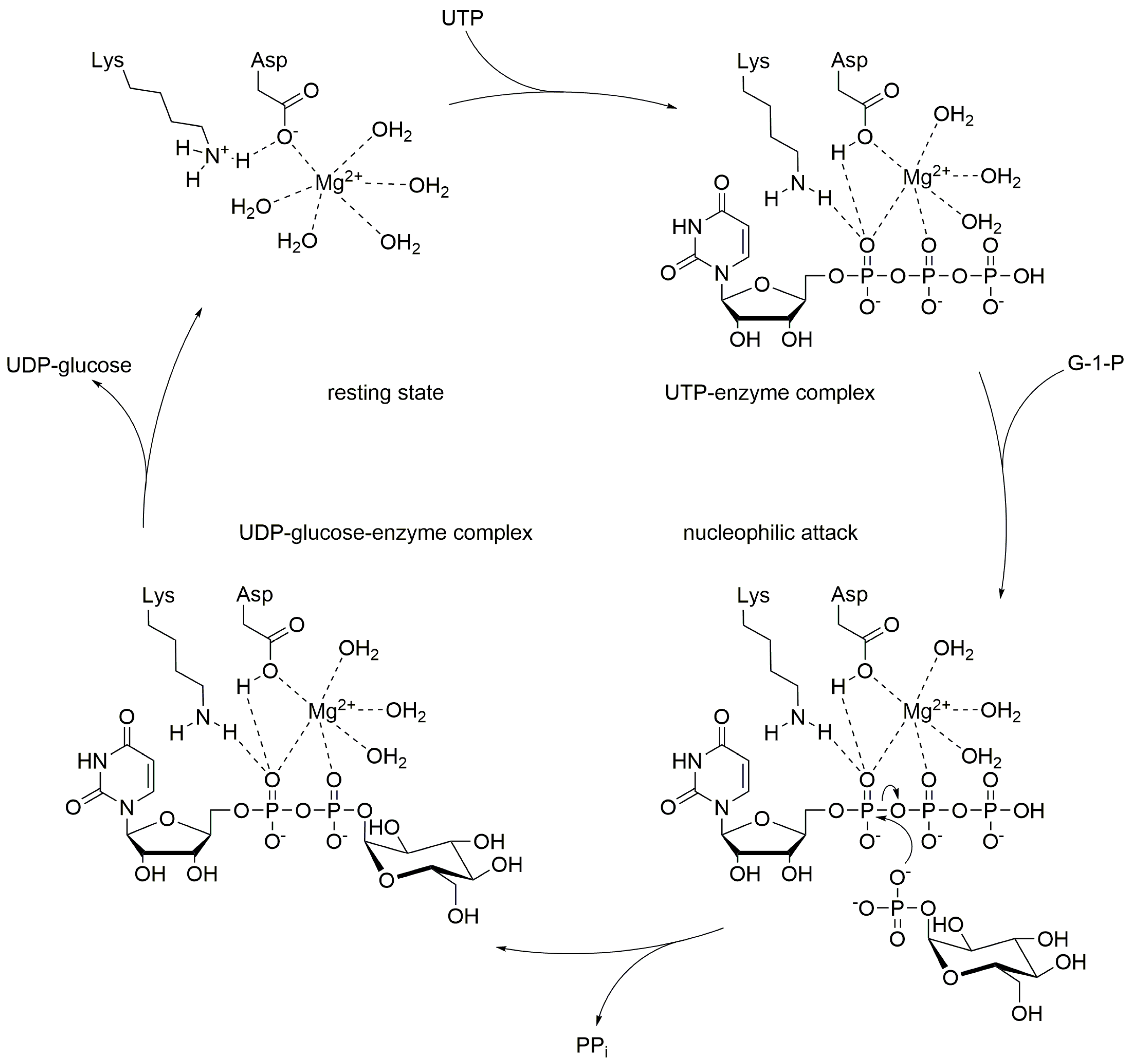
Figure \(\PageIndex{4}\) shows an interactive iCn3D model of the human UDP-glucose pyrophosphorylase tetramer (3R2W).
Figure \(\PageIndex{4}\): Human UDP-glucose pyrophosphorylase tetramer (3R2W). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/icn3d/share.html?9yq9STPGvq87Wnbt8
Substrate binding and active site residues are shown as CPK-colored sticks and labeled in one of the subunits. Three key loops whose correct positioning is required for catalysis are colored as follows:
- Latch Loop: 406-416, yellow, which contains Glu 412 (shown in spacefill)
- SB Loop: 275-282, red
- 309 Loop: 309-311, cyan, which contains Ser 309 (shown in spacefill).
Only four of the eight subunits are shown for clarity. Note the proximity of the interacting loops positioned between the brown and magenta subunits. The SB and 309 loops at that location are part of the brown subunit, while the yellow latch loop is part of the magenta subunit. The latch loop is between the SB and 309 loops. The cyan 309 loop hence is prevented from interacting with the substrate and with the movements of the SB and 309 loops needed for catalysis. In other species, it has been observed the SB loop moves down UDP-glucose on binding to the active site. The structure above is for the apo-enzyme without bound substrate, and hence the inactive or closed form of the protein.
Mutations in the 309 loop (S309N/S311R) still had 84% of normal activity. Glu412 in the latch loop is highly conserved in vertebrates (but not present in yeast). Mutations in Glu 412 didn't affect the formation of oligomers of the enzyme but did affect activity.
- replacing E412 with a short aspartate (E412D) significantly increase activity (176%)
- E412Q, which eliminates the charge while retaining the approximate size of the side chain showed just a marginal increase in activity (19%)
- E412K, which flips the charge and increases the length of the side chain decreased activity to 22%.
These mutations generally suggest that steric effects in the region of subunit interaction are most important in activity.
Glycogen Synthase
UDP-glucose is then utilized by glycogen synthase (GS) to extend the main chain of glycogen by one glucose residue. In this reaction, the 4’-OH group of the glycogen main chain attacks the anomeric carbon of UDP-glucose (Figure \(\PageIndex{5}\)). The UDP functional group serves as a good leaving group allowing for the formation of the alpha 1 --> 4 bond.
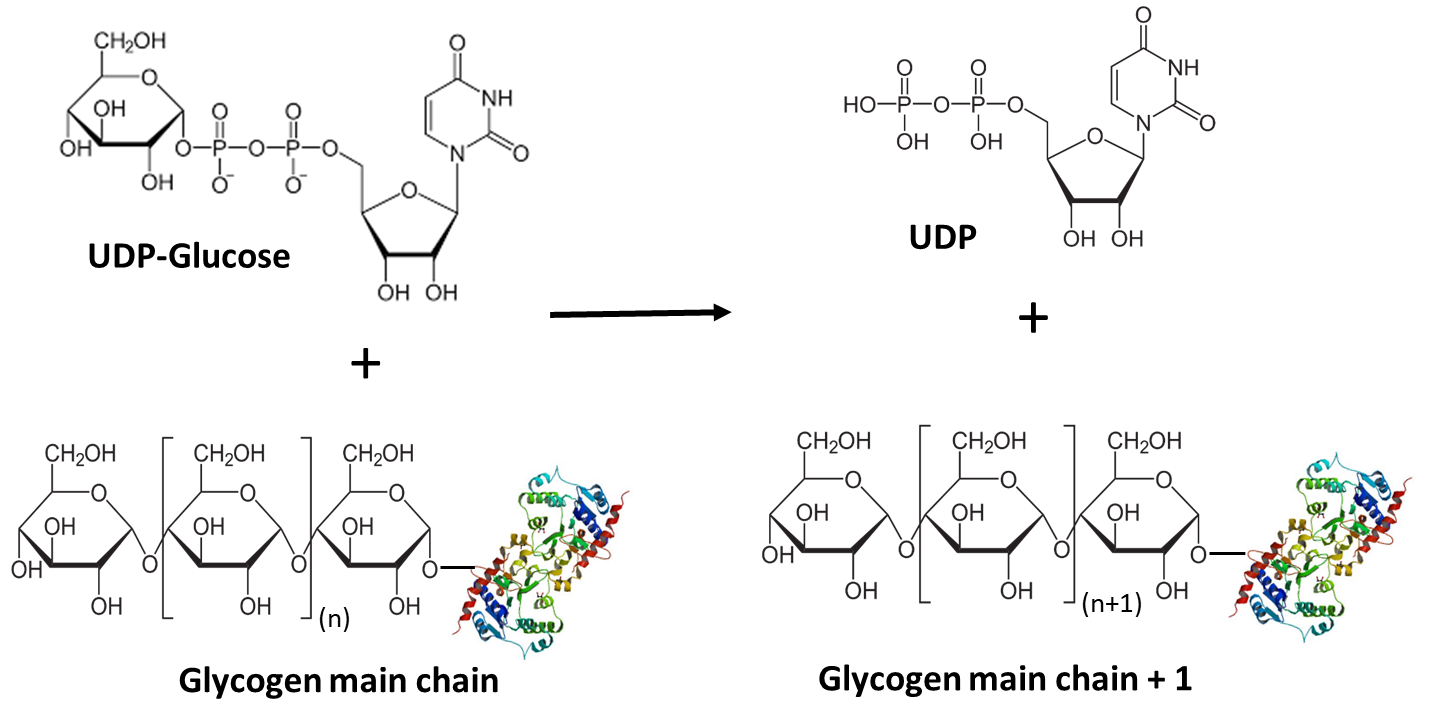
Glycogen synthase has two isoforms, GS1 expressed in tissue where glycogen is present (for example muscle) and GS2 expressed only in the liver. We will explore its mechanism more fully below.
Glycogenin
Previously, we mentioned that GS requires a glycogen primer of 4 – 6 glucose residues to begin adding new residues to the main chain. This primer is provided by the small docking protein, Glycogenin (GN or GYG). This protein is a homodimer that self-catalyzes its own glycosylation at amino acid Tyr-194. In this reaction, UDP-glucose is coordinated by a Mn2+ metal cofactor and critical aspartate residues (Figure \(\PageIndex{6}\)). The –OH group of Tyr-194 then mediates nucleophilic attack on the anomeric carbon of UDP-glucose. Thus, glycogenin is tethered to the reducing end of the glycogen molecule.
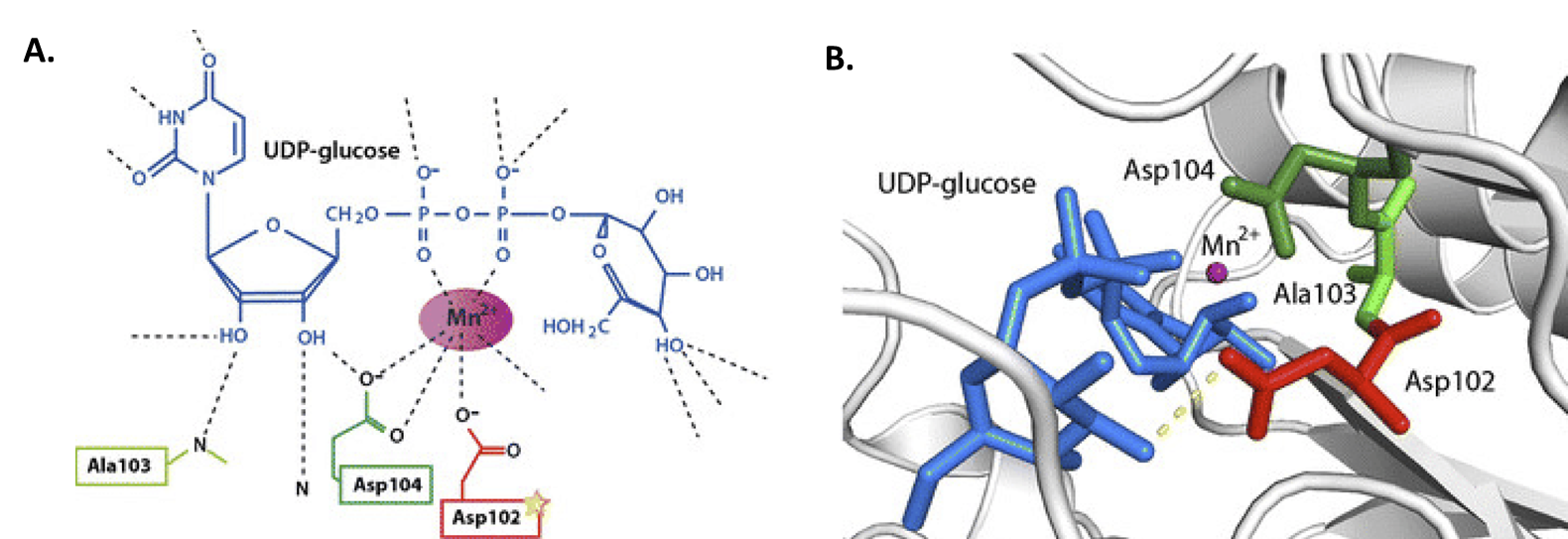
As with glycogen synthase, glycogenin (GN or GYG) has two isoforms, with GN2 (or GYG2) expressed mostly in the liver, pancreas, and heart.
Mechanisms for glycogen synthesis by glycogenin and glycogen synthase
How do glycogenin and glycogen synthase cooperate in the synthesis of glycogen? Structures of the complex of glycogenin-1 (GYG1), which seeds the molecule by starting glycogen synthesis by autoglucosylation, and glycogen synthase-1 (GS1 or GYS1), which extends the molecule, show allosteric transitions between three main states, the closed/inactive, partially open, and open active complex, as shown in Figure \(\PageIndex{7}\):
Figure \(\PageIndex{7}\): GYS1 chains A, B, C, and D are colored orange, turquoise, purple, and navy respectively. GYG1 globular domains and the GYG1-tail fragment are colored in gray shades for all chains. The left bottom structure shows the apo GYS1:GYG1 mobile complex. The bottom middle structure is the apo GYS1:GYG1 ordered complex. The bottom right structure is the +G6P GYG1:GYS1 complex. Fastman et al., 2022, Cell Reports 40, 111041 July 5, 2022, 2022. https://doi.org/10.1016/j.celrep.2022.111041. Creative Commons Attribution (CC BY 4.0)
It makes sense that both enzymes bind to each other and cooperate in the synthesis of glycogen. In the closed state, GS (GYS) is a tetramer, which like hemoglobin can be described as a T or closed/inactive state. In a slight difference, one of the GS (GYS) subunits appears to display an asymmetric conformation which leads to close interactions with GN (GYG) allowing the glycogen seed polymer on GN (GYG) to move to GS (GYS) for elongation (a partially open state). Multiple conformations of the complex have been resolved. Further conformational changes lead to the open/active state and a more open binding groove for GN (GYG).
Figure \(\PageIndex{8}\) shows an interactive iCn3D model of the Human glycogenin-1 and glycogen synthase-1 complex in the presence of glucose-6-phosphate (8CVX). Glucose-6-phosphate is an allosteric activator of glycogen synthase.
Figure \(\PageIndex{8}\): Human glycogenin-1 and glycogen synthase-1 complex in the presence of glucose-6-phosphate (8CVX). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...ivh4gZCupGcXZ7
The larger glycogen synthase subunits in the tetramer are shown in various colors while the glycogenin fragments bound to each monomer of GS (GYS) are shown in gray.
Figure \(\PageIndex{9}\) illustrate in greater detail the conformational changes induced on the binding glucose-6-phosphate to glycogen synthase.
Figure \(\PageIndex{9}\): G6P binding induces a conformational change across the GYS1 tetramer leading to an open conformation for all four active sites. Fastman et al, ibid.
Panel (A) shows a cartoon representation of the +G6P GYS1:GYG1 complex (left) with GYS1 and GYG1 chains colored as in the previous figure. G6P is shown as spheres colored by heteroatom. Blue and yellow lines and boxes (right) indicate relative positions for perpendicular views. A black dotted circle indicates the oligomeric interface between the CTD-loop region and tetramerization core domains. The CTD-loop, residues 484–488, forms cross-protomer interactions on the other side of the active site. Green dashed semicircles indicate open active sites.
Panel (B) shows G6P-binding site interactions. G6P (white) and proximal interacting residues are shown as sticks and colored by heteroatom. Polar interactions are indicated by yellow dotted lines.
Panel (C) shows changes at the G6P-binding site across conformations. A G6P-bound protomer is highlighted in orange with G6P shown as spheres. The adjacent active state protomer (chain C) is shown as a green cartoon. A basal state protomer (chain C) is modeled relative to the orange G6P-bound protomer and shown in yellow. Key changes to the G6P-sensing loop and a regulatory helix are highlighted by outline and non-transparent representation with arrows indicating the relevant motions. The rest of each chain is shown in a transparent cartoon representation.
We will return to the mechanism after exploring the last enzyme in the pathway.
Glycogen Branching Enzyme
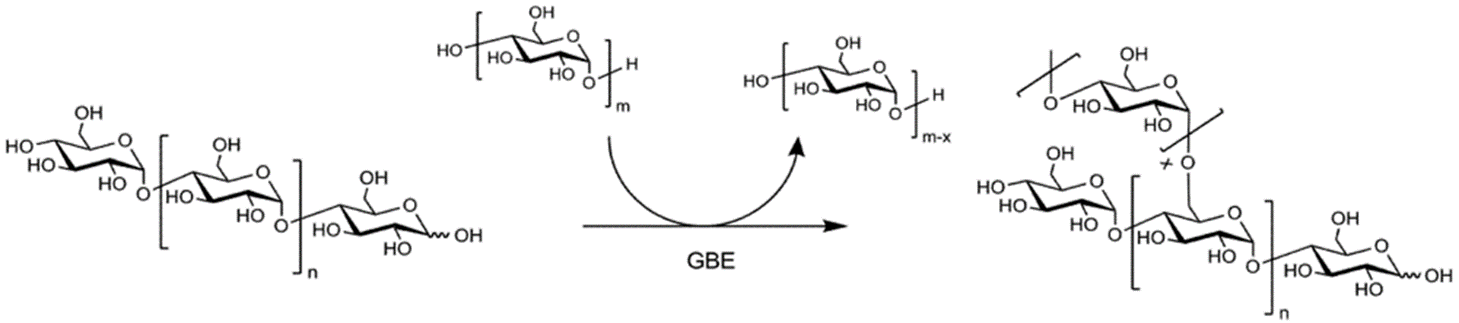
The final enzyme, the glycogen branching enzyme (GBE), catalyzes the hydrolytic cleavage of an α(1→4) glycosidic linkage and subsequent inter- or intra-chain transfer of the non-reducing terminal fragment to the C6 hydroxyl position of an α-glucan (Figure \(\PageIndex{10}\)). In this example, an inter-chain transfer is occurring. At the top of the scheme, above the arrow, you can see that the GBE enzyme transiently removes several glucose residues (usually around 7) from one linear glycogen chain and then attaches it as an alpha 1→ 6 branch to the other chain. In this process, an additional non-reducing end is created which can act as a primer site for Glycogen Phosphorylase (the main enzyme that breaks down glycogen). Thus, glucose residues can be released very quickly when needed.
Details of the structure and domain organization of the human glycogen branching enzyme are shown in Figure \(\PageIndex{11}\). The four domains include N1, CBM48, the catalytic domain, and the C-terminal domain.
Figure \(\PageIndex{11}\): Crystal structure of hGBE1. Froese et al. Human Molecular Genetics, Volume 24, Issue 20, 15 October 2015, Pages 5667–5676, https://doi.org/10.1093/hmg/ddv280. Creative Commons Attribution License (http://creativecommons.org/licenses/by/4.0/)
Panels (A and B) show orthogonal (perpendicular) views of hGBE1 showing the N-terminal helical segment (orange), CBM48 (pink), central catalytic domain (green), and C-terminal domain (blue). The catalytic triad Asp357-Glu412-Asp481 is shown as red sticks. Numbers refer to domain boundaries. N- and C-termini are labeled as grey spheres.
Panel (C) shows the superposition of branching enzyme structures from human (hGBE1, this study), O. sativa SBE1, and M. tuberculosis GBE, highlighting the conserved domain architecture and three regions of structural variation.
Panel (D) shows the domain organization of hGBE1, O. sativa SBE1, and M. tuberculosis GBE revealing differences in the N-terminus between prokaryotic and eukaryotic polypeptides. Prokaryotic GBEs contain two N-terminal carbohydrate-binding domains (N1, N2) whereas eukaryotes contain only one (CBM48) and replace the prokaryotic N1 domain with a helical extension.
Figure \(\PageIndex{12}\) shows an interactive iCn3D model of the human glycogen branching enzyme (GBE1)
Figure \(\PageIndex{12}\): human glycogen branching enzyme (GBE1). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...47aqw5qto8hkU7
The domains in the iCn3D module are colored similarly to that of the previous figure.
The enzyme core is similar to amylase with the conserved active site. Diseases of glycogen storage often result from mutations in the amylase core domain. For example, late-onset adult polyglucosan body disease (APBD) arises from a common mutation, Y329S. The effect of this mutation may arise from misfolding. A tetrapeptide, Leu-Thr-Lys-Glu, given to patients, increased activity twofold, probably by acting as a chaperone to facilitate proper folding.
Putting it all together: Glycogen Synthesis and Its Regulation
Figure \(\PageIndex{13}\) shows the steps involved in the addition of glucose from the donor UDP-glucose to glycogenin and through glycogen synthase to the growing glycogen polymer which becomes branched through the activity of the glycogen branching enzyme.
Figure \(\PageIndex{13}\): Summary of glycogen synthesis. Marr, L., Biswas, D., Daly, L.A. et al. Nat Commun 13, 3372 (2022). https://doi.org/10.1038/s41467-022-31109-6. Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/
The top reaction shows the step catalyzed by glycogenin (GN), while the bottom reactions are those catalyzed by glycogen synthase and the glycogen branching enzyme.
Regulation of glycogen synthesis occurs in part through the phosphorylation of key residues in both GS (GYS) and GN (GYG). Those sites are shown in Figure \(\PageIndex{14}\).
Figure \(\PageIndex{14}\): Domain and phosphorylation sites for glycogen synthase 1 (GS1) and glycogenin 1 (GN1). Marr et al, ibid
Panel c shows the domain architecture of human GS (top) and GN (bottom). Known in vivo phosphorylation sites of GS are shown in red and are labeled with residue number and classical nomenclature (in bold). GN tyrosine 195 that becomes auto-glucosylated and was mutated to phenylalanine (Y195F) in this study is indicated. Not to scale. Panel e shows the cartoon representation of GN WT and Y195F.
As we showed above, glycogen synthase is allosterically activated on the binding of glucose-6-phosphate, which can be thought to activate the enzyme by a T to R state change. The enzyme is inactivated by phosphorylation at multiple sites as shown above. Activation hence can also occur through dephosphorylation. Phosphorylation of a site can provide a binding site that leads to more phosphorylations. This can lead to a flexible "spike" of hyperphosphorylated residues forming from two of the monomers. Particularly important is pSer641 (site 3A) which interacts with a series of arginine residues in a regulatory helix of glycogen synthase. This arginine cluster has been called the arginine cradle. The interaction sites are illustrated in Figure \(\PageIndex{15}\).
Figure \(\PageIndex{15}\): The phosphoregulatory region of human GS. Marr et al, ibid
Panel a shows the human (Hs)GS-GN34 structure in ribbons (top left). The N- and C- terminal tails of one GS protomer (chain A) lie next to one another and move towards the adjacent protomer, meeting the N- and C-terminal tails from chain B. Arrows indicate a continuation of cryo-EM density (top right). Electron density (C1 symmetry) for phosphorylated S641 (pS641) interacting with R588 and R591 on the regulatory helices α22 (bottom left). Residues that are interacting with the N- and C-terminal tails that are mutated in this study are shown (bottom right).
Panel b shows a comparison of distances between regulatory helices of adjacent monomers of HsGS (reported here), low activity inhibited mimic (PDB ID 5SUL), basal state (PDB ID 3NAZ), and G6P activated (PDB ID 5SUK) yeast GS (yGS) crystal structures. Quoted distances were measured from Cα of Arg591 (chain A) and -Cα of Arg580 (chain B) of HsGS and corresponding yeast residues.
The strong electrostatic arginine-pSer interactions lock the tetramer into the inactive T state.
A cartoon model illustrating the regulation of glycogen synthase by phosphorylation/dephosphorylation and interconversion between T and R state is shown in Figure \(\PageIndex{16}\).
Figure \(\PageIndex{16}\): GS and GN cooperate to synthesize glycogen. Marr et al, ibid
The inhibition by phosphorylation can be relieved by binding the allosteric effector glucose-6-phosphate and does not require phosphatases,
References
1. Kumpf, A., Partzsch, A., Pollender, A., Bento, I., and Tischler, D. (2019) Two Homologous Enzymes of the GalU Family in Rhodococcus opacus 1CP-RoGalU1 and RoGalU2. Int. J. Mol. Sci. 20(22), 5809. https://doi.org/10.3390/ijms20225809




.png?revision=1&size=bestfit&width=391&height=344)
.png?revision=1&size=bestfit&width=461&height=423)
.png?revision=1&size=bestfit&width=557&height=345)