8.3: Nucleic Acids - Comparison of DNA and RNA
- Page ID
- 72660
Now that we have an understanding of the structures of DNA and the structures and various functions of RNA, we can now more fully explore how their chemical similarities and difference contribute to different functions.
Chemical modifications of DNA and RNA
Post-translation modifications of proteins alter their structural/functional properties. Likewise, intentional chemical modifications of nucleic acid bases alter both their structures and potentially their transcriptional and translational status. Figure \(\PageIndex{1}\) shows common modifications of bases in DNA.
Likewise, RNA is chemically modified. Figure \(\PageIndex{2}\) shows common modifications of bases in RNA. Methylation and subsequent hydroxylation to hydroxymethyl are common to both DNA and RNA. Methylation of DNA often represses the transcription of the DNA into RNA. Hence it has huge potential to alter gene transcription. Such changes to the DNA are called epigenetic modifications. These changes can be passed down to future generations as well and affect the phenotype of a cell. Histone proteins involved in DNA packing into nucleosomes can also be methylated and acetylated, altering the interaction of the DNA with the nucleosome core and further packing, again affecting transcription.
The chemical modification to RNA also can change the reading out of the genome. The epitranscriptome refers to the collective chemical modifications to RNA, and its understanding is part of new field called epitranscriptomics.
Mutations
Mutation can arise from the chemical modification of bases. Uracil in RNA is a demethylated form of thymine in DNA. In RNA, AU base pairs replace AT base pairs. Why the need for uracil in RNA? The question could be rephrased as to why the need for thymine, with its extra methyl group, in DNA. It's useful to think about the consequence of replacing a single H in a molecule with a -CH3. Take HOH, water, as an example. Our bodies are over 60% water. We drink liters of water of concentration 55 M each day. Yet if we drink 0.07 L of methanol, CH3OH, half of us would die! Let's probe some consequences of the U (no -CH3) and T (with -CH3) changes in DNA. It can get confusing but just remember that the normal base pairs in DNA are AT, but AU base pairs also form (they norm in RNA). The -CH3 substituent on thymine does not affect its base pairing.
a. Spontaneous deamination of cytosine in DNA
Why are we now discussing cytosine in DNA? One reason is that the most common mutation in DNA is a C to T replacement. One way that happens is through the spontaneous hydrolytic deamination of cytosine in DNA to uracil, which we have presumed to be found only in RNA. The mechanism for this deamination and subsequence conversion of a GC to an AT base pair is shown in Figure \(\PageIndex{3}\). The inset box shows a simplified mechanism for spontaneous deamination.
Hence a possible consequence of the deamination reaction is a GC to AT base pair mutation if the uracil in DNA is not removed before DNA replication. Fortunately, the enzyme uracil-DNA glycosylases can remove any uracils found in DNA, leaving an abasic site, which can be fixed with DNA repair enzymes.
We can now ask the question, why T and not U in DNA? Pretend you are a DNA repair enzyme and you see a UA base pair in DNA. How can you tell if the UA base pair is correct and intended to be there or if it should be a CG base pair that underwent deamination? The most common uracil-DNA glycosylases remove the uracil whether it is across from guanine, the correct base but which can not hydrogen bond with uracil (in the green oval in Figure \(\PageIndex{3}\)), or if is across from adenine, the wrong base (in red oval), which is present after a round of replication. Evolution has addressed this problem by adding a methyl group to uracil to form thymine and using that base, which forms a base pair with adenine. Now no decision on which base across from a uracil (guanine if the uracil arose from deamination) or across from a "uracil-like" thymine (adenine) is correct.
b. Other mutations
Since we are considering chemical modifications to DNA and mutations, it is appropriate to give a more expanded background on them. In addition to mutations caused by spontaneous hydrolytic deamination of cytosine, mutations can also arise through the addition of a wrong base during DNA replication, by chemical damages caused by radiation or chemical modifying agents. How many mistakes in replication are made? If you received a 99% on an examination, you would be ecstatic. That's not good enough for DNA replication. In Cell Biology by the Numbers, they calculate it this way. Assume the replication /repair is so good that it takes 108 replications to make a mistake (an error rate of 10-8/BP). Assume also there are 3 x 109 base pairs in the human genome. This leads to a mutation rate 10-100 mutations/genome/generation or about 0.1-1 mutations/genome/replication. Not bad!
Figure \(\PageIndex{4}\) shows how common point mutations might arise just randomly.
Chemical agents also can cause point mutations. Figure \(\PageIndex{5}\) shows point mutations arising from oxidative deaminations (not hydrolytic) by nitrous acid/nitrosamines and from alkylating agents.
Figure \(\PageIndex{6}\) shows a variety of alkylating agents with mutagenic potential.
Finally, large-scale changes in chromosome structure can also occur as shown in Figure \(\PageIndex{7}\), usually with profound consequences.
Why DNA and RNA - A chemical perspective
Asking a "why" question (like above) in the sciences is really not appropriate as such teleological questions are more philosophical or religious. Yet we will in this section, in part, to be in the company of Alexander Rich, who wrote a very cool article entitled "Why RNA and DNA have different Structures".
Given that RNA expresses catalytic activities and can carry genetic information (some viruses have ds and ss RNA as their genome), it has been suggested that early life might have been based on RNA. DNA would evolve later as a more secure carrier of genetic information. An inspection of the chemical properties of DNA, RNA, and proteins shows them to have attributes needed for their expressed function. Let's examine each for structural features that might be important for function.
a. Why does DNA lack a 2' OH group (found in RNA), which has been replaced with hydrogen? This required the evolutionary creation of a new enzyme, ribonucleotide reductase, to catalyze the replacement of the OH in a ribonucleotide monomer to form the deoxyribonucleotide form. One possible explanation is offered in the figure below. DNA, the main carrier of genetic information, must be an extremely stable molecule. An OH present on C'2 could act as a nucleophile and attack the proximal P in the phosphodiester bond, leading to a nucleophilic substitution reaction and potential cleavage of the link. RNA, an intermediary molecule, whose concentration (at least as mRNA) should rise and fall based on the need for a potential transcript, should be more labile to such hydrolysis. Figure \(\PageIndex{8}\) shows a possible reaction diagram for the internal cleavage of RNA. (The reaction would probably proceed with no actual intermediate, but just a transition state.
b. Why do both DNA and RNA contain a phosphodiester link between adjacent monomers instead of more "traditional" links such as carboxylic acid esters, amides, or anhydrides? One possible explanation is given below. Nucleophilic attack on the sp3 hybridized P in a phosphodiester is much more difficult than for a more open sp2 hybridized carboxylic acid derivative. In addition, the negative charge on the O in the phosphodiester link would decrease the likelihood of a nucleophilic attack. The negative charges on both strands in ds-DNA probably help keep the strands separated allowing the traditional base pairing and double-stranded helical structure observed. The cleavage of the phosphodiester link in DNA and a hypothetic ester link is shown in Figure \(\PageIndex{9}\). Again, the reaction of the phosphodiester shows a pentavalent intermediate, but most like the reaction proceeds directly from the transition state.
c. Why is DNA found as a repetitive double-stranded helix but RNA is usually found as a single-stranded molecule that can form complicated tertiary structures with some ds-RNA motifs?
Another reason for the absence of the 2' OH in DNA is that it allows the deoxyribose ring in DNA to pucker in just the right way to sterically allow extended ds-DNA helices (B type). The pucker in deoxyribose and ribose can be visualized by visualizing a single plane in the sugar ring defined by the ring atoms C1', O, and C4'. If a ring atom is pointing in the same direction as the C4'-C5' bond, the ring atom is defined as endo. If it is pointing in the opposite direction, it is defined as exo. In the most common form of double-stranded DNA, B-DNA, which is the iconic extended double helix you know so well, C2' is in the endo form. It can also adopt the C3' endo form, leading to the formation of another less common helix, a more open ds-A helix. In contrast, steric interference prevents ribose in RNA from adopting the 2'endo conformation, and allows only the 3'endo form, precluding the occurrences of extended ds-B-RNA helices but allowing more open, A-type helices.
Figure \(\PageIndex{10}\) shows another comparison between the A-RNA and B-DNA double helices and the C'3 and C'2 endo forms of the ribose
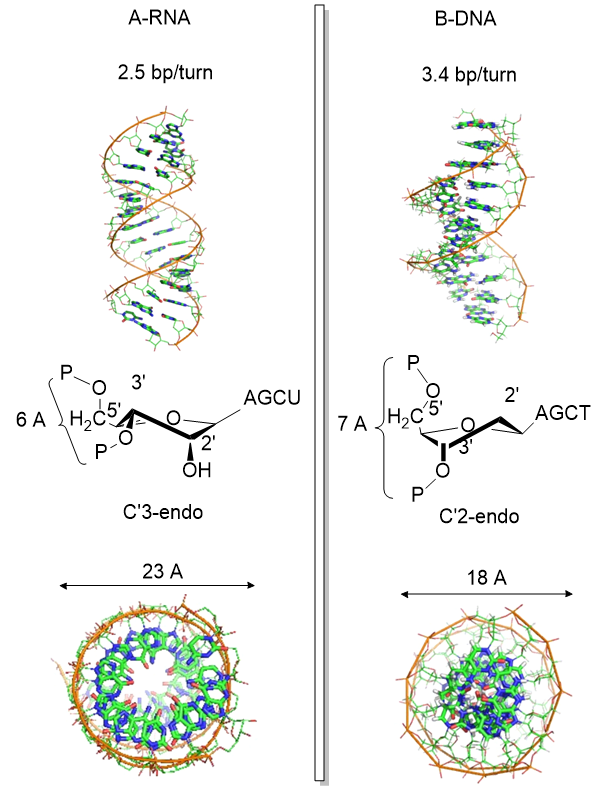
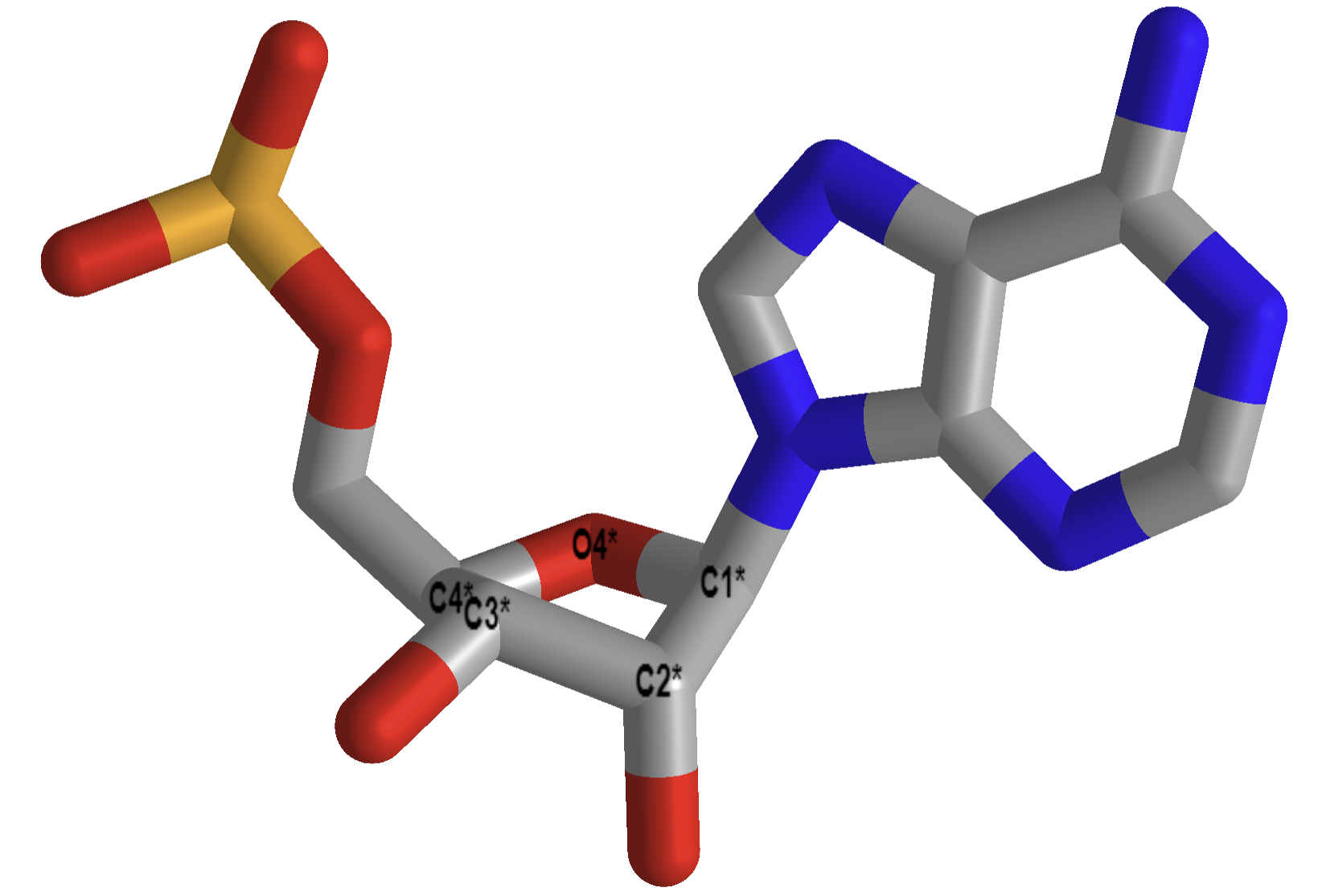
Figure \(\PageIndex{5}\) shows interactive iCn3D models of the pentoses in a strand of A-RNA (413D), double-stranded, left, and B-DNA (1BNA), double-stranded, right.
| C'3-endo ribose, A-RNA (413D, double stranded) | C'2 endo ribose, B-DNA (1BNA, double stranded) |
|
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...KPueqrBADczh26 |
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...BEn5nqsCQG2JH6 |
d. What about the molecular dynamics of A-RNA and B-DNA?
The information above suggests that the sugar ring of DNA is conformationally more flexible than the ribose ring of RNA. This can be inferred from the observation that dsDNA can adopt B and A forms, which requires a switch from the 2' endo in the B form to the 3'endo form in the A form. The smaller H on the 2'C would offer less steric interference with such flexibility. The rigidity in ribose is associated with a smaller 5'O to 3'O distance in RNA leading to a compression of the nucleotides into a helix with a smaller number of base pairs/turn.
The increased flexibility in DNA allows rotation around the C1'-N glycosidic bond connecting the deoxyribose and base in DNA, allowing different orientations of AT and GC base pairs with each other. The normal "anti" orientation allows "Watson-Crick" (WC) base pairing between AT and GC base pairs while the altered rotation allows "Hoogsteen" (Hoog) base pairs. The different orientations for an AT base pair are shown in Figure \(\PageIndex{11}\).
The Watson-Crick (WC) and Hoogsteen (HG) base pairs in B-DNA are in a dynamic equilibrium with the equilibrium greatly favoring the WC form as indicated by the arrows in the figure above. In a DNA:protein complex, the WC ↔ HG equilibrium can favor the WG form for AT and GC+ forms (in the latter, the C is protonated) when those base pairs are also involved in protein recognition. They can also occur more frequently in damaged DNA. In contrast, molecular dynamic studies show that the HG base pairs A-U and GC+ are strongly disfavored in ds A-RNA.
One type of DNA damage is methylation on N1-adenosine and N1-guanosine. This modification prevents normal Watson-Crick base pairing but for DNA, these modified bases can still engage in Hoogsteen base pairing, preserving the overall structure of dsDNA and its ability to stably carry genetic information. This same methylation occurs normally in post-transcriptional modified RNA. Hence, N1 adenosine and N1 guanosine methylation prevent any type of base pairing in the modified RNA. These properties make DNA a better carrier of molecular information and offer another way to regulate the structural and functional properties of RNA.
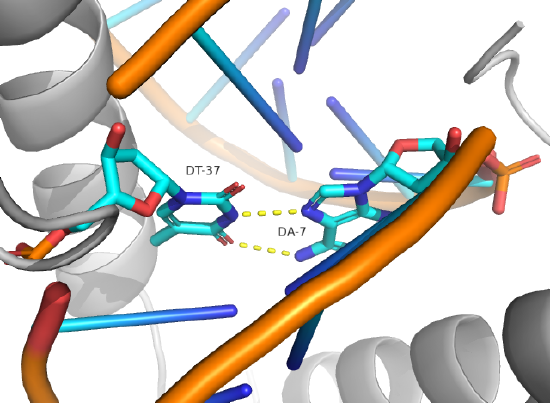
Hoogsteen base pairs can be found in distorted dsDNA structures (caused by protein:DNA interactions) but also in normal B-DNA. Figure \(\PageIndex{12}\) shows a Hoogsteen base pair between dA7 and dT37 in the MAT α 2 homeodomain:DNA complex (pdb 1K61). Note that the dA base in the Hoogsteen base pair is rotated syn (with respect to the deoxyribose ring) instead of the usual anti, allowing the Hoogsteen base pair.
A Structural Comparison
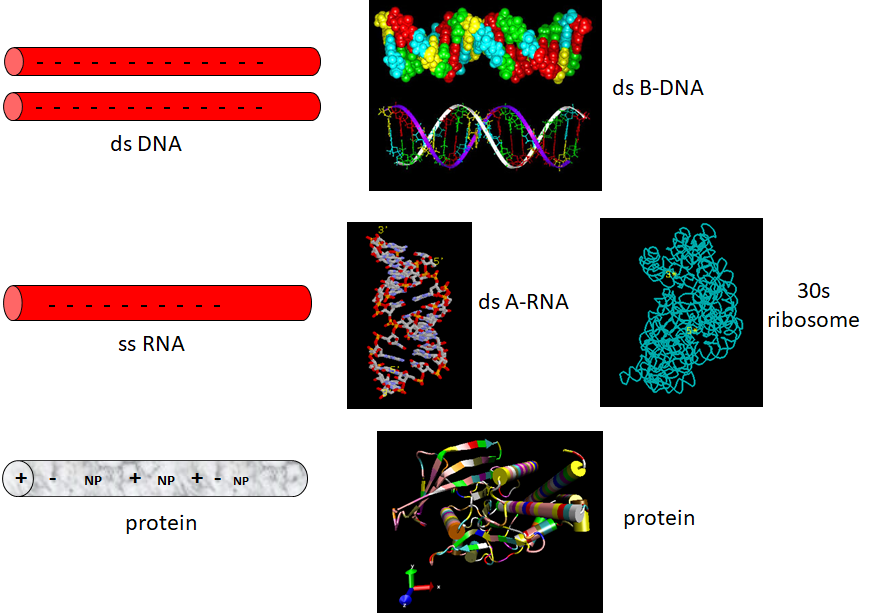
Now let's review the kinds of structures adopted by the 3 major macromolecules, DNA, RNA, and proteins. DNA predominately adopts the classic ds-BDNA structure, although this structure is wound around nucleosomes and "supercoiled" in cells since it must be packed into the nucleus. This extended helical form arises in part from the significant electrostatic repulsions of two strands of this polyanion (even in the presence of counter-ions). Given its high charge density, it is not surprising that it forms complexes with positive proteins and does not adopt complex tertiary structures. RNA, on the other hand, can not form long B-type double-stranded helices (due to steric constraints of the 2'OH and the resulting 3'endo ribose pucker). Rather it can adopt complex tertiary conformations (albeit with significant counter-ion binding to stabilize the structure) and in doing so can form regions of secondary structure (ds-A RNA) in the form of stem/hairpin forms. Proteins, with their combination of polar charged, polar uncharged, and nonpolar side chains have little electrostatic hindrance in the adoption of secondary and tertiary structures. RNA and proteins can both adopt tertiary structures with potential binding and catalytic sites, making them ideal catalysts for chemical reactions. RNA, given its 4 nucleotide alphabet, can also carry genetic information, making it an ideal candidate for the first evolved macromolecules enabling the development of life. Proteins with a great abundance of organic functionalities would eventually supplant RNA as a better choice for life's catalyst. DNA, with its greater stability, would supplant RNA as the choice for the main carrier of genetic information (Figure \(\PageIndex{13}\)):
A final note on the simplicity of the dsDNA structure. A mutation causing a single base pair change in DNA does not change the iconic ds-stranded DNA structure. If it did, DNA would not be a reliable molecule to store and read out the genetic blueprint. In contrast, a single mutation in the DNA leading to a single amino acid substitution may lead to a protein with altered structure and function. On one hand that could be deleterious or even fatal to the organism. On the other hand, the new protein structure might have new functionalities that allow adaptation to new environments or allow new types of reactions. Evolution would favor the latter.
References
Börner, R., Kowerko, D., Miserachs, H.G., Shaffer, M., and Sigel, R.K.O. (2016) Metal ion induced heterogeneity in RNA folding studied by smFRET. Coordination Chemistry Reviews 327 DOI: 10.1016/j.ccr.2016.06.002 Available at: https://www.researchgate.net/publication/303846502_Metal_ion_induced_heterogeneity_in_RNA_folding_studied_by_smFRET
Hardison, R. (2019) B-Form, A-Form, and Z-Form of DNA. Chapter in: R. Hardison’s Working with Molecular Genetics. Published by LibreTexts. Available at: https://bio.libretexts.org/Bookshelves/Genetics/Book%3A_Working_with_Molecular_Genetics_(Hardison)/Unit_I%3A_Genes%2C_Nucleic_Acids%2C_Genomes_and_Chromosomes/2%3A_Structures_of_Nucleic_Acids/2.5%3A_B-Form%2C_A-Form%2C_and_Z-Form_of_DNA
Lenglet, G., David-Cordonnier, M-H., (2010) DNA-destabilizing agents as an alternative approach for targeting DNA: Mechanisms of action and cellular consequences. Journal of Nucleic Acids 2010, Article ID: 290935, DOI: 10.4061/2010/290935 Available at: https://www.hindawi.com/journals/jna/2010/290935/
Mechanobiology Institute (2018) What are chromosomes and chromosome territories? Produced by the National University of Singapore. Available at: https://www.mechanobio.info/genome-regulation/what-are-chromosomes-and-chromosome-territories/
National Human Genome Research Institute (2019) The Human Genome Project. National Institutes of Health. Available at: https://www.genome.gov/human-genome-project
Wikipedia contributors. (2019, July 8). DNA. In Wikipedia, The Free Encyclopedia. Retrieved 02:41, July 22, 2019, from https://en.Wikipedia.org/w/index.php?title=DNA&oldid=905364161
Wikipedia contributors. (2019, July 22). Chromosome. In Wikipedia, The Free Encyclopedia. Retrieved 15:18, July 23, 2019, from en.Wikipedia.org/w/index.php?title=Chromosome&oldid=907355235
Wikilectures. Prokaryotic Chromosomes (2017) In MediaWiki, Available at: https://www.wikilectures.eu/w/Prokaryotic_Chromosomes
Wikipedia contributors. (2019, May 15). DNA supercoil. In Wikipedia, The Free Encyclopedia. Retrieved 19:40, July 25, 2019, from en.Wikipedia.org/w/index.php?title=DNA_supercoil&oldid=897160342
Wikipedia contributors. (2019, July 23). Histone. In Wikipedia, The Free Encyclopedia. Retrieved 16:19, July 26, 2019, from en.Wikipedia.org/w/index.php?title=Histone&oldid=907472227
Wikipedia contributors. (2019, July 17). Nucleosome. In Wikipedia, The Free Encyclopedia. Retrieved 17:17, July 26, 2019, from en.Wikipedia.org/w/index.php?title=Nucleosome&oldid=906654745
Wikipedia contributors. (2019, July 26). Human genome. In Wikipedia, The Free Encyclopedia. Retrieved 06:12, July 27, 2019, from en.Wikipedia.org/w/index.php?title=Human_genome&oldid=908031878
Wikipedia contributors. (2019, July 19). Gene structure. In Wikipedia, The Free Encyclopedia. Retrieved 06:16, July 27, 2019, from en.Wikipedia.org/w/index.php?title=Gene_structure&oldid=906938498




.png?revision=1)