5.5: Protein Interactions Modulated by Chemical Energy- Actin, Myosin, and Molecular Motors
- Page ID
- 14943
9/21/21: The material below is derived from (shortened and summarized): Overview of the mechanism of cytoskeletal motors based on structure. Kato et al. Biophys Rev. 2018 Apr; 10(2): 571–581. Published online 2017 Dec 12. doi: 10.1007/s12551-017-0368-1. Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0).
Muscle Contraction
The biochemistry of muscle structure and contraction involves many proteins, including myosin heavy and light chains, actin, three types of troponins, tropomyosin, and other regulatory proteins. These form complexes which engage in cyclic conformation changes leading to muscle contraction in a process that requires chemical energy in the form of ATP hydrolysis. In the process, chemical energy is converted to mechanical energy. Those who view biochemistry through a more chemical lens might find the biology somewhat confusing. We'll provide a short introduction before looking at the proteins in more detail.
First let's start with a simplified version of the structure of myosin and actin to the basic repeating structural unit of the muscle, the sarcomere, which is shown in panel A of Figure \(\PageIndex{1}\). Myosin forms a function dimer of two intertwined heavy chains (blue), each associated with two different light chains. These are named the essential light chain (red) and the regulatory light chain (yellow). Overall the functional dimer of myosin is a hetero 6-mer.
The heavy chain of myosin consists of two major domains and other regions as shown in panel A of Figure \(\PageIndex{1}\). It contains a
- motor (head) domain (blue ellipse), which binds and cleaves ATP (i.e. it is an ATPase) and which also binds actin;
- an alpha-helical tail domain that binds the tail domain of another myosin heavy chain to form the functional a coiled-coil structure in the hetero 6-mer.
- an α-helical lever arm, which binds the two light chains.
In the muscle sarcomere, many hetero 6-mers aggregate through their coiled-coil tails to form the large blue structure shown in panel B of Figure \(\PageIndex{1}\). The protruding heads that radiate away from the filament axis interact with actin filaments. This is shown in panel B of Figure \(\PageIndex{1}\). The actin filaments are shown as two intertwined actin strands (small gray circles).
Note also that in panel B, the myosin thick filaments appear to be bipolar as their motor head domain are pointed in opposite direction (antiparallel) along the center of the thick filament that has no heads in the center. The central region without motor domains has aggregated hetero 6-mers pointed in opposite directions. However, the rest are packed in a parallel fashion and extend the rest of the way.
The head domain of the myosin heavy chain can be removed, along with the associated light chains, from the rest of the using selective proteolysis to form the S1 (head), or S1 and part of the myosin light chain (called heavy meromyosin -HMM). This has enabled studies of these molecules under simpler conditions. The myosin heavy chain fragments are shown in Figure \(\PageIndex{2}\).
Going in the direction from the rod domain to the motor domain, the myosin heavy chain has a long alpha-helical lever arm which binds the two myosin light chains (the regulatory light chain (yellow above) and the essential light chain (green above). This is followed by the converter domain and finally the motor domain, which binds ATP.
The actin thin filament consists of two intertwined antiparallel strands of many G-actin (globular) monomers (1atn, molecular weight of 42K), which self-assemble in a process requiring ATP hydrolysis, to form the F (filamentous) actin structure. The structure of G-actin and the overall arrangement of the asymmetric monomers in the F filament are shown in Figure \(\PageIndex{3}\).
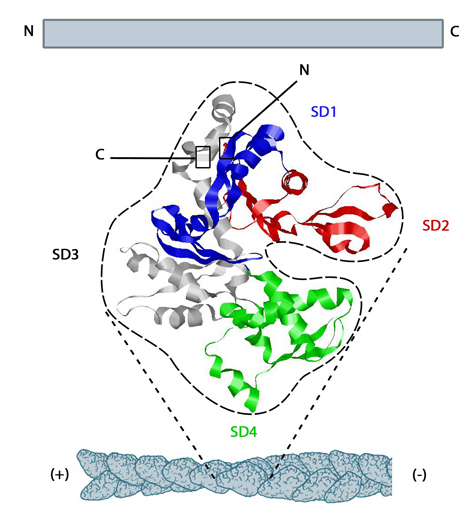
Figure \(\PageIndex{3}\): The structure of monomeric actin in the F-actin thin filament. https://www.mechanobio.info/cytoskel...ilaments-grow/. Creative Commons Attribution-NonCommercial 4.0 International License.
It would be nice if the structure of the sarcomere were as simple as illustrated in Figure \(\PageIndex{1}\). Many more proteins are involved which regulate its contraction. Let's add some more proteins (troponin subunits T, C and I as well as tropomyosin) to get a deeper understanding of the structure as shown in the cartoon representation shown in Figure \(\PageIndex{4}\).
Figure \(\PageIndex{4}\): Modified from de Tombe PP et al. Global Cardiology Science and Practice 2016:21
http://dx.doi.org/10.21542/gcsp.2016.21. Creative Commons Attribution 4.0 International License.
The trimeric troponin complex consists of
- troponin-C which binds Ca2+ ions
- troponin-I which inhibits contraction
- troponsin-T which attaches the trimeric troponin to the thin actin filament.
Ca2+ ions released from stores in the sarcoplasmic reticulum (the muscle equivalent of the endoplasmic reticulum) on neural stimulation of muscle cells by the neurotransmitter acetylcholine initiates sarcomere contraction. The Ca2+ ions binds to troponin-C (dark red) which leads to conformational changes that rotates tropomyosin, which occupies the groove between the two actin chains in the thin filament. This exposes a binding site on actin for the motor domain heads on the myosin thick filament.
An interactive iCn3D model of the Actin-Myosin-Tropomyosin ADP complex (6X5Z) is shown in Figure \(\PageIndex{5}\). - actin magenta, myosin cyan, tropomyosin gray, ADP and Mg spacefill
Figure \(\PageIndex{5}\): Actin-Myosin-Tropomyosin ADP complex (6X5Z). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...Q8KSdXLhjtNzt6
Now let's show an even larger cartoon view, which shows what is observed under a microscope. Figure \(\PageIndex{6}\) shows a cartoon representation of a sarcomere.
- the myosin thick filament is shown as a yellow rectangle;
- the actin thin filament is shown as a orange rectangle with tropomysin shown as a red line and the trimeric troponin complex shown in blue
The M line provides anchors for the thick filament while the Z disc attaches to and anchors the thin filaments. The myosin filaments are crosslinked and anchored at the end M band, which contains a series of proteins including myomesin, creatine kinase and M-protein. The Z disc/line contains many proteins including alpha-actinin (predominant), actin, many other proteins, and the end of titin. This cartoon shows the features shown in an electron micrograph of the vertebrate striated muscle sarcomere, which extends between the two Z-lines (or disks), as shown in Figure \(\PageIndex{7}\). A single bidirectional myosin thick filament is shown as an orangish rectangle, while single thin actin filaments are shown in blue (attached to the Z disc).
Figure \(\PageIndex{7}\): Electron micrograph of the sarcomere. John Squire. Int. J. Mol. Sci. 2019, 20, 5715; doi:10.3390/ijms20225715. Creative Commons Attribution
(CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Again note that the rod domains of the thick filaments are packed in an antiparallel fashion (ie. pointing in opposite directions) in the middle of the sarcomere near the end M line band, so in this region that are no myosin heads. However, the rest are packed in a parallel fashion and extend through the rest of the A-band. These myosin heavy chains are packed in a somewhat (quasi) symmetrical fashion, which repeats. Looking down the myosin fiber, the motor domains radiate outward with a 3-fold rotational symmetry and are staggered at 1200.
Now let's visualize the contraction of the sarcomere since it's hard to display in a single figure. Imagine it contracting somewhat like an accordion. The M line, devoid of myosin head groups and where the myosin heavy chains extend in different directions toward the left and right Z-lines, remains stationary. Since the myosin chains in the thick filament are attached to it, they remain stationary as well. Imagine the left and right Z line (disk), which anchor the actin thin filaments, moving towards the center line as the actin thin filaments slide along the stationary myosin thick filaments. This contracts the sarcomere. The sliding is powered, of course, by ATP hydrolysis in the motor heads of myosin in a series of steps involving attachment, detachment, and reattachment of the flexible myosin head domain further along with the actin thin filament.
Especially note the elongated protein called titin shown in cartoon form in Figure \(\PageIndex{6}\). This links the ends of the myosin filament and the Z-line. Titin is the largest single polypeptide known. Its molecular weight is close to 4 million with over 34,000 amino acids. It is highly elastic. It helps balance the forces on each side of the sarcomere. Its repetitive structure is shown in Figure \(\PageIndex{8}\). A series of structured Ig domains, comprised mostly of beta-strands are separated by flexible hinge structures which allow rearrangements and stretching.
The chemical cycle of adenine nucleotides consists of discrete steps which repeat: binding of ATP, hydrolysis, and dissociation of ADP and Pi. These are linked to a mechanical cycle involving attachment and detachment of myosin heads with the actin thin filaments and sliding of the thin filaments with respect to the thick myosin filaments.
Figure \(\PageIndex{9}\) shows the linked chemical and mechanical states in the cycle detailing how force is generated.
The myosin S1 head is shown at different angles. Starting with the bottom left in the ATP bound form, the head is bent backward to the left toward the rod domain. On hydrolysis of bound ATP, the head rotates into a vertical "cocked" position (top left). In this orientation, the S1 head binds weakly to actin (top right). A power stroke ensues as ADP and Pi dissociate from the myosin S1 head, reorienting the head to an orientation similar to the ATP-bound form.
As it is a cycle, you can start any place along it to follow it. Figure \(\PageIndex{10}\) shows a "linear" presentation of the cycle which starts with Ca2+ binding to troponin C. This exposes the myosin binding site on actin and allows the actin binding to the myosin S1 head to form the weakly bound M.ADP.Pi state. which is shown in the upper left of Figure \(\PageIndex{9}\).

In this diagram:
- In the resting (extended) sarcomere state, ADP and Pi are bound to the motor head of myosin. On neural stimulation of the muscle, Ca2+ ions are released from the sarcoplasmic reticulum and bind troponin-C. This leads to the movement of tropomyosin, which exposes the myosin motor head binding site on actin.
- On binding of actin, the power stroke results, as the actin filaments slide in contraction, leading to the dissociation of ADP and Pi.
- ATP then binds to the motor head, leading to the detachment of the myosin heads from the actin filament and ATP hydrolysis
- The motor head cocks to a position where it can interact again with actin filaments and start the process again.
More states also exist including an ATP-bound state that is detached from actin (post rigor), an ADP-bound state (M·ADP) without Pi, and a nucleotide-free myosin state that is bound to actin (rigor). Pi dissociation is the key step that leads to the movement of the lever arm when bound to actin, resulting in the power stroke. ADP dissociates slowly after this so the M·ADP lasts longer.
Figure \(\PageIndex{11}\) shows one last alternative figure with some of the states missing from the diagrams above and with corresponding structural features.
To make matters more complicated, there are 35 different classes of myosin (MW 227K) and it is involved in functions (vesicle transport for example) other than muscle contraction. We'll explore the structure of the S1 head more closely when we discuss another motor protein, kinesin.
No two-dimensional illustration of the process of sarcomere/muscle contraction can rival the insight gained by viewing a quality simulation of the molecular process. The video found below offers such a simulation. We don't routinely incorporate public videos into this book, in part since the links might not last and they often come with advertisements. This video, which is not narrated but richly annotated so clearly illustrates the ideas presented above, that we choose to incorporate it.
Here is a link to another video: Sliding movement of actin filaments
The cytoskeleton: An Overview
Before we explore two other motor proteins, kinesin and dynein, whose structures are very similar to myosin, we will offer some background on the proteins that make up the interior skeleton of the cell. Once again, the structure of the cytoskeleton is a bit amorphous and constantly changing. It structurally supports the inner organelles and other components of the cell. It facilitates the movement of these structures within the cell. It must dramatically rearrange when cells divide or when they move. More importantly for us, they interact with the motor proteins kinesin and dynein. If we don't know a bit about them, our understanding of motor proteins would lack context. Much of the following overview comes, unless otherwise noted, from a BioLibre introductory biology text by Easlon.
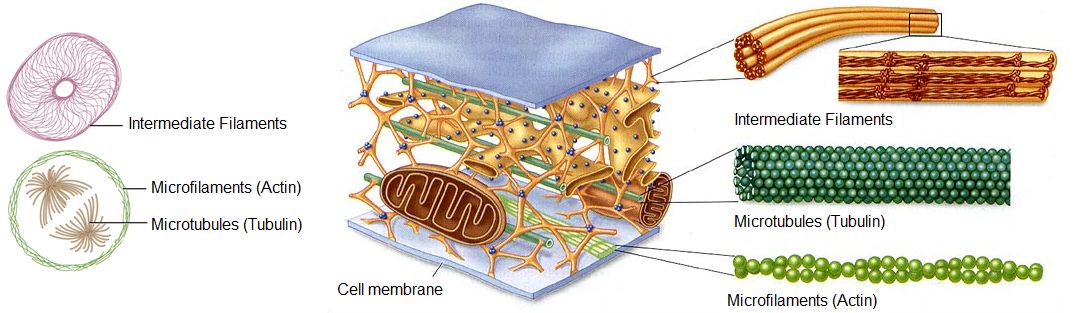
The cytoskeleton is a network of different protein fibers that provides many functions: it maintains or changes the shape of the cell; it secures some organelles in specific positions; it enables movement of cytoplasm and vesicles within the cell; and it enables the cell to move in response to stimuli. There are three types of fibers within the cytoskeleton: microfilaments, intermediate filaments, and microtubules, as shown in Figure \(\PageIndex{12}\). Some of the cytoskeletal fibers work in conjunction with molecular motors, which move along the fibers within the cell to carry out a diverse set of functions. There are two main families of cytoskeletally-associated molecular motors: dyneins and kinesins.
How can the cell purposely move and control the location of materials between these organelles? More specifically, how can a eukaryotic cell transport compounds from their place of origin (in most cases the cytoplasm) to where they are needed (perhaps the nucleus, the mitochondria, or the cell surface)?
One possible solution is for the cell to create a network that can connect all the different parts of the cell together. This network could be used not only as a scaffold to hold components in place but also as a reference for direction. For example, we can use a map to determine the direction we need to travel and the roads to connect and travel from home to campus. Likewise, an interconnecting network inside the cell can be used to direct and move compounds from one location to a final destination. Some of the required characteristics of this network are listed below. Can you add to this list?
Here are some characteristics of the cytoskeleton:
- The network needs to be extensive, and connect every area of the cell.
- The network needs to be flexible, able to change and adapt as the cell grows larger, divides into two cells, or physically moves from one environment to another.
- The network needs to be strong, and able to hold up to mechanical pressure from inside or outside of the cell.
- The network needs to be composed of different fibers and each of these fibers needs to be for a specific connection in the cell. For example, certain fibers might be involved in holding organelles in place, and other fibers would be involved in connecting two different organelles.
- The fibers need to have directionality (or polarity), meaning they need to have a defined starting point and a defined end to help direct movement from one location to another.
- The fibers need to work with proteins that can convert chemical energy into kinetic energy, to actively transport compounds along the fibers.
Figure \(\PageIndex{13}\) is a bovine pulmonary artery endothelial cells that as been stained to show thin filaments composed of actin (red), and microtubules (green), composed of tubulin.
Figure \(\PageIndex{13}\): Actin and microtubules in endothelial cells. https://sitn.hms.harvard.edu/art/201...tin-four-ways/. Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Microfilaments
Microfilaments are cytoskeleton fibers composed of actin subunits. Actin is the most abundant protein in the cytosol of eukaryotic cells and comprises 20% of total cellular protein by weight in muscle cells. The actin amino acid sequence is highly conserved in eukaryotic cells, meaning that the protein amino acid sequence, and therefore its final 3-D shape, has changed little over the course of evolution, maintaining more than 80% similarity between algae and humans. We have already seen actin filaments as part of the sarcomere.
Actin can be present as either a free monomer called G-actin (globular) or as part of a polymer microfilament called F-actin ("F" for filamentous). Actin must be bound to ATP to assemble into its filamentous form and maintain the structural integrity of the filament. (Note that we discussed above ATP binding to the myosin head and its subsequent hydrolysis during the chemomechanical sarcomere contraction, but we didn't discuss the role of ATP in F-actin formation.)
Recent Updates: Actin - 7/16/23
G-actin has 2 outer subdomains (1 and 2) and two inner ones (3 and 4). Between the outer and inner subdomains are a nucleotide-binding cleft and an opposing hydrophobic cleft involved in actin/actin and actin/actin-binding protein interactions necessary for filament formation.
An interactive iCn3D model of the rabbit F-Actin-ADP complex (2ZWG) is shown in Figure \(\PageIndex{14}\) below.
Figure \(\PageIndex{14}\): Rabbit F-Actin-ADP complex (2ZWG). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...n4poXDwJuqmASA
The large cleft between subdomains 2 (yellow) and 4 (cyan) is the nucleotide-binding cleft. The hydrophobic cleft is between subdomain 1 (salmon) and 3 (magenta). Many of the hydrophobic amino acid side chains in hydrophobic cleft are shown as gray sticks. Note that this form of Actin-ADP is flat on rotation.,
Figure \(\PageIndex{15}\) below shows an animation of the transition from G-Actin-ATP/Ca2+ complex (magenta) to F-Actin-ADP/Ca2+ complex (cyan)
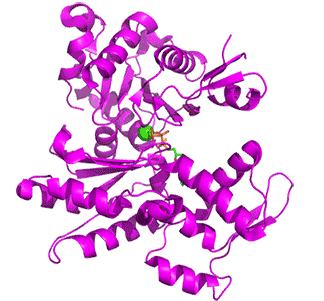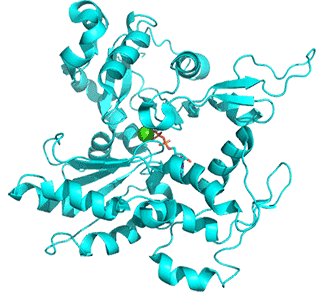
Figure \(\PageIndex{15}\) below shows an animation of the transition from G-Actin-ATP/Ca2+ complex (magenta) to F-Actin-ADP/Ca2+ complex (cyan)
Note the conformational changes and the changes in the cleft on the conversion of the G-ATP complex to the F-ADP one. Although the G in G-actin is for "globular", it is not really spherically. Both G- and F-actin are flat. Imagine the proteins as resembling the letter "H" but with the top "cleft" of the H (representing the nucleotide-binding domain) bigger than the bottom "cleft" (representing the hydrophobic cleft). In G actin, one of the long sides of the letter "H" is tilted compared to the other, while in F-actin, both sides are parallel, as shown in Figure \(\PageIndex{16}\) below. Hence F-actin is flatter than G-actin.
Figure \(\PageIndex{16}\): cartoon showing structural changes in G- to F-Actin transition (after https://www.science.org/doi/10.1126/science.adg6812)
The actin filament itself has structural polarity implying that it has two distinct ends to the filament. These ends are called the "(-)" end and the "(+)" end. At the "(+)" or "barbed" end, actin.ATP subunits are added to the elongating filament. At the "(-)" or "pointed" end, actin.ADP subunits dissociate from the filament. Actin hence is an ATPase. G-actin is a slow ATPase while F-actin is a fast ATPase due to a conformational change in the protein forming a flatter subunit conformation with a more catalytically productive active site.
This process of assembly and disassembly is controlled by the ATP to ADP ratio in the cytoplasm. The general structure of bare F-actin from Oryctolagus cuniculus is shown in an interactive iCn3D model Figure \(\PageIndex{17}\). The bottom strand (repeating subunits shown in cyan and light cyan) winds around the top strand (subunits shown in magenta and light magenta). They twist around each other in a right-handed sense. A complete turn is formed after in a 13-mer. You can see the twist in the figure below which shows only 4 monomers in each strand.
Figure \(\PageIndex{17}\): The structure of bare F Actin (6bno). (Copyright; author via source).
Click the image for a popup (slow load) or use this external link: https://structure.ncbi.nlm.nih.gov/i...QcXoreeyVaHhB7
Figure \(\PageIndex{18}\) below shows the free barbed end (magneta, 8F8R) and the free pointed end (cyan, 8F8S) of actin.
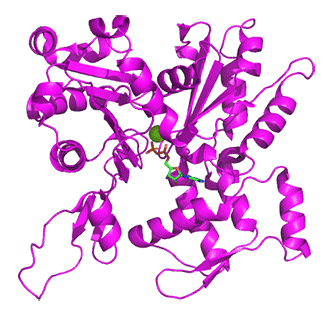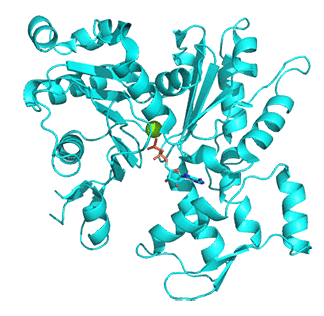
Figure \(\PageIndex{18}\): Free barbed end (magneta, 8F8R) and the free pointed end (cyan, 8F8S) of actin
Proteins interacting at the barbed and pointed ends include CapZ and tropomodulin, respectively. The CapZ binds to the F-actin flat "H" form with the CapZ undergoing significant conformational changes and minor one for the actin end. At the free pointed end, the monomers are more like G-actin in the twisted "H" form. On binding of tropomodulin, the second to the end subunit forms a more F actin conformation.
In the process of filament "treadmilling" (polymerization), actin-ATP binds to the + (barbed) end of F-actin. The bound ATP is quickly hydrolyzed with the Pi slowly released. Actin-ADP dissociates from the - (pointed_ end. Free actin can bind ATP and be ready to add to the + (barbed) end. Proteins like forming can catalyze the addition of actin monomers to the + (barbed) end while proteins like cofillin can promote dissociation from the - (pointed) end. Also both ends can be capped to stop actin addition at the + (barbed) end (by CapZ) or removal from the - (pointed) end (by tropomodulin).
As with myosin-actin interactions, hydrolysis of ATP and release of Pi are key steps. Multiple nucleotide states of actin include A.ATP, A.ADP.Pi, and A.ADP. As the polymer form, a gradient of these nucleotide states exist. On fiber formation, it appears that the key general base, His161, is positioned closer to the labile terminal phosphate on ATP, enabling hydrolysis.
Actin participates in many cellular processes, including muscle contraction, cell motility, cytokinesis during cell division, vesicle, and organelle movement, and the maintenance of cell shape. Actin filaments serve as a track for the movement of a family of motor proteins called myosins discussed above
Intermediate filaments
Intermediate filaments are made of several strands of fibrous proteins that are wound together as shown in Figure \(\PageIndex{19}\). These elements of the cytoskeleton get their name from the fact that their diameter, eight to ten nm, is between those of the smaller microfilaments and the larger microtubules. The intermediate filaments are the most diverse group of cytoskeletal elements. Several types of fibrous proteins are found in the intermediate filaments. You are probably most familiar with keratin, the fibrous protein that strengthens your hair, nails, and the epidermis of the skin.
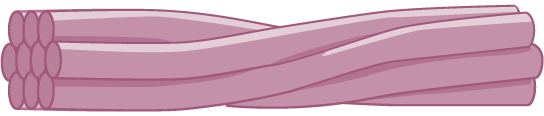
Intermediate filaments have no role in cell movement. Their function is purely structural. They bear tension, thus maintaining the shape of the cell, and anchor the nucleus and other organelles in place. The figure above shows how intermediate filaments create a cable-like supportive scaffolding inside the cell.
Microtubules
Microtubules are the largest component of the cytoskeleton and are found throughout the cytoplasm. These polymers are made up of globular protein subunits called α-tubulin and β-tubulin. Microtubules are found not only in eukaryotic cells but in some bacteria as well.
Both the α-tubulin and β-tubulin subunits bind to GTP instead of GTP. When bound to GTP, the microtubule starts to form with the first tubulins "nucleating" the growth of the growing tubule. As more GTP tubulin dimers assemble onto the filament, GTP is slowly hydrolyzed by β-tubulin to form GDP. Tubulin bound to GDP is less structurally robust and can lead to disassembly of the microtubule.
Much like the actin filaments discussed above, microtubules also have a distinct polarity that is critical for their biological function. Tubulin polymerizes end to end, with the β-subunits of one tubulin dimer contacting the α-subunits of the next dimer. These differences lead to different subunits being exposed on the two ends of the filament. The ends are designated the "(−)" and "(+)" ends. Unlike actin filaments, microtubules can elongate at both the "(+)" and "(-)" ends, but elongation is significantly more rapid at the "(+)" end. The structure of tubulin is shown in Figure \(\PageIndex{20}\). One turn of the tubulin (7diz) in cortical microtubules from the human parasite Toxoplasma gondii with proteins inside the tubulin polymers is shown. The alpha subunits are shown in cyan, beta in magenta and the inner proteins specific to this tubulin polymer in gray.
Figure \(\PageIndex{20}\): The structure of microtubules. Microtubules are hollow. Their walls consist of 13 polymerized dimers of α-tubulin and β-tubulin (right image). The left image shows the molecular structure of the tube.
Microtubules help the cell resist compression, provide a track along which vesicles move through the cell, pull replicated chromosomes to opposite ends of a dividing cell, and are the structural elements of flagella, cilia, and centrioles (the latter are the two perpendicular bodies of the centrosome). In fact, in animal cells, the centrosome is the microtubule organizing center. In eukaryotic cells, flagella and cilia are quite different structurally from their counterparts in bacteria, as discussed below.
A key function of microtubules is to move molecular "cargo" along the microtubule which acts like a "railroad" track. Two key motor proteins are involved in binding to target cargos, and hauling them along the polymer. These motor proteins are kinesin, which moves cargo towards the (+) end, and dynein, which moves it to the (-) end. These interactions are illustrated in Figure \(\PageIndex{21}\).
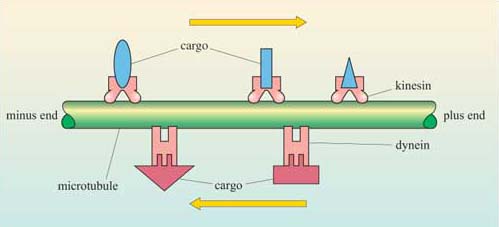
Figure \(\PageIndex{21}\): Kinesins and dyneins carry cargo along microtubules. Kinesin moves towards the plus end of the microtubule, whereas the dyneins move towards the minus end.
Here is an animation of the dynamics of micotubule and cargo movement.
Motor Proteins - Cargo movent along microtubule tracks
We will explore the motors protein kinesin and dynein in more detail. As they serve similar functions, you might expect them to have similar structures. Let's see!
Some of the material is from https://www.open.edu/openlearn/scien...nt-section-5.2
Kinesins
Motor proteins bind to vesicles and organelles and use energy from ATP to move them along the microtubule or microfilament network. Two families of motor proteins, the kinesins and dyneins, move vesicles along microtubules, and members of the myosin family move them along microfilaments. The myosin family is also important in cell movement.
The kinesin superfamily (KIF) consists of over 40 different proteins, divided into about 16 families. They bind both microtubules and ATP and are molecular motors. The first ones identified were involved in cargo (organelles, mRNAs, proteins) transport along the long axons of neuronal cells. This method of transport would be vastly superior than simple diffusion down the axons. The kinesin-1 family (members include KIF5A, KIF5B and KIF5C) are heterotetramers with two heavy and two light chains, somewhat similar to myosin.
As with myosin, kinesins have a motor head domain, which binds and hydrolyses ATP. They have a microtubule-binding domain instead of an actin-binding region. The N-kinesins have their motor domain at the end terminal, while the domain for M- and C- kinesins are in the middle or carboxy end, respectively. Both N-kinesins and C-kinesins are responsible for plus end and minus end-directed motility, and M-kinesins are used for depolymerization of microtubules in tubulin molecules. As with myosin, the neck stalk and tail are coiled coils of the heavy chains. Cargos bind through intermediary adapter proteins although some bind directly to kinesin tails.
The direction of movement of bound vesicles along the cytoskeleton is absolutely dependent on the polarity of the microfilaments and microtubules. Some motor proteins move from the minus end to the plus end and others in the opposite direction. For example, of the various myosins that have been discovered throughout the animal and plant kingdoms, all but one (myosin VI) move towards the plus end of the filament.
Kinesins have a tertiary structure that is similar to myosin II, even though there is no significant similarity in the primary structure. Both molecules have two heads with motor domains formed around an ATP-binding core, and a coiled tail that binds to the cargo, as shown in Figure \(\PageIndex{30}\). A number of other molecules are related to kinesin, and all of them share the kinesin motor domain, but very little else. These are the kinesin-related proteins. Kinesin itself moves towards the plus end of microtubules, but other members of the kinesin family move to the plus or minus end depending on the protein. Some of the kinesin-related proteins are involved in moving microtubules during mitosis – in this way the motor protein and the microtubule act in an analogous way to myosin and microfilaments in cell movement.
Figure \(\PageIndex{22}\) below shows an interactive iCn3D model of the neck and motor domains of dimeric Kinesin-3 KIF13B from rat (6A1Z)
Figure \(\PageIndex{22}\): Neck and motor domains of dimeric Kinesin-3 KIF13B from rat (6A1Z). (Copyright; author via source).
Click the image for a popup (slow load) or use this external link: https://structure.ncbi.nlm.nih.gov/i...LTKsQN4xcN9B99
One monomer is shown as a blue surface and the other as a gray cartoon. ADP is shown in colored spacefill. Amino acid side chains involved in nucleotide and microtubule binding are shown as coloreds sticks and labeled.
(Dyneins, which we will explore below, are unrelated to either kinesins or myosins, and they move towards the minus end of microtubules. Each is composed of two or three heavy chains, with the cytoplasmic dyneins having two chains, each of which forms a large motor domain. In nerve cells, the axonemal dyneins, which have two or three motor domains, transport vesicles along microtubule bundles in the axons.)
The speed of the movement mediated by dyneins and kinesins is signficant. In vitro, kinesins can move along microtubules at 2 μm s−1 and dyneins at up to 14 μm s−1. Although these high rates of movement would not be achieved in the complex environment of a cell, they can explain, for example, how caveolar transcytosis of molecules (from the luminal side of the blood vessel to the subendothelial space) across an endothelial cell can occur in 1–2 minutes. Movement and force generation by both classes of proteins involves ATP hydrolysis and allosteric shifts in the orientation of the motor domains, so that the proteins are thought to ‘step’ progressively down the microtubule.
Figure \(\PageIndex{23}\) shows a cartoon of a kinesin dimer attached to a microtubule.
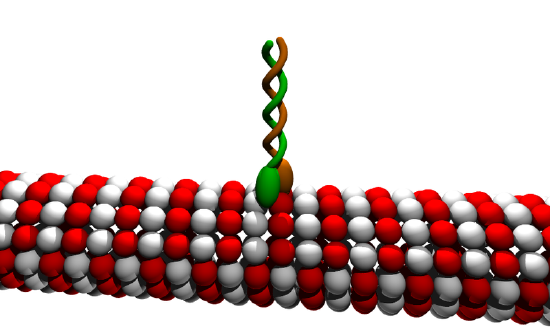
Figure \(\PageIndex{23}\): Cartoon of kinesin dimer attached to a microtubule. Moez. https://commons.wikimedia.org/wiki/F...in_cartoon.png. Creative Commons Attribution-Share Alike 3.0 Unported
Figure \(\PageIndex{24}\) shows an animated cartoon of Kinesin walking on microtubule.
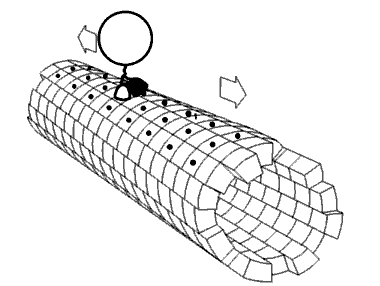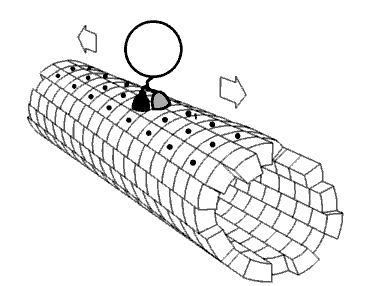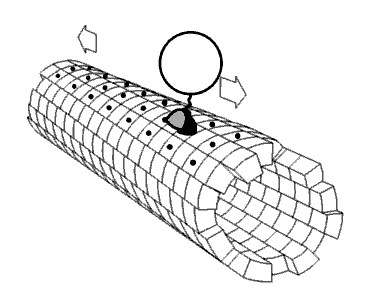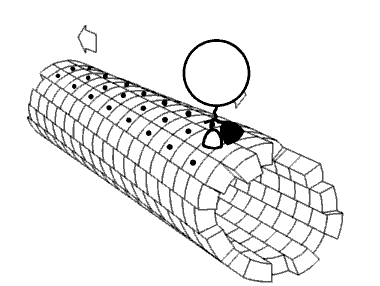
Figure \(\PageIndex{24}\): Cartoon of Kinesin walking on microtubule. By Jzp706 - Own work, CC0, https://commons.wikimedia.org/w/inde...=13188271jdkfj
Figure \(\PageIndex{25}\) shows an animation showing a kinesin dimer bound to a tubulin microtubule .

Figure \(\PageIndex{26}\) shows two motor domains of kinesin-1 bind to the microtubule lattice simultaneously.
Figure \(\PageIndex{26}\): Two motor domains of kinesin-1 bind to the microtubule lattice simultaneously. Qin, J.; Zhang, H.; Geng, Y.; Ji, Q. How Kinesin-1 Utilize the Energy of Nucleotide: The Conformational Changes and Mechanochemical Coupling in the Unidirectional Motion of Kinesin-1. Int. J. Mol. Sci. 2020, 21, 6977. https://doi.org/10.3390/ijms21186977. Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/)
The neck linkers of the kinesin-1 dimer are colored green. The leading head is in the nucleotide-free state and has an undocked neck linker, which points to the minus end of the microtubule. The trailing head is in the ADP·Pi/ADP-bound state and has a plus-end pointed neck linker.
Notice however that the ATP-binding site of kinesin is located at the distal tip of the motor domain, whereas in myosin the equivalent site is deep within the motor domain and covered by the myosin's actin-binding site. Therefore the mechanism of stepping is different in the two molecules. In particular, the α-helical linking region connecting the two motor domains of kinesin appears to transfer allosteric changes between them to coordinate ATPase activity and hence the stepping motion of the protein. It is interesting that the motor domains of kinesin-related proteins that move to the plus end and those that move to the minus end of the microtubule are similar but the linkage between them is quite different. Kinesin has its motor domain near the N-terminus, whereas the molecule Ncd, which moves to the minus end of the microtubule, has its motor domain located near the C-terminus. It seems that whether the protein is directed to the plus or the minus end is dependent on the configuration of domains and the coordination.
Figure \(\PageIndex{27}\): shows a comparison of structures of the motor head domain of myosin (panel A) and kinesin.ADP (panel B) and kinesin.ATP (panel C). T the structure of these proteins varies with the form of the bound nucleotide.
The myosin head has a 7-stranded central β-sheet and 3 α-helices on each side of the sheet. Nucleotides bind in the β-sheet regions which as a P (phosphate binding)-loop and two "switches". On the back side of the β-sheet are two 50K domains, in between which actin binds when exposed. A converter and lever arm follow leading to the coiled rod domain. The motor domain of kinesin is smaller but homologous with the tubulin binding site also on the back side of the β-sheet. These structures suggest similar mechanisms of action for myosin and kinesins. However, in contrast to the importance of Pi release in the power stroke for the sarcomere, it appears that ADP/ATP are the key players. When ADP is released, the central β-sheet twists, leading to binding to microtubules, implying that microtubules act to exchange ADP/ATP to generate the force. The linkage of conformational change and binding in microtubule:kinesin chemomechanical coupling is shown in Figure \(\PageIndex{28}\).
Kinesin is permanently attached to the microtubule, by either one head or the other. By comparison, the myosin heads (which are arranged in bundles in myofibrils) are only in contact with actin filaments for about 5% of each movement cycle.
Dynein
Like myosin and kinesin, dynein is a motor protein that undergoes chemomechanical cycles to move cargo on microtubules in the direction of the (-) end. Less is known about dynein given its much larger size (MW 1.4M) than myosin and kinesin, and the difficulty in studying it. There are cytoplasmic and flagellar versions of the protein. The latter generates a force that leads to flagellar motion and propulsion of the cell. They are part of the ATPases Associated with diverse cellular Activities (AAA+ proteins) superfamily. These often form hexameric ring complexes that are involved in the nucleotide-dependent changes in protein structure. Some acts as protease, chaperone, transcription factors, etc. Dynein is different from other AAA+ proteins as it is a single protein that includes six domains that arrange to form a hexameric ring structure. The domain structure and model of dynein bound to a microtubule are shown in Figure \(\PageIndex{29}\).
Figure \(\PageIndex{29}\): Overall structure of dynein. Upper panel Domain composition of dynein. The numbers of AAA+ modules (AAA1–AAA6) are indicated. ‘C’ indicates the C sequence. Lower panel The overall structure of dynein in the post-power stroke state. CC1 and CC2 indicate coiled-coil helices 1 and 2 of the stalk, respectively. Kato et al., ibid.
The structure contains:
- six AAA+ module domains with different sequences that nevertheless form a hexameric ring (labeled 1-6 in Figure \(\PageIndex{x}\) above;
- a lever-like linker domain that stabilizes the ring domains, connects to the tail domain and produces the power stroke;
- a coiled-coil antiparallel stalk domain contains two intertwined alpha-helices;
- a globular microtubule-binding domain (MTBD) that in the primary sequences of the protein separates the two alpha-helices in the stalk and which binds the "track" (microtubule).
- a tail-domain that binds to cargos, or if multiple dyneins interact, other dyneins.
It is quite amazing that dynein has a 3D structure (globular cytoskeleton binding domain - stalk - tail) that mimics that of myosin and kinesin, but all in a single protein. The power stroke is linked to lever motion and ATP hydrolysis. We must use this structural information to understand ATP binding and hydrolysis, conformational changes in the linker during the power stroke, transmission of conformational changes through the stalk and ensuing changes in the interaction of the microtubule-binding domain with the microtubule.
Nucleotide binding and hydrolysis
- the lever-like linker swings between AAA+2 and the stalk base(Roberts et al. 2009).
- AAA+ modules 1-4 can bind an cleave ATP so they acts as ATPases. This contrasts to both myosin and kinesis which have only 1 ATP binding site per heavy chain. AAA+ 5 and 6 have mutations, which prevent ATP hydrolysis.
- The ATPase site of AAA1 is between AAA1 and AAA2 binds ADP with higher affinity than ATP since an arginine side chain on AAA2 is not close enough to interact with the gamma-phosphate of ATP. The ADP-bound form is called the open form.
- Crystals structure of ADP.vanadate (an equivalent ATP analog) shows a closed active site in which the arginine side chains interact with the vanadate, a proxy for the gamma-phosphate of ATP. Hence ATP binding closes this active site. A
Figure \(\PageIndex{30}\) shows the open (ADP bound) and closed (ATP equivalent) bound state of dynein.
Linker conformational changes in power stroke
Linker motion produces the power stroke and the ADP bound form represents a post-stroke state in which the linker has minimal contact with the ring domains and is mostly detached. A long α-helix, H10, within the linker appears to be involved in the power stroke as it extended in th ADP bound state, but bent in the ATP proxy state, where it is posed to initiate the power stroke. These conformations are illustrated in Figure \(\PageIndex{31}\).
Communication between the domains for ATP hydrolysis and track binding
Obviously, there needs to be coordination and correcting timing between changes in the ATPase hexameric ring and its communication through the linker to the stalk and to the microtubule-binding domain (MTBD), which is 140 Å distant. This communication is easier for kinesins and myosins since the motor domain, where ATP cleavage occurs, is also the domain that binds actin and tubulin, respectively. The communication between the distal domains occurs through the coiled-coiled stalk. This makes sense as the MTBD is in between the two helices of the stalk.
The two antiparallel helices presumably slide over each other to form different alignments or registries between the helices. Indeed different alignments appear to affect the affinity of the MTBD for tubulin. This was experimentally verified by creating a new protein with the coiled-coil stalk of dynein with another protein, seryl-tRNA synthetase, as illustrated in Figure \(\PageIndex{32}\).
Interactions with tubulin
There also appear to be two main conformations for the MTBD for its interaction with tubulin. Figure \(\PageIndex{x}\) below shows the domain structure of dynein and various synthetic constructs used in experiments (panel A), a model structure for the interaction of dynein with the α/β-tubulin dimer (panel B) and a detailed rendering of the strong (alpha-helix registry) and weak (beta structure registry) of the MTBD and the α/β-tubulin dimer. During each full cycle, the MTBD must dissociate from and reassociate with the microtubule as it marches along the polymer (hence the need for weak and strong binding interactions). Figure \(\PageIndex{33}\) below shows the domain organization of dynein and low- and high-affinity bound states of the MTBD.
Figure \(\PageIndex{33}\): Domain organization of cytoplasmic dynein and low- and high-affinity MT-bound states of the dynein MTBD. a Organization of the full-length cytoplasmic dynein heavy chain (HC) with an N-terminal Halo-Tag, Halo-Dyn1471kDa (a.a. 1–4092) and the tail-truncated monomeric constructs, GFP-Dyn1331kDa and Dyn1331kDa-GFP (a.a. 1219–4092). b Dynein MD bound to α/β-tubulin in the strong binding state (merged from PDB entries 3VKG and 3J1T; see Supplementary Note 1). c MT-bound MTBD in the weak MT-binding β-registration of the stalk helices (top, PDB entry 3J1U) and in the strong MT-binding α-registration (bottom, PDB entry 3J1T). Lu Rao et al. Nature Communications (2019) 10:3332 | https://doi.org/10.1038/s41467-019-11231-8. Creative Commons Attribution 4.0 International License.




.png?revision=1&size=bestfit&width=465&height=244)
.png?revision=1)
.png?revision=1&size=bestfit&width=425)