1.1: Cellular Foundations
- Page ID
- 14913
Introduction
You have probably studied the cell many times, either in high school or in college biology classes. There are many websites available that review both prokaryotic (bacterial and archaeal cell types) and eukaryotic cells (protist, fungi, plant, and animal cell types). All cells have some similar structural components, including genetic material in the form of chromosomes, a membrane-bound lipid bilayer that separates the inside of the cell from the outside of the cell, and ribosomes that are responsible for protein synthesis. This tutorial is designed specifically from the viewpoint of chemistry. It explores four classes of biomolecules that are also present in all cell types (lipids, proteins, nucleic acids and carbohydrates) and describes in a simplified pictorial manner where they are found, made, and degraded in a typical eukaryotic, animal cell (i.e. their history). This cell review focuses on the organelle structures common in eukaryotic cells. Subsequent chapters will concentrate on the structure and function of specific biomolecules.
Let’s think of a cell as a chemical factory that designs, imports, synthesizes, uses, exports, and degrades a variety of chemicals (in the case of the cell, these include lipids, proteins, nucleic acids, and carbohydrates). It also must determine or sense the amount of raw and finished chemicals it has available and respond to its own and external needs by ramping up or shutting off production. Biochemistry is the branch of science dedicated to the study of these chemical processes within a cell. Understanding these processes can also lend insight into disease states and the pharmacological effects of toxins, drugs, and other medicines within the body.
The building and breaking down of life-sustaining chemicals within an organism is known as Metabolism. Overall, the three main purposes of metabolism are: (1) the conversion of food to energy to run cellular processes; (2) the conversion of food/fuel to building blocks for the production of primary metabolites, such as proteins, lipids, nucleic acids, and other secondary metabolites; and (3) the elimination of waste products. These enzyme-catalyzed reactions allow organisms to grow and reproduce, maintain their structures, and respond to their environments.
Metabolic reactions may be categorized as catabolic– the breaking down of compounds (for example, the breaking down of proteins into amino acids during digestion); or anabolic – the building up (synthesis) of compounds (such as proteins, carbohydrates, lipids, and nucleic acids). Usually, catabolism releases energy, and anabolism consumes energy.
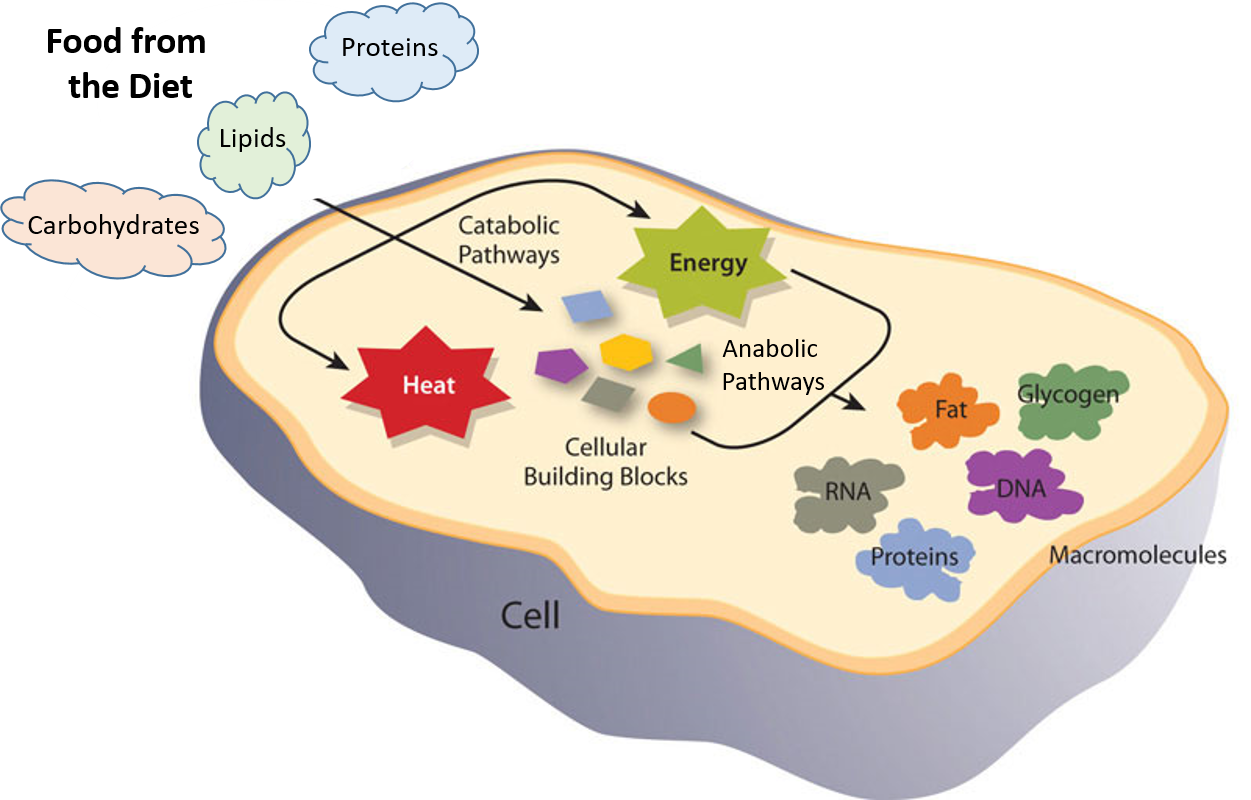
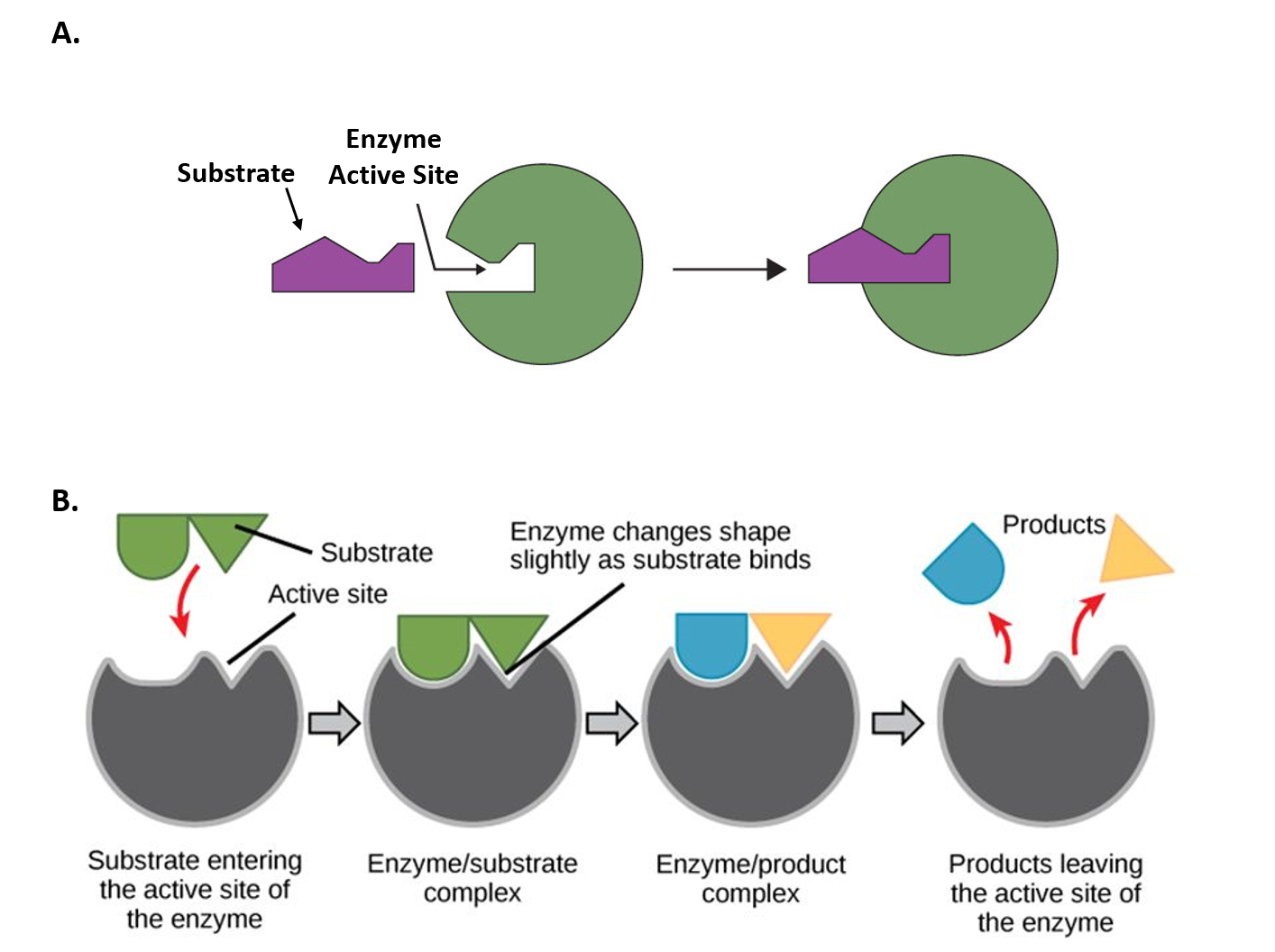
The chemical reactions of metabolism are organized into metabolic pathways, in which one chemical is transformed through a series of steps into another chemical, each step often being facilitated by a specific enzyme. Enzymes are crucial to metabolism because enzymes act as catalysts – they allow a reaction to proceed more rapidly. In addition, enzymes can provide a mechanism for cells to regulate the rate of a metabolic reaction in response to changes in the cell’s environment or to signals from other cells, through the activation or inhibition of the enzyme’s activity. Enzymes can also allow organisms to drive desirable reactions that require energy that will not occur by themselves, by coupling them to spontaneous reactions that release energy. Enzyme shape is critical to the function of the enzyme as it determines the specific binding of a reactant. This can occur by a lock and key model where the reactant is the exact shape of the enzyme binding site, or by an induced fit model, where the contact of the reactant with the protein causes the shape of the protein to change in order to bind to the reactant. The catalytic mechanisms, kinetics, and regulatory pathways of enzymes will be studied in detail within this text.
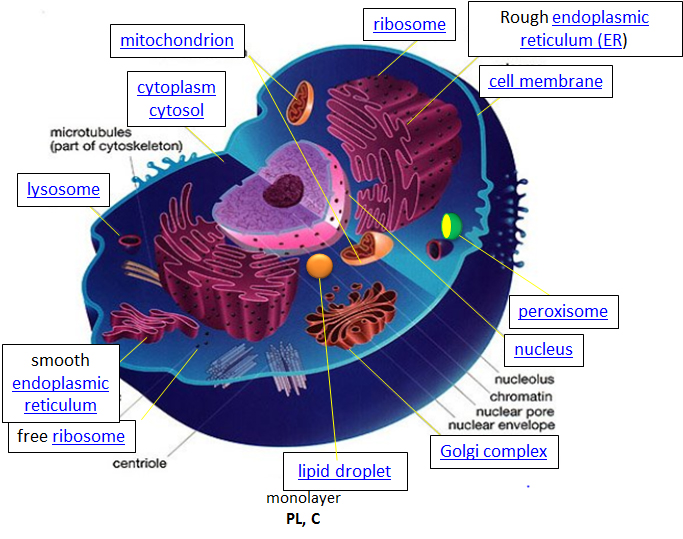
Within eukaryotic cells, the metabolic machinery present allows for the construction of membrane-bound organelle structures that help to compartmentalize cellular functions. Therefore, organelles can be thought of as ‘little organs’ within the cell having discrete cellular functions. The figure of the cell below and in the other linked sites based on it was made available with the kind permission of Liliana Torres. Click on the blue hyperlinks for some of the organelles for more detailed information on them.
Design – The design for a cell mostly resides in the blueprint for the cell, the genetic code, which is comprised of the DNA in the cell nucleus and a small amount in the mitochondria. Of course, the DNA blueprint must be read out (transcribed) by ribosomes which themselves were encoded by the DNA and contain a combination of RNA and protein subunits. The genetic code has the master plan that determines the sequence of all cellular proteins, which then catalyze almost all other activities in the cell, including catalysis, motility, architectural structure, etc. In contrast to DNA, RNA, and protein polymers, the length and sequence of polysaccharide polymers and lipids are not driven by such a template but rather by the enzymes that catalyze the synthesis.
Import/Export: Many of the chemical constituents of the cell arise not from direct synthesis but from the import of both small and large molecules. The imported molecules must pass through the cell membrane and in some cases through additional membranes if they need to reside inside membrane-bound organelles. Molecules can move into the cell by passive diffusion across the membrane but usually, their movement is “facilitated” by a membrane transporter protein. Molecules can also move against a concentration gradient in a process called “active transport”. Given the amphiphilic nature of the bilayer (polar head group exterior, nonpolar interior), you would expect that polar molecules like glucose would have difficulty in moving across the membrane by passive diffusion. Typically, only small nonpolar molecules move across the membrane via passive transport. Membrane-bound transport proteins are involved in the movement of both nonpolar and polar molecules.
- transporters, carrier proteins, and permeases: These membrane proteins move specific ligand molecules across a membrane, typically down a concentration gradient. Computer simulations of the facilitated diffusion of lactose across the membrane are shown in the following link. Animation of lactose diffusion through the LacY receptor (The link above and immediately below are from the Theoretical and Computational Biophysics group at the Beckman Institute, the University of Illinois at Urbana-Champaign. These molecular dynamic simulations were made with VMD/NAMD/BioCoRE/JMV/other software support developed by the Group with NIH support.)
- ion channels – These membrane proteins allow the flow of ions across membranes. Some are permanently open (nongated) while others are gated open or closed depending on the presence of ligands that bind the protein channel and the local environment of the protein in the membrane. The flow of ions through the channel proceeds in a thermodynamically favored direction, which depends on their concentration and voltage gradients across the membrane.
- pores: Some membranes (nuclear, mitochondria) assemble proteins (such as porins) to form large, but regulated pores. Porins are found in mitochondrial membranes while nucleoporins are found in the nuclear membrane. Small molecules can generally pass through these membrane pores while large ones are selected based on their tendency to form transient intermolecular attractive forces with the pore proteins. The following link shows the diffusion of water through aquaporin. animation of water diffusion through the aquaporin channel,
- endocytosis: Very large particles [for example, Low Density Lipoproteins (LDL) and viruses] can enter a cell through a process called endocytosis. Initially, the LDL or virus binds to a receptor on the surface of the cell. This triggers a series of events that leads to the invagination of the cell membrane at that point. This eventually pinches off to form an endosomal vesicle which is surrounded by a protein called clathrin. “Early” endosomes can pick up new proteins and other constituents as well as shed them as they move and mature through the cell. During this maturation process, protein pumps in the endosome lead to a decrease in the endosomal pH which can lead to conformation changes in protein structure and shedding of proteins. Eventually, the “late” endosome reaches and fuses with the lysosome, an internal organelle that contains degradative enzymes. Undegraded components like viral nucleic acids or cholesterol are delivered to the cell. This transport can also go in the reverse direction (called exocytosis) and recycle receptors to the cell membrane. Likewise, vesicles pinched off from the Golgi complex can fuse with endosomes, with some components surviving the process to reenter the Golgi.
Synthesize/Degrade: Cells have to synthesize and degrade small molecules as well as larger polymeric proteins, carbohydrates, lipids, and nucleic acids. The anabolic (synthetic) and catabolic (degradative) pathways are often compartmentalized in time and space within a cell. For example, fatty acid synthesis is carried out in the cytoplasm but fatty acid oxidation is carried out in the mitochondria. Proteins are synthesized in the cytoplasm or completed in the endoplasmic reticulum (for membrane and exported proteins) while they are degraded in the lysosome or more importantly in a large multimolecular structure in the cell called the proteasome.
Key Characteristics of a Cell
Let’s consider some key characteristics of a cell before we get into the details in later chapters.
Cells and their internal compartments have regulated concentrations of ions and hydronium ions.
As expected the pH of the cytosol (the aqueous substance surrounding all the organelles within the cell) varies from about 7.0-7.4, depending on the metabolic state of the cell. Some organelles have proton transporters that can significantly alter the pH inside an organelle. For example, the pH inside the lysosome, a degradative organelle, is about 4.8. Furthermore, the creation of a pH gradient across the inner mitochondrial membrane is sufficient to drive the thermodynamically unfavored synthesis of ATP.
Compared to the extracellular fluid, the concentration of potassium ions is higher inside the cell, while concentrations of sodium, chloride, and calcium ions are higher on the outside of the cell (see table below). These concentration gradients are maintained by ion transporters and channels and require energy expenditure ultimately in the form of ATP hydrolysis. Changes in these concentrations are integral to the signaling system used by the cell to sense and respond to changes in its external and internal environments. The table below shows approximate ion concentrations in the cell.
| Ion | Inside (mM) | Outside (mM) |
|---|---|---|
| Na+ | 140 | 5 |
| K+ | 12 | 140 |
| Cl- | 4 | 15 |
| Ca2+ | 1 uM | 2 |
Cells have an internal framework that provides architectural and internal structural support
The “cytoskeletal” architecture of a (with molecular “cables”- and “girder-like” structures) is not dissimilar from a factory. The internal framework of a cell or cytoskeleton, is composed of microfilaments, intermediate filaments, and microtubules. These are comprised of monomeric proteins which self-assemble to form the internal architecture. Parts of the cytoskeleton can be seen in Figure 1.4.

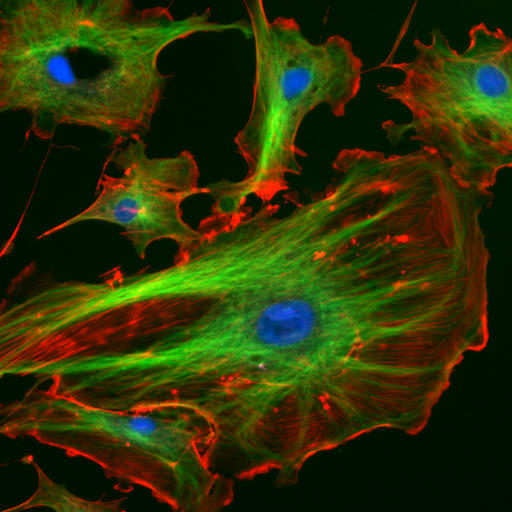
Microfilaments of actin monomers (which are stained with a red/orange fluorophore) and microtubules which offer more structural support made of tubulin monomers (stained green) along with the blue-stained nucleus are shown in the image. Organelles are supported and organized by the cytoskeleton (primarily microtubules). Even the cell membrane is supported underneath the inner leaflet by actin (stained orange) and spectrin microfilaments. Motor proteins like myosin (that moves along actin microfilaments) and dynein and kinesin (that move along tubulin microtubules) carry cargo (vesicles, organelles) in a directional fashion. The cell is not a disorganized collection of molecules and organelles. Rather it is highly organized for optimal chemical production, use, and degradation.
Cells have a variety of shapes. Some circulating immune cells must slip through the cells that line capillary walls to migrate to sites of infection. The same process occurs when tumor cells metastasize and escape to other sites in the body. In order to do so, the cell must drastically change shape, a response that requires the dissociation of the cytoskeleton polymers into monomers which are available later for repolymerization. The following video shows the mobility and flexibility of a Killer T-Cell as it attacks and kills a cancerous cell.
Video 1.1 Killer T Cell Attacking Cancer. Video available on YouTube through creative commons by Cambridge University
The cell is an amazingly crowded place
In chemistry labs, we typically work with dilute solutions of solute molecules in a solvent. You have probably heard that the body is comprised of 68% water, but the water concentration is obviously dependent on the cellular environment. Solute molecules like protein and carbohydrates are densely packed. Cells are so crowded that the space between larger molecules like proteins is typically smaller than that of a single protein. Studies have shown that the stability of a protein is increased in such conditions, which would help keep the protein in the correctly folded, native state. Another consequence of high intracellular concentrations is that it limits the diffusion of molecules throughout the cell, as would be expected from an equilibrium perspective in dilute solutions. Thus, cytoplasmic cellular functions can be highly localized within specific regions of the cell creating unique microenvironments and higher differentiation potential within a single cell.
Hence the study of biomolecules in dilute solutions in the lab may not reveal the actual complexities of interactions and activities of the same molecule in vivo. Recently investigators have added a neutral copolymer of sucrose and epichlorohydrin to cells in vitro. These particles induced the organization of extracellular molecules secreted by the cell, forming an organized extracellular “matrix” which induced the organization of the microfilaments on the inside of the cell as well as inducing changes in cell activity.1 Furthermore, in vitro enzyme activity of a key enzyme in glycolysis dramatically increases under crowded conditions.2 Another result of crowding may be the spatial and temporal association of key enzymes involved in specific metabolic pathways, allowing for the coordinated passage of substrates and products within the colocalized enzyme system.
Cell components undergo phase transitions to form substructures within the cell.
A perplexing question is how substructures form within a cell. This includes not only the biogenesis of organelles like mitochondria but also smaller particle such as polysaccharide granules, lipid droplets, protein/RNA particles (including the ribosome) as well as the nucleolus of the cell nucleus. It might be easiest to consider this problem using two examples from the lipid world, lipid droplets and membrane rafts. You are very familiar with phase transitions that occur when a sparing soluble nonpolar liquid is added to water. At a high enough concentration, the solubility of the nonpolar liquid is exceeded and a phase transition occurs as evidenced by the appearance of two separate liquid phases. The same process occurs when triglycerides coalesce into lipid droplets with proteins associated on their outside. Another example occurs within a cell membrane when lipids with saturated alkyl chains self-associate with membrane cholesterol (which contains a rigid planar ring system) to form a membrane microdomain called a lipid raft. Lipid rafts are characterized by greater packing efficiency, rigidity, and thickness than other parts of the membrane. These lipid rafts often recruit proteins involved in signaling processes within the cell membranes. This process of phase separation is also called liquid/liquid demixing as two “liquid-like” substances separate.
In a similar manner, it appears that proteins that interact with RNA are composed of less diverse amino acid sequences and have more flexible (“more liquid-like) structures allowing their preferential interaction with RNA to form large RNA-protein particles (like the ribosome and other RNA processing structures) in a fashion that mimics liquid/liquid demixing. All of these interactions are just manifestations of the various intermolecular forces that can exist between molecules. These include ionic interactions, ion-dipole interactions, dipole-dipole interactions, and London dispersion forces (A review of intermolecular forces can be found by Kahn Academy on YouTube).



