8.10: Genome Editing (CRISPR)
- Page ID
- 10448
The development of tools that would allow scientists to make specific, targeted changes in the genome has been the Holy Grail of molecular biology. An ingenious new tool that is both simple and effective in making precise changes is poised to revolutionize the field, much as PCR did in the 1980s. Known as the CRISPR/Cas9 system, and often abbreviated simply as CRISPR, it is based on a sort of bacterial immune system that allows bacteria to recognize and inactivate viral invaders.
CRSPR
CRISPR stands for Clustered Regularly Interspaced Short Palindromic Repeats, short repeated sequences found in prokaryotic DNA, separated by spacer sequences derived from past encounters with, for example, a bacteriophage. Like the glass slipper left behind by Cinderella that was later used to identify her, the pieces of the invader's sequences are a way for the bacteria to identify the virus if it attacks again. Inserted into the bacterial genome, these sequences can later be transcribed into a guide RNA that matches, and base-pairs with, sections of the viral genome if it was encountered again. A nuclease associated with the guide RNA then cleaves the sequence base-paired with the guide RNA. (The nucleases are named Cas for CRISPR-associated.)
The essential elements of this system are a guide RNA that homes in on the target sequence and a nuclease that can make a cut in the sequence that is bound by the guide RNA. By engineering guide RNAs complementary to a target gene, it is possible to target the nuclease to cleave within that gene. In the CRISPR/Cas9 system, the Cas9 endonuclease cuts both strands of the gene sequence targeted by the guide RNA (Figure \(\PageIndex{1}\)). This generates a double-strand break that the cell attempts to repair.
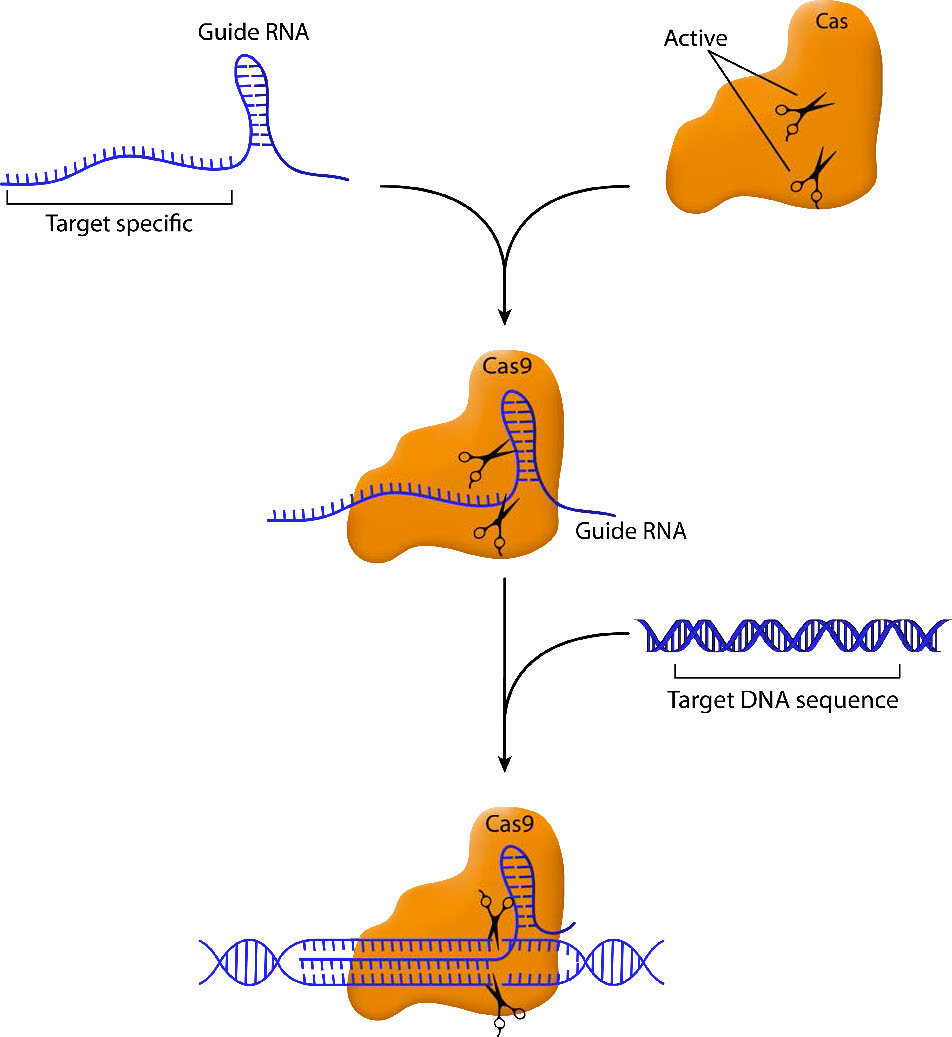
As you may remember, double-strand breaks in DNA can be repaired by simple, nonhomologous end joining (NHEJ) or by homologous recombination. When a break is fixed by NHEJ, there is good chance that there will be deletions or insertions that will inactivate the gene they are in. Thus, targeted cleavage of a site by CRISPR/Cas9 can easily and specifically inactivate a gene, making it easy to characterize the gene's function.
But, what if you wished to simply mutate the gene at a specific site to study the effect of the mutation? This, too, can be achieved. If a homologous sequence bearing the specific mutation is provided, homologous recombination can repair the break, and at the same time insert the exact mutation desired. It is obvious that if you can insert a mutation as just described, it should be possible to correct a mutation in the genome by cleaving at the appropriate spot and providing the correct sequence as a template for repair by homologous recombination. The simplicity of the system holds great promise for curing genetic diseases.
Scientists have also come up with some creative variations on the CRISPR/Cas9 system. For instance, one variant inactivates the nuclease activity of Cas9. The guide RNA in this system pairs with the target sequence, but the Cas9 does not cleave it. Instead, the Cas9 blocks the transcription of the downstream gene (Figure \(\PageIndex{2}\)) This method allows specific genes to be turned off without actually altering the DNA sequence.
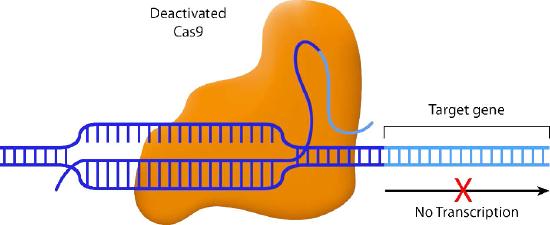
Another variation also uses a disabled Cas9, but this time, the Cas9 is fused to a transcriptional activation domain. In this situation, the guide RNA positions the Cas9-activator domain in a place where it can enhance transcription from a specific promoter (Figure \(\PageIndex{3}\)). Other variations on this theme attach histone-modifying enzymes or DNA methylases to the inactive Cas9. Again, the guide RNA positions the Cas9 in the desired spot, and the enzyme attached to Cas9 can methylate the DNA or modify the histones in that region.
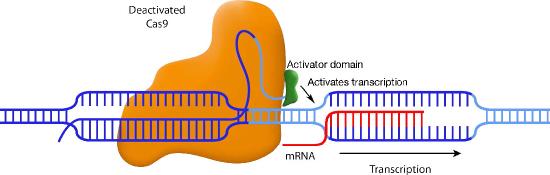
CRISPR has already been used to edit genomes in a wide variety of species (and in human cell cultures). It may not be long before the technique is approved for clinical use. In the meanwhile, CRISPR is transforming molecular biology.


