8.4: Detection, identification and quantitation of specific nucleic acids and proteins
- Page ID
- 10442
While gel electrophoresis can be used to resolve molecules in a mixture, by itself, the technique does not permit the detection and identification of specific nucleic acid sequences or proteins. For example, the 2-D gel shown above clearly separates a large number of proteins in a sample into individual spots. However, if we wanted to know whether a specific protein was present, we could not tell by simply looking at the gel. Likewise, in an agarose gel, while bands of DNA could be assigned a size, one could not distinguish between two DNAs of different sequence if they were both the same length in base-pairs. One way to detect the presence of a particular nucleic acid or protein is dependent on transferring the separated molecules from the gels onto a membrane made of nitrocellulose or nylon to create a “blot” and probing for the molecule(s) of interest using reagents that specifically bind to those molecules. The next section will discuss how this can be done for nucleic acids as well as for proteins.
Southern and Northern Blots
The Southern blot is named for its inventor, Oxford professor, Edwin Southern, who came up with a protocol for transferring DNA fragments from a gel onto a nitrocellulose sheet and detecting a specific DNA sequence among the bands on the blot. As shown in Figure 8.21, the method works as follows. A mixture of DNA molecules (often DNA that has been cut into smaller fragments using restriction endonucleases) is loaded on an agarose gel, as usual. After the gel run is complete, the DNA bands are transferred from the gel onto a membrane. This can be achieved by capillary transfer, where the gel is placed in contact with a piece of membrane and buffer is pulled through the gel by wicking it up into a stack of absorbent paper placed above the membrane. As the buffer moves, it carries with it the DNA fragments. The DNA binds to the membrane leaving a “print” of DNA fragments that exactly mirrors their positions in the gel. The blotting membrane may be treated with UV light, heat, or chemicals to firmly attach the DNA to the membrane.
Next, a probe, or visualizing agent specific for the molecule of interest is added to the membrane. In Figure 8.21, this is called a labeled probe. The probes in a Southern blot are pieces of DNA designed to be complementary to the desired target sequence. If the sequence of interest is present on the blot, the probe, which is complementary to it, can base-pair (hybridize) with it. The blot is then washed to remove all unbound probe. Probes are labeled with radioactivity or with other chemical reagents that allow them to be easily detected when bound to the blot, so it is possible to visually determine whether the probe has bound to any of the DNA bands on the blot. Given that the Southern blot relies on specific base-pairing between the probe and the target sequence, it is easy to adapt the technique to detect specific RNA molecules, as well. The modification of this method to detect RNAs was jokingly named a “northern” blot.
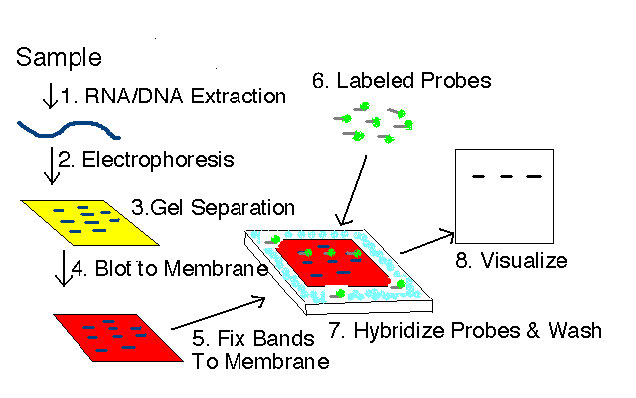
Figure 8.21 - Northern or Southern blot scheme. Southern blotting adds strand denaturation between steps 4 and 5. Wikipedia
Western Blots
Proteins cannot, for obvious reasons, be detected through base-pairing with a DNA probe, but protein blots, made by transferring proteins, separated on a gel, onto a membrane, can be probed using specific antibodies against a particular protein of interest. Protein detection usually employs two antibodies, the first of which is not labeled. The label is on the second antibody, which is designed to recognize only the first antibody in a piggyback fashion. The first antibody specifically binds to the protein of interest on the blot and the second antibody recognizes and binds the first antibody. The second antibody commonly carries an enzyme or reagent which can cause a reaction to produce a color upon further treatment. In the end, if the molecule of interest is in the original mixture, it will “light” up and reveal itself on the blot. This variation on the blotting theme was dubbed a western blot (Figure 8.22).
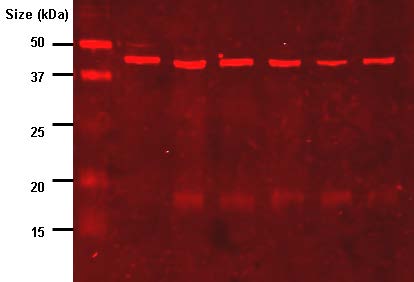
Figure 8.22 - Result of a western blot analysis. Wikipedia
In each of the blots described above, binding of the probe to the target molecule allows one to determine whether the target sequence or protein was in the sample. Although blots are designed to be used for detection, rather than for precise quantitation, it is possible to obtain estimates of the abundance of the target molecule from densitometry measurements of signal intensity.
Microarrays
2-D gels are a way of surveying a broad spectrum of protein molecules simultaneously. One approach to doing something similar with DNA or RNA involves what are called microarrays. Microarrays are especially useful for monitoring the expressions of thousands of genes, simultaneously. Where a northern blot would allow the identification of a single mRNA from a mixture of mRNAs, a microarray experiment can allow the simultaneous identification of thousands of mRNAs that may be made by a cell at a given time. It is also possible to perform quantitation much more reliably than with a blot.
Microarrays employ a glass slide, or chip, to which are attached short sequences of single-stranded DNA, arranged in a grid, or matrix (Figure 8.23) Each position in the grid corresponds to a unique gene. That is, the DNA sequence at this spot is part of the sequence of a specific gene. Each spot on the grid has multiple identical copies of the same sequence. The gene sequence immobilized at each position in the grid is recorded.
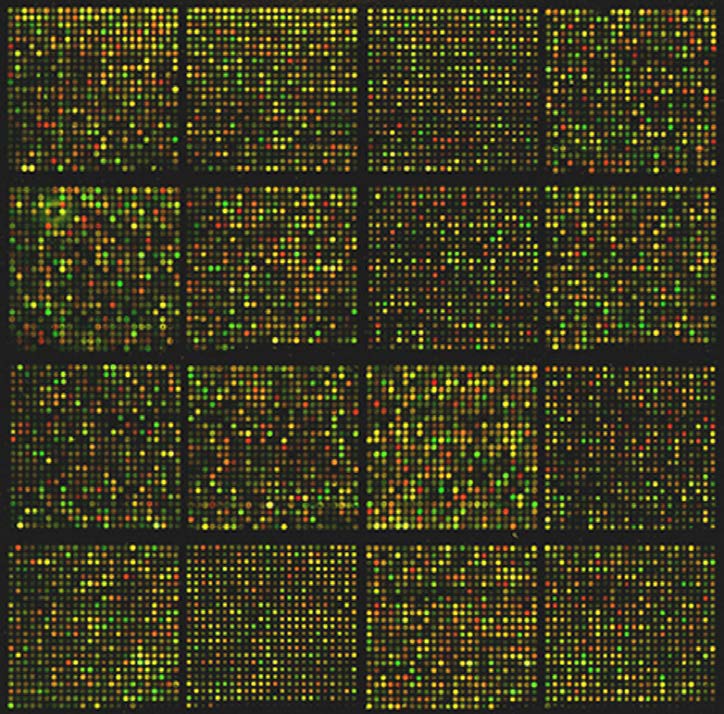
Figure 8.23 - Microarray design. Image by Taralyn Tan
To the slide are added a mixture of sample molecules, some of which will recognize and bind specifically to the sequences on the slide. Binding between the sample molecules and the sequences attached to the slide occurs by base pairing, in the case of DNA microarrays. The slide is then washed to remove sample molecules that are not specifically bound to the sequences in the grid.
Sample molecules are tagged with a fluorescent dye, allowing the spots where they bind to be identified. The grid is analyzed spot by spot for binding of the sample molecules to the immobilized sequences. The more sample molecules are bound at a spot, the greater the intensity of dye fluorescence that will be observed. Information from this analysis can give information about the presence/absence/abundance of molecules in the sample that bind to the sequences in the grid.
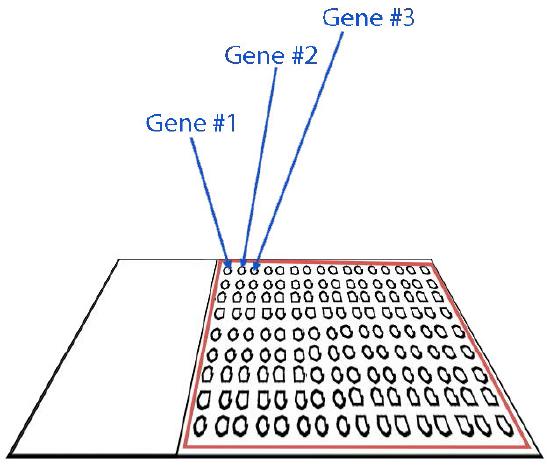
Figure 8.24 - Large scale microarray analysis of mouse transcriptome. Wikipedia


