8.2: Fractionation and Chromatography Techniques
- Page ID
- 10439
Fractionation of samples, as the name suggests, is a process of separating out the components or fractions of the lysate. Fractionation typically begins with centrifugation of the lysate. Using low-speed centrifugation, one can remove cell debris, leaving a supernatant containing the contents of the cell. By using successively higher centrifugation speeds (and resulting g forces) it is possible to separate out different cellular components, like nuclei, mitochondria, etc., from the cytoplasm. These may then be separately lysed to release molecules that are specific to the particular cellular compartment. The soluble fraction of any lysate can, then, be further separated into its constituents using various methods.
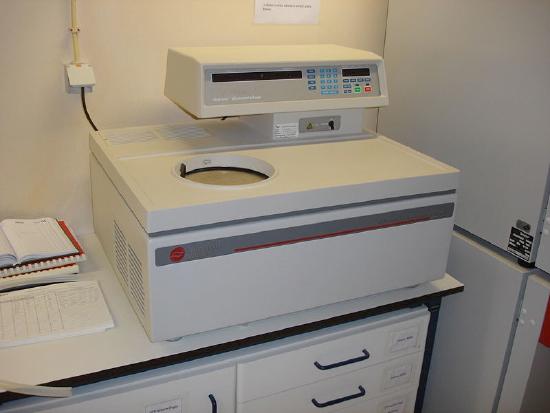

Column Chromatography
One powerful method used for this purpose is chromatography. We will consider several chromatographic approaches. Chromatography is used to separate out the components of a mixture based on differences in their size, charge or other characteristics. During chromatography, the mobile phase (buffer or other solvent) moves through the stationary phase (usually a solid matrix) carrying the components of the mixture. Separation of the components is achieved, because the different components move at different rates, for reasons that vary, depending on the type of chromatography used. We will consider several different kinds of chromatography to illustrate this process.
- Ion exchange chromatography
- Gel exclusion chromatography
- Affinity chromatography
- HPLC

These variations on chromatography are performed with the stationary phase held within so-called columns (Figure \(\PageIndex{1}\)). These are tubes containing the stationary phase (also called the “support” or solid phase).
Supports are composed of tiny beads suspended in buffer (Figure \(\PageIndex{3}\)) and are designed to exploit the chemistry or size differences of the components of the samples and thus provide a means of separation. Columns are “packed” or filled with the support, and a buffer or solvent carries the mixture of compounds to be separated through the support. Molecules in the sample interact differentially with the support and, consequently, travel through it at different speeds, thus enabling separation.

Ion exchange chromatography
In ion exchange chromatography, the support consists of tiny beads to which are attached chemicals possessing a charge. Before use, the beads are equilibrated in a solution containing an appropriate counter-ion to the charged molecule on the bead. Figure \(\PageIndex{5}\) shows the repeating unit of polystyrolsulfonate, a compound used as a cation exchange resin. As you can see, this molecule is negatively charged, and thus the beads would be equilibrated in a buffer containing a positively charged ion, say sodium. In the suspension, the negatively charged polystyrolsulfonate is unable to leave the beads, due to its covalent attachment, but the counter-ions (sodium) can be “exchanged” for molecules of the same charge.
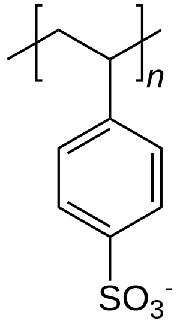
Exchanges
Thus, a cation exchange column will have positively charged counter-ions and negatively charged molecules covalently attached to the beads. Positively charged compounds from a cell lysate passed through the column will exchange with the counter-ions and “stick” to the negatively charged compounds covalently attached to the beads. Molecules in the sample that are neutral in charge or negatively charged will pass quickly through the column. At this point, only positively charged molecules from the original sample would be bound to the column. These may then be washed off, or eluted, by using buffers containing high concentrations of salt. Under these conditions, the interaction between the positively charged molecules and the polystyrosulfonate would be disrupted, allowing the molecules that were bound to the column to be recovered.
Anion exchange
On the other hand, in anion exchange chromatography, the chemicals attached to the beads are positively charged and the counterions are negatively charged (chloride, for example). Negatively charged molecules in the cell lysate will “stick” and other molecules will pass through quickly. To remove the molecules “stuck” to a column, one simply needs to add a high concentration of counter-ions to release them.
Uses
Ion exchange resins are useful for separating charged from uncharged, or oppositely charged, biomolecules in solution. The resins have a variety of other applications, including water purification and softening. Figure \(\PageIndex{5}\) shows use of a polystyrolsulfonate polymer in removing calcium for water softening.
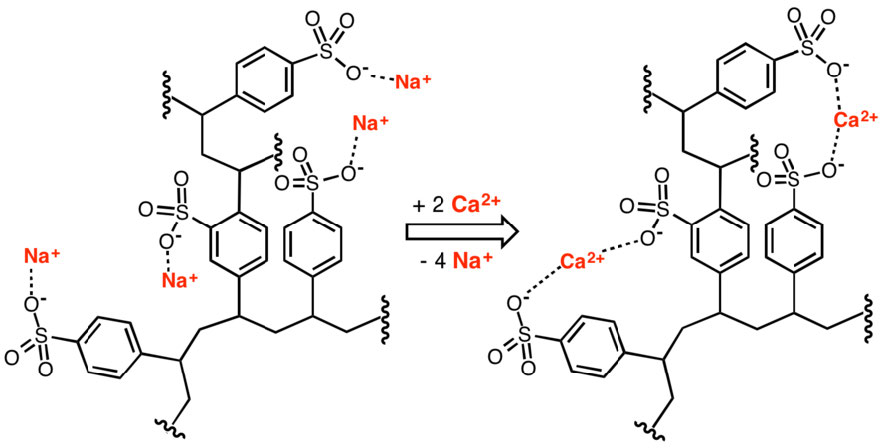
Figure \(\PageIndex{5}\): Removal of calcium ions by an ion exchanger. Wikipedia
Size Exclusion Chromatography
Size exclusion chromatography (also called molecular exclusion chromatography, gel exclusion chromatography, or gel filtration chromatography) is a low resolution separation method that employs beads with tiny “tunnels” in them that each have a precise opening. The size of the opening is referred to as an “exclusion limit,” which means that molecules above a certain molecular weight will not be able to pass through the tunnels. Molecules with physical sizes larger than the exclusion limit do not enter the tunnels and pass through the column relatively quickly, in the spaces outside the beads. Smaller molecules, which can enter the tunnels, do so, and thus, have a longer path that they take in passing through the column and elute last (Figure \(\PageIndex{6}\)).
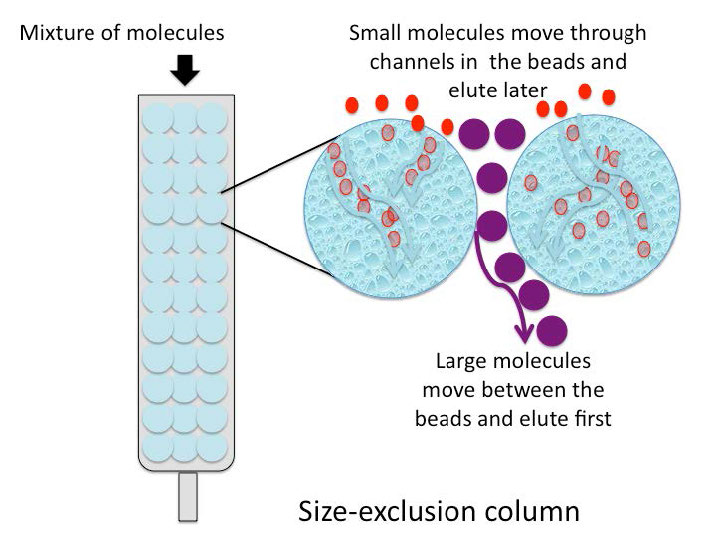
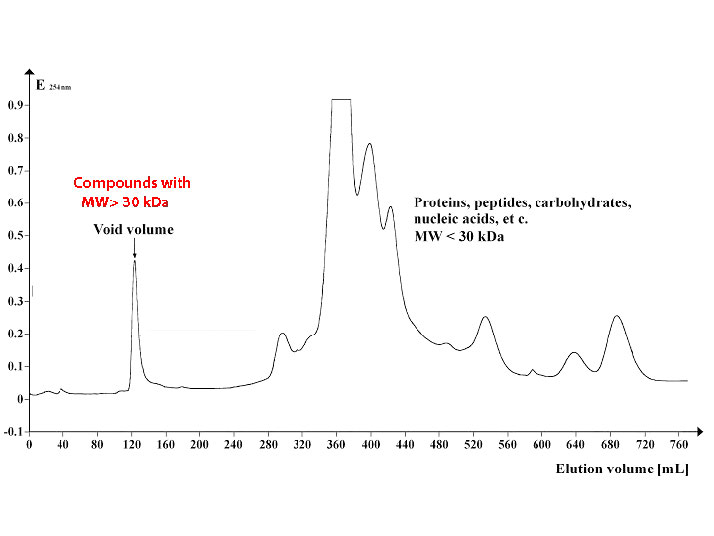
Figure \(\PageIndex{7}\) shows a profile of a group of proteins separated by size exclusion chromatography using beads with an exclusion limit of about 30,000 Daltons. Proteins 30,000 in molecular weight or larger elute in the void volume (left) while smaller proteins elute later (middle and right).
Affinity chromatography
Affinity chromatography is a very powerful and selective technique that exploits the binding affinities of sample molecules (typically proteins) for molecules covalently linked to the support beads. In contrast to ion-exchange chromatography, where all molecules of a given charge would bind to the column, affinity chromatography exploits the specific binding of a protein or proteins to a ligand that is immobilized on the beads in the column.
For example, if one wanted to separate all of the proteins in a cell lysate that bind to ATP from proteins that do not bind ATP, one could use a column that has ATP attached to the support beads and pass the sample through the column. All proteins that bind ATP will “stick” to the column, whereas those that do not bind ATP will pass quickly through it. The bound proteins may then be released from the column by adding a solution of ATP that will displace the bound proteins by competing, for the proteins, with the ATP attached to the column matrix.
Histidine tagging
Histidine tagging (His-tagging) is a special kind of affinity chromatography and is a powerful tool for isolating a recombinant protein from a cell lysate. His-tagging relies on altering the DNA coding region for a protein to add a series of at least six histidine residues to the amino or carboxyl terminal of the encoded protein. This “His-Tag” is useful in purifying the tagged protein because histidine side chains can bind to nickel or cobalt ions. Separation of His-tagged proteins from a cell lysate is relatively easy (Figure \(\PageIndex{8}\)).Passing the crude cell lysate through a column with nickel or cobalt attached to beads allows the His-tagged proteins to “stick,” while the remaining cell proteins all pass quickly through. The His-tagged proteins are then eluted by addition of imidazole to the column. Imidazole, which resembles the side chain of histidine, competes with the His-tagged proteins and displaces them from the column. Although non-tagged proteins in the lysate may also contain histidine as part of their sequence, they will not bind to the column as strongly as the His-tagged protein and will, thus, be displaced at lower imidazole concentrations than needed to elute the His-tagged protein. Surprisingly, many His-tagged proteins appear to function normally despite the added histidines, but if needed, the histidine tags may be cleaved from the purified protein by treatment with a protease that excises the added histidines, allowing the recovery of the desired protein with its native sequence.
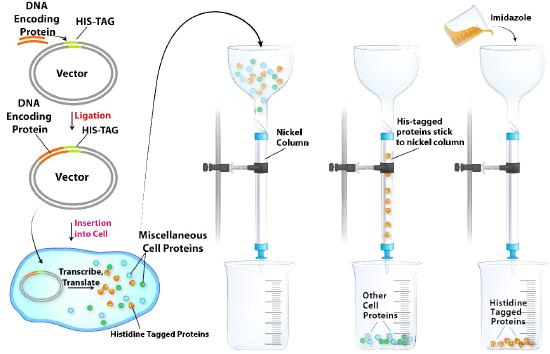
Figure \(\PageIndex{8}\): Affinity chromatographic purification of a protein by histidine tagging.Image by Aleia Kim
HPLC
High performance liquid chromatography (HPLC) is a powerful tool for separating a variety of molecules based on their differential polarities (Figure \(\PageIndex{9}\)). A more efficient form of column chromatography, it employs columns with tightly packed supports and very tiny beads such that flow of solvents/buffers through the columns requires high pressures. The supports used may be polar (normal phase separation) or non-polar (reverse phase separation). In normal phase separations, non-polar molecules elute first followed by the more polar compounds. This order is switched in reverse phase chromatography. Of the two, reverse phase is much more commonly employed due to more reproducible chromatographic profiles (separations) that it typically produces.
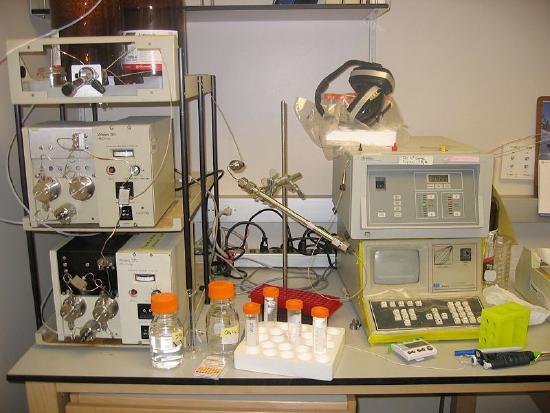

Figure \(\PageIndex{9}\): HPLC: Pumps on left/Column in center/Detector on the right. Wikipedia


